Rex Ying
DYSTIL: Dynamic Strategy Induction with Large Language Models for Reinforcement Learning
May 06, 2025



Abstract:Reinforcement learning from expert demonstrations has long remained a challenging research problem, and existing state-of-the-art methods using behavioral cloning plus further RL training often suffer from poor generalization, low sample efficiency, and poor model interpretability. Inspired by the strong reasoning abilities of large language models (LLMs), we propose a novel strategy-based reinforcement learning framework integrated with LLMs called DYnamic STrategy Induction with Llms for reinforcement learning (DYSTIL) to overcome these limitations. DYSTIL dynamically queries a strategy-generating LLM to induce textual strategies based on advantage estimations and expert demonstrations, and gradually internalizes induced strategies into the RL agent through policy optimization to improve its performance through boosting policy generalization and enhancing sample efficiency. It also provides a direct textual channel to observe and interpret the evolution of the policy's underlying strategies during training. We test DYSTIL over challenging RL environments from Minigrid and BabyAI, and empirically demonstrate that DYSTIL significantly outperforms state-of-the-art baseline methods by 17.75% in average success rate while also enjoying higher sample efficiency during the learning process.
Towards A Universal Graph Structural Encoder
Apr 15, 2025Abstract:Recent advancements in large-scale pre-training have shown the potential to learn generalizable representations for downstream tasks. In the graph domain, however, capturing and transferring structural information across different graph domains remains challenging, primarily due to the inherent differences in topological patterns across various contexts. Additionally, most existing models struggle to capture the complexity of rich graph structures, leading to inadequate exploration of the embedding space. To address these challenges, we propose GFSE, a universal graph structural encoder designed to capture transferable structural patterns across diverse domains such as molecular graphs, social networks, and citation networks. GFSE is the first cross-domain graph structural encoder pre-trained with multiple self-supervised learning objectives. Built on a Graph Transformer, GFSE incorporates attention mechanisms informed by graph inductive bias, enabling it to encode intricate multi-level and fine-grained topological features. The pre-trained GFSE produces generic and theoretically expressive positional and structural encoding for graphs, which can be seamlessly integrated with various downstream graph feature encoders, including graph neural networks for vectorized features and Large Language Models for text-attributed graphs. Comprehensive experiments on synthetic and real-world datasets demonstrate GFSE's capability to significantly enhance the model's performance while requiring substantially less task-specific fine-tuning. Notably, GFSE achieves state-of-the-art performance in 81.6% evaluated cases, spanning diverse graph models and datasets, highlighting its potential as a powerful and versatile encoder for graph-structured data.
Position: Beyond Euclidean -- Foundation Models Should Embrace Non-Euclidean Geometries
Apr 11, 2025Abstract:In the era of foundation models and Large Language Models (LLMs), Euclidean space has been the de facto geometric setting for machine learning architectures. However, recent literature has demonstrated that this choice comes with fundamental limitations. At a large scale, real-world data often exhibit inherently non-Euclidean structures, such as multi-way relationships, hierarchies, symmetries, and non-isotropic scaling, in a variety of domains, such as languages, vision, and the natural sciences. It is challenging to effectively capture these structures within the constraints of Euclidean spaces. This position paper argues that moving beyond Euclidean geometry is not merely an optional enhancement but a necessity to maintain the scaling law for the next-generation of foundation models. By adopting these geometries, foundation models could more efficiently leverage the aforementioned structures. Task-aware adaptability that dynamically reconfigures embeddings to match the geometry of downstream applications could further enhance efficiency and expressivity. Our position is supported by a series of theoretical and empirical investigations of prevalent foundation models.Finally, we outline a roadmap for integrating non-Euclidean geometries into foundation models, including strategies for building geometric foundation models via fine-tuning, training from scratch, and hybrid approaches.
HyperCore: The Core Framework for Building Hyperbolic Foundation Models with Comprehensive Modules
Apr 11, 2025



Abstract:Hyperbolic neural networks have emerged as a powerful tool for modeling hierarchical data across diverse modalities. Recent studies show that token distributions in foundation models exhibit scale-free properties, suggesting that hyperbolic space is a more suitable ambient space than Euclidean space for many pre-training and downstream tasks. However, existing tools lack essential components for building hyperbolic foundation models, making it difficult to leverage recent advancements. We introduce HyperCore, a comprehensive open-source framework that provides core modules for constructing hyperbolic foundation models across multiple modalities. HyperCore's modules can be effortlessly combined to develop novel hyperbolic foundation models, eliminating the need to extensively modify Euclidean modules from scratch and possible redundant research efforts. To demonstrate its versatility, we build and test the first fully hyperbolic vision transformers (LViT) with a fine-tuning pipeline, the first fully hyperbolic multimodal CLIP model (L-CLIP), and a hybrid Graph RAG with a hyperbolic graph encoder. Our experiments demonstrate that LViT outperforms its Euclidean counterpart. Additionally, we benchmark and reproduce experiments across hyperbolic GNNs, CNNs, Transformers, and vision Transformers to highlight HyperCore's advantages.
Beyond Feature Importance: Feature Interactions in Predicting Post-Stroke Rigidity with Graph Explainable AI
Apr 10, 2025Abstract:This study addresses the challenge of predicting post-stroke rigidity by emphasizing feature interactions through graph-based explainable AI. Post-stroke rigidity, characterized by increased muscle tone and stiffness, significantly affects survivors' mobility and quality of life. Despite its prevalence, early prediction remains limited, delaying intervention. We analyze 519K stroke hospitalization records from the Healthcare Cost and Utilization Project dataset, where 43% of patients exhibited rigidity. We compare traditional approaches such as Logistic Regression, XGBoost, and Transformer with graph-based models like Graphormer and Graph Attention Network. These graph models inherently capture feature interactions and incorporate intrinsic or post-hoc explainability. Our results show that graph-based methods outperform others (AUROC 0.75), identifying key predictors such as NIH Stroke Scale and APR-DRG mortality risk scores. They also uncover interactions missed by conventional models. This research provides a novel application of graph-based XAI in stroke prognosis, with potential to guide early identification and personalized rehabilitation strategies.
Mixture-of-Personas Language Models for Population Simulation
Apr 07, 2025



Abstract:Advances in Large Language Models (LLMs) paved the way for their emerging applications in various domains, such as human behavior simulations, where LLMs could augment human-generated data in social science research and machine learning model training. However, pretrained LLMs often fail to capture the behavioral diversity of target populations due to the inherent variability across individuals and groups. To address this, we propose \textit{Mixture of Personas} (MoP), a \textit{probabilistic} prompting method that aligns the LLM responses with the target population. MoP is a contextual mixture model, where each component is an LM agent characterized by a persona and an exemplar representing subpopulation behaviors. The persona and exemplar are randomly chosen according to the learned mixing weights to elicit diverse LLM responses during simulation. MoP is flexible, requires no model finetuning, and is transferable across base models. Experiments for synthetic data generation show that MoP outperforms competing methods in alignment and diversity metrics.
MTBench: A Multimodal Time Series Benchmark for Temporal Reasoning and Question Answering
Mar 21, 2025


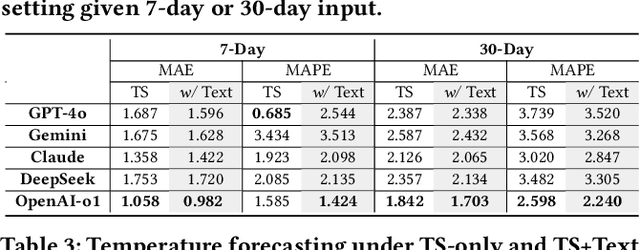
Abstract:Understanding the relationship between textual news and time-series evolution is a critical yet under-explored challenge in applied data science. While multimodal learning has gained traction, existing multimodal time-series datasets fall short in evaluating cross-modal reasoning and complex question answering, which are essential for capturing complex interactions between narrative information and temporal patterns. To bridge this gap, we introduce Multimodal Time Series Benchmark (MTBench), a large-scale benchmark designed to evaluate large language models (LLMs) on time series and text understanding across financial and weather domains. MTbench comprises paired time series and textual data, including financial news with corresponding stock price movements and weather reports aligned with historical temperature records. Unlike existing benchmarks that focus on isolated modalities, MTbench provides a comprehensive testbed for models to jointly reason over structured numerical trends and unstructured textual narratives. The richness of MTbench enables formulation of diverse tasks that require a deep understanding of both text and time-series data, including time-series forecasting, semantic and technical trend analysis, and news-driven question answering (QA). These tasks target the model's ability to capture temporal dependencies, extract key insights from textual context, and integrate cross-modal information. We evaluate state-of-the-art LLMs on MTbench, analyzing their effectiveness in modeling the complex relationships between news narratives and temporal patterns. Our findings reveal significant challenges in current models, including difficulties in capturing long-term dependencies, interpreting causality in financial and weather trends, and effectively fusing multimodal information.
Large-Scale AI in Telecom: Charting the Roadmap for Innovation, Scalability, and Enhanced Digital Experiences
Mar 06, 2025



Abstract:This white paper discusses the role of large-scale AI in the telecommunications industry, with a specific focus on the potential of generative AI to revolutionize network functions and user experiences, especially in the context of 6G systems. It highlights the development and deployment of Large Telecom Models (LTMs), which are tailored AI models designed to address the complex challenges faced by modern telecom networks. The paper covers a wide range of topics, from the architecture and deployment strategies of LTMs to their applications in network management, resource allocation, and optimization. It also explores the regulatory, ethical, and standardization considerations for LTMs, offering insights into their future integration into telecom infrastructure. The goal is to provide a comprehensive roadmap for the adoption of LTMs to enhance scalability, performance, and user-centric innovation in telecom networks.
HiPoNet: A Topology-Preserving Multi-View Neural Network For High Dimensional Point Cloud and Single-Cell Data
Feb 11, 2025



Abstract:In this paper, we propose HiPoNet, an end-to-end differentiable neural network for regression, classification, and representation learning on high-dimensional point clouds. Single-cell data can have high dimensionality exceeding the capabilities of existing methods point cloud tailored for 3D data. Moreover, modern single-cell and spatial experiments now yield entire cohorts of datasets (i.e. one on every patient), necessitating models that can process large, high-dimensional point clouds at scale. Most current approaches build a single nearest-neighbor graph, discarding important geometric information. In contrast, HiPoNet forms higher-order simplicial complexes through learnable feature reweighting, generating multiple data views that disentangle distinct biological processes. It then employs simplicial wavelet transforms to extract multi-scale features - capturing both local and global topology. We empirically show that these components preserve topological information in the learned representations, and that HiPoNet significantly outperforms state-of-the-art point-cloud and graph-based models on single cell. We also show an application of HiPoNet on spatial transcriptomics datasets using spatial co-ordinates as one of the views. Overall, HiPoNet offers a robust and scalable solution for high-dimensional data analysis.
Flow Matching for Collaborative Filtering
Feb 11, 2025Abstract:Generative models have shown great promise in collaborative filtering by capturing the underlying distribution of user interests and preferences. However, existing approaches struggle with inaccurate posterior approximations and misalignment with the discrete nature of recommendation data, limiting their expressiveness and real-world performance. To address these limitations, we propose FlowCF, a novel flow-based recommendation system leveraging flow matching for collaborative filtering. We tailor flow matching to the unique challenges in recommendation through two key innovations: (1) a behavior-guided prior that aligns with user behavior patterns to handle the sparse and heterogeneous user-item interactions, and (2) a discrete flow framework to preserve the binary nature of implicit feedback while maintaining the benefits of flow matching, such as stable training and efficient inference. Extensive experiments demonstrate that FlowCF achieves state-of-the-art recommendation accuracy across various datasets with the fastest inference speed, making it a compelling approach for real-world recommender systems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge