Qiming Yang
Thousand-GPU Large-Scale Training and Optimization Recipe for AI-Native Cloud Embodied Intelligence Infrastructure
Mar 11, 2026Abstract:Embodied intelligence is a key step towards Artificial General Intelligence (AGI), yet its development faces multiple challenges including data, frameworks, infrastructure, and evaluation systems. To address these issues, we have, for the first time in the industry, launched a cloud-based, thousand-GPU distributed training platform for embodied intelligence, built upon the widely adopted LeRobot framework, and have systematically overcome bottlenecks across the entire pipeline. At the data layer, we have restructured the data pipeline to optimize the flow of embodied training data. In terms of training, for the GR00T-N1.5 model, utilizing thousand-GPU clusters and data at the scale of hundreds of millions, the single-round training time has been reduced from 15 hours to just 22 minutes, achieving a 40-fold speedup. At the model layer, by combining variable-length FlashAttention and Data Packing, we have moved from sample redundancy to sequence integration, resulting in a 188% speed increase; π-0.5 attention optimization has accelerated training by 165%; and FP8 quantization has delivered a 140% speedup. On the infrastructure side, relying on high-performance storage, a 3.2T RDMA network, and a Ray-driven elastic AI data lake, we have achieved deep synergy among data, storage, communication, and computation. We have also built an end-to-end evaluation system, creating a closed loop from training to simulation to assessment. This framework has already been fully validated on thousand-GPU clusters, laying a crucial technical foundation for the development and application of next-generation autonomous intelligent robots, and is expected to accelerate the arrival of the era of human-machine integration.
RL-VLA$^3$: Reinforcement Learning VLA Accelerating via Full Asynchronism
Feb 05, 2026Abstract:In recent years, Vision-Language-Action (VLA) models have emerged as a crucial pathway towards general embodied intelligence, yet their training efficiency has become a key bottleneck. Although existing reinforcement learning (RL)-based training frameworks like RLinf can enhance model generalization, they still rely on synchronous execution, leading to severe resource underutilization and throughput limitations during environment interaction, policy generation (rollout), and model update phases (actor). To overcome this challenge, this paper, for the first time, proposes and implements a fully-asynchronous policy training framework encompassing the entire pipeline from environment interaction, rollout generation, to actor policy updates. Systematically drawing inspiration from asynchronous optimization ideas in large model RL, our framework designs a multi-level decoupled architecture. This includes asynchronous parallelization of environment interaction and trajectory collection, streaming execution for policy generation, and decoupled scheduling for training updates. We validated the effectiveness of our method across diverse VLA models and environments. On the LIBERO benchmark, the framework achieves throughput improvements of up to 59.25\% compared to existing synchronous strategies. When deeply optimizing separation strategies, throughput can be increased by as much as 126.67\%. We verified the effectiveness of each asynchronous component via ablation studies. Scaling law validation across 8 to 256 GPUs demonstrates our method's excellent scalability under most conditions.
Research on Improved U-net Based Remote Sensing Image Segmentation Algorithm
Aug 22, 2024



Abstract:In recent years, although U-Net network has made significant progress in the field of image segmentation, it still faces performance bottlenecks in remote sensing image segmentation. In this paper, we innovatively propose to introduce SimAM and CBAM attention mechanism in U-Net, and the experimental results show that after adding SimAM and CBAM modules alone, the model improves 17.41% and 12.23% in MIoU, and the Mpa and Accuracy are also significantly improved. And after fusing the two,the model performance jumps up to 19.11% in MIoU, and the Mpa and Accuracy are also improved by 16.38% and 14.8% respectively, showing excellent segmentation accuracy and visual effect with strong generalization ability and robustness. This study opens up a new path for remote sensing image segmentation technology and has important reference value for algorithm selection and improvement.
A Self-attention Knowledge Domain Adaptation Network for Commercial Lithium-ion Batteries State-of-health Estimation under Shallow Cycles
Apr 11, 2023



Abstract:Accurate state-of-health (SOH) estimation is critical to guarantee the safety, efficiency and reliability of battery-powered applications. Most SOH estimation methods focus on the 0-100\% full state-of-charge (SOC) range that has similar distributions. However, the batteries in real-world applications usually work in the partial SOC range under shallow-cycle conditions and follow different degradation profiles with no labeled data available, thus making SOH estimation challenging. To estimate shallow-cycle battery SOH, a novel unsupervised deep transfer learning method is proposed to bridge different domains using self-attention distillation module and multi-kernel maximum mean discrepancy technique. The proposed method automatically extracts domain-variant features from charge curves to transfer knowledge from the large-scale labeled full cycles to the unlabeled shallow cycles. The CALCE and SNL battery datasets are employed to verify the effectiveness of the proposed method to estimate the battery SOH for different SOC ranges, temperatures, and discharge rates. The proposed method achieves a root-mean-square error within 2\% and outperforms other transfer learning methods for different SOC ranges. When applied to batteries with different operating conditions and from different manufacturers, the proposed method still exhibits superior SOH estimation performance. The proposed method is the first attempt at accurately estimating battery SOH under shallow-cycle conditions without needing a full-cycle characteristic test.
Unified Normalization for Accelerating and Stabilizing Transformers
Aug 02, 2022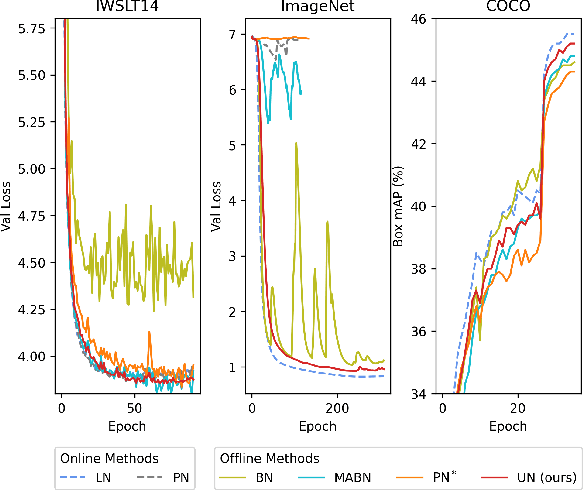
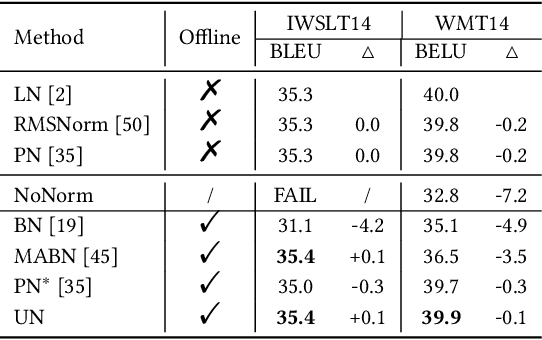
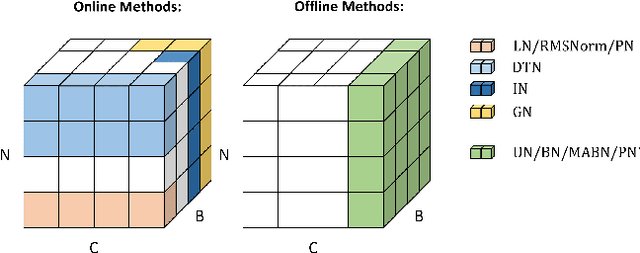
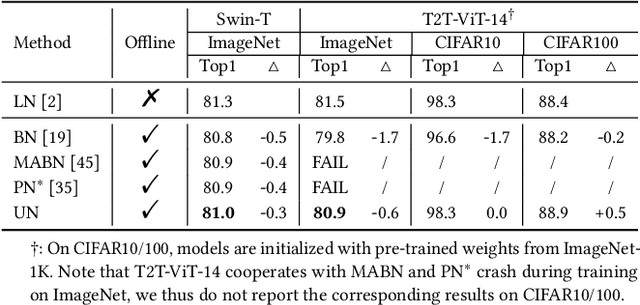
Abstract:Solid results from Transformers have made them prevailing architectures in various natural language and vision tasks. As a default component in Transformers, Layer Normalization (LN) normalizes activations within each token to boost the robustness. However, LN requires on-the-fly statistics calculation in inference as well as division and square root operations, leading to inefficiency on hardware. What is more, replacing LN with other hardware-efficient normalization schemes (e.g., Batch Normalization) results in inferior performance, even collapse in training. We find that this dilemma is caused by abnormal behaviors of activation statistics, including large fluctuations over iterations and extreme outliers across layers. To tackle these issues, we propose Unified Normalization (UN), which can speed up the inference by being fused with other linear operations and achieve comparable performance on par with LN. UN strives to boost performance by calibrating the activation and gradient statistics with a tailored fluctuation smoothing strategy. Meanwhile, an adaptive outlier filtration strategy is applied to avoid collapse in training whose effectiveness is theoretically proved and experimentally verified in this paper. We demonstrate that UN can be an efficient drop-in alternative to LN by conducting extensive experiments on language and vision tasks. Besides, we evaluate the efficiency of our method on GPU. Transformers equipped with UN enjoy about 31% inference speedup and nearly 18% memory reduction. Code will be released at https://github.com/hikvision-research/Unified-Normalization.
Temporal Link Prediction via Adjusted Sigmoid Function and 2-Simplex Sructure
Jun 20, 2022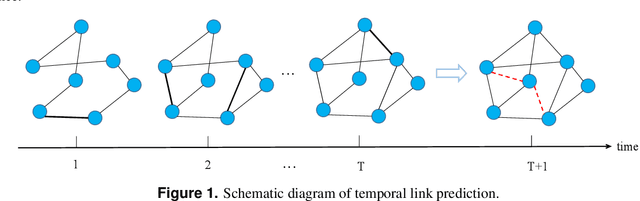
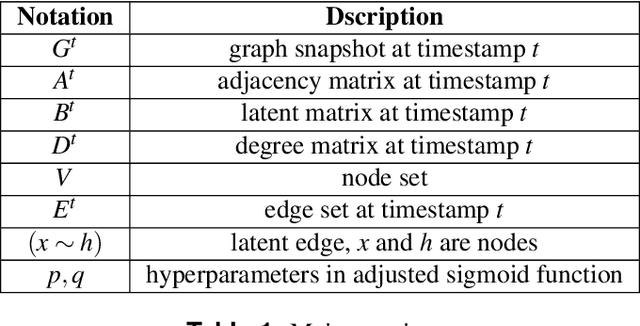
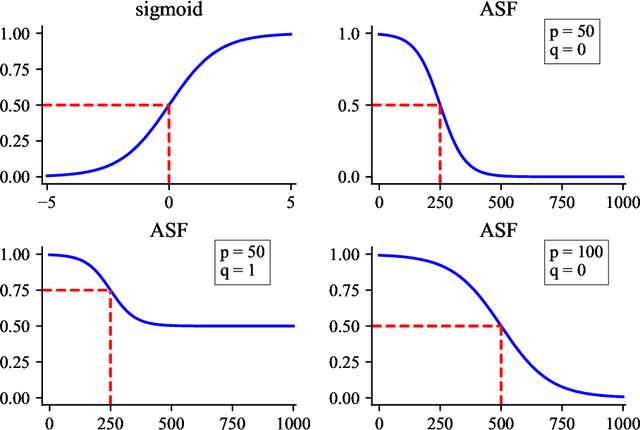
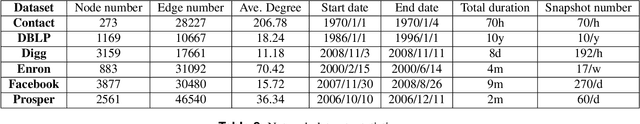
Abstract:Temporal network link prediction is an important task in the field of network science, and has a wide range of applications in practical scenarios. Revealing the evolutionary mechanism of the network is essential for link prediction, and how to effectively utilize the historical information for temporal links and efficiently extract the high-order patterns of network structure remains a vital challenge. To address these issues, in this paper, we propose a novel temporal link prediction model with adjusted sigmoid function and 2-simplex structure (TLPSS). The adjusted sigmoid decay mode takes the active, decay and stable states of edges into account, which properly fits the life cycle of information. Moreover, the latent matrix sequence is introduced, which is composed of simplex high-order structure, to enhance the performance of link prediction method since it is highly feasible in sparse network. Combining the life cycle of information and simplex high-order structure, the overall performance of TLPSS is achieved by satisfying the consistency of temporal and structural information in dynamic networks. Experimental results on six real-world datasets demonstrate the effectiveness of TLPSS, and our proposed model improves the performance of link prediction by an average of 15% compared to other baseline methods.
Enhance Ambiguous Community Structure via Multi-strategy Community Related Link Prediction Method with Evolutionary Process
Apr 28, 2022Abstract:Most real-world networks suffer from incompleteness or incorrectness, which is an inherent attribute to real-world datasets. As a consequence, those downstream machine learning tasks in complex network like community detection methods may yield less satisfactory results, i.e., a proper preprocessing measure is required here. To address this issue, in this paper, we design a new community attribute based link prediction strategy HAP and propose a two-step community enhancement algorithm with automatic evolution process based on HAP. This paper aims at providing a community enhancement measure through adding links to clarify ambiguous community structures. The HAP method takes the neighbourhood uncertainty and Shannon entropy to identify boundary nodes, and establishes links by considering the nodes' community attributes and community size at the same time. The experimental results on twelve real-world datasets with ground truth community indicate that the proposed link prediction method outperforms other baseline methods and the enhancement of community follows the expected evolution process.
Mining Domain Knowledge: Improved Framework towards Automatically Standardizing Anatomical Structure Nomenclature in Radiotherapy
Dec 04, 2019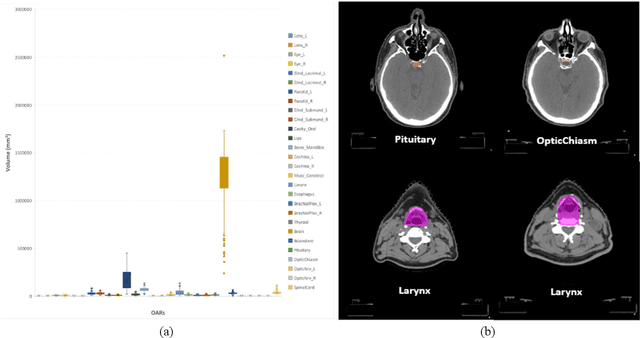
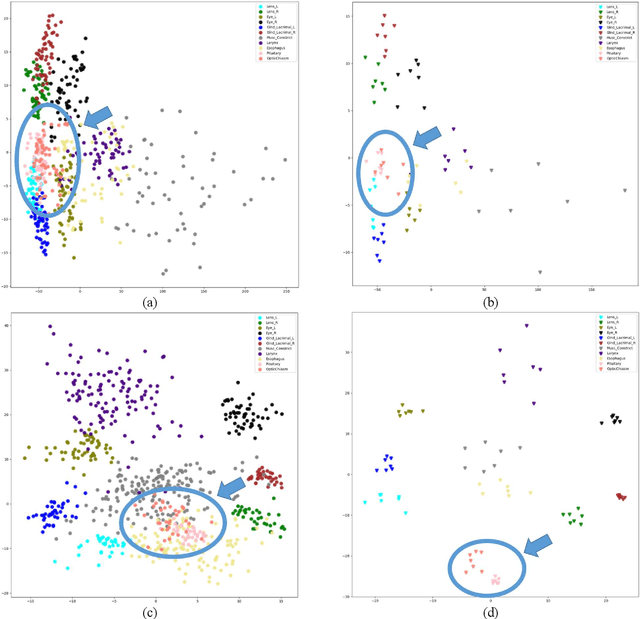
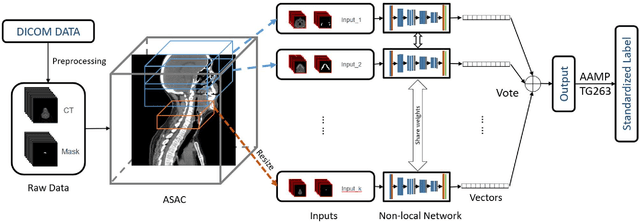
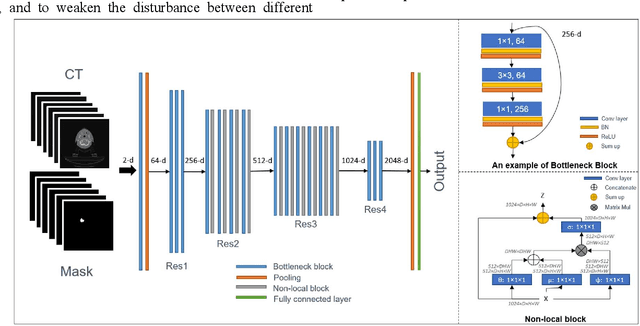
Abstract:Automatically standardizing nomenclature for anatomical structures in radiotherapy (RT) clinical data is an unmet urgent need in the era of big data and artificial intelligence. Existing methods either can hardly handle cross-institutional datasets or suffer from heavy imbalance and poor-quality delineation in clinical RT datasets. To solve these problems, we propose an automated structure nomenclature standardization framework, 3DNNV, which consists of an improved data processing strategy (ASAC/Voting) and an optimized feature extraction module to simulate clinicians' domain knowledge and recognition mechanisms to identify heavily imbalanced small-volume organs at risk (OARs) better than other methods. We used partial data from an open-source head-and-neck cancer dataset (HN_PETCT) to train the model, then tested the model on three cross-institutional datasets to demonstrate its generalizability. 3DNNV outperformed the baseline model (ResNet50), achieving a significantly higher average true positive rate (TPR) on the three test datasets (+8.27%, +2.39%, +5.53%). More importantly, the 3DNNV outperformed the baseline, 28.63% to 91.17%, on the F1 score of a small-volume OAR with only 9 training samples, when tested on the HN_UTSW dataset. The developed framework can be used to help standardizing structure nomenclature to facilitate data-driven clinical research in cancer radiotherapy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge