Nassir Navab
Computer Aided Medical Procedures, Technische Universit Munchen, Germany, Johns Hopkins University, Baltimore MD, USA
Unsupervised Pre-Training on Patient Population Graphs for Patient-Level Predictions
Mar 23, 2022
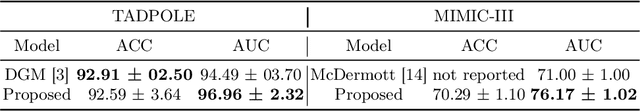

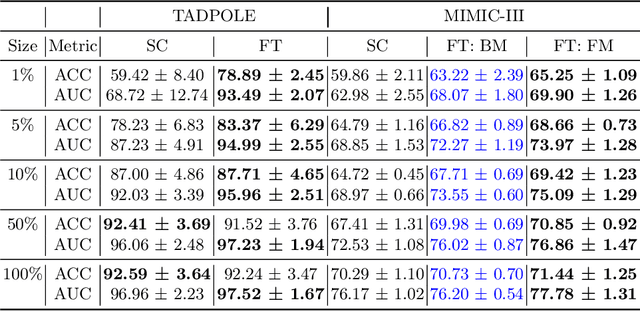
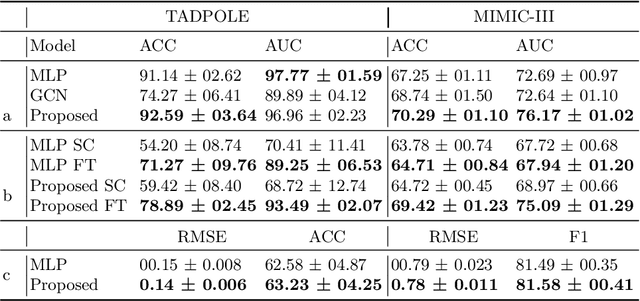
Abstract:Pre-training has shown success in different areas of machine learning, such as Computer Vision (CV), Natural Language Processing (NLP) and medical imaging. However, it has not been fully explored for clinical data analysis. Even though an immense amount of Electronic Health Record (EHR) data is recorded, data and labels can be scarce if the data is collected in small hospitals or deals with rare diseases. In such scenarios, pre-training on a larger set of EHR data could improve the model performance. In this paper, we apply unsupervised pre-training to heterogeneous, multi-modal EHR data for patient outcome prediction. To model this data, we leverage graph deep learning over population graphs. We first design a network architecture based on graph transformer designed to handle various input feature types occurring in EHR data, like continuous, discrete, and time-series features, allowing better multi-modal data fusion. Further, we design pre-training methods based on masked imputation to pre-train our network before fine-tuning on different end tasks. Pre-training is done in a fully unsupervised fashion, which lays the groundwork for pre-training on large public datasets with different tasks and similar modalities in the future. We test our method on two medical datasets of patient records, TADPOLE and MIMIC-III, including imaging and non-imaging features and different prediction tasks. We find that our proposed graph based pre-training method helps in modeling the data at a population level and further improves performance on the fine tuning tasks in terms of AUC on average by 4.15% for MIMIC and 7.64% for TADPOLE.
4D-OR: Semantic Scene Graphs for OR Domain Modeling
Mar 22, 2022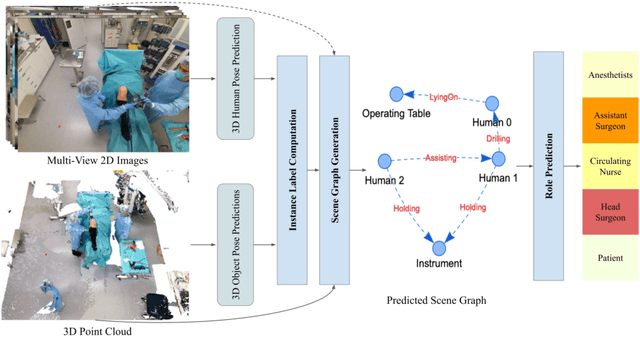
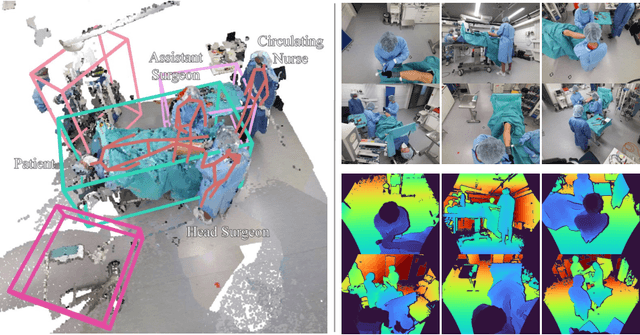


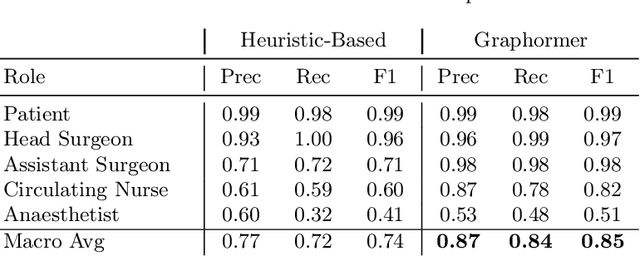
Abstract:Surgical procedures are conducted in highly complex operating rooms (OR), comprising different actors, devices, and interactions. To date, only medically trained human experts are capable of understanding all the links and interactions in such a demanding environment. This paper aims to bring the community one step closer to automated, holistic and semantic understanding and modeling of OR domain. Towards this goal, for the first time, we propose using semantic scene graphs (SSG) to describe and summarize the surgical scene. The nodes of the scene graphs represent different actors and objects in the room, such as medical staff, patients, and medical equipment, whereas edges are the relationships between them. To validate the possibilities of the proposed representation, we create the first publicly available 4D surgical SSG dataset, 4D-OR, containing ten simulated total knee replacement surgeries recorded with six RGB-D sensors in a realistic OR simulation center. 4D-OR includes 6734 frames and is richly annotated with SSGs, human and object poses, and clinical roles. We propose an end-to-end neural network-based SSG generation pipeline, with a rate of success of 0.75 macro F1, indeed being able to infer semantic reasoning in the OR. We further demonstrate the representation power of our scene graphs by using it for the problem of clinical role prediction, where we achieve 0.85 macro F1. The code and dataset will be made available upon acceptance.
Conditional Generative Data Augmentation for Clinical Audio Datasets
Mar 22, 2022
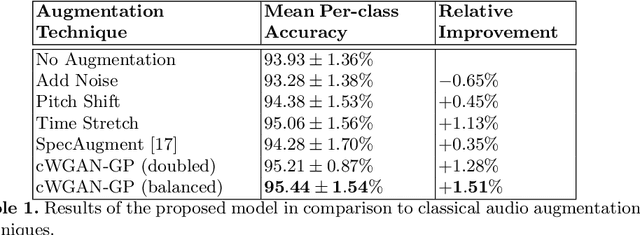
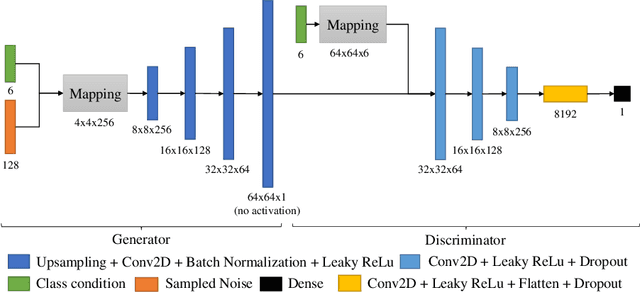
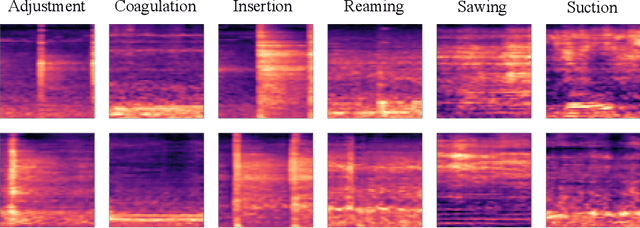
Abstract:In this work, we propose a novel data augmentation method for clinical audio datasets based on a conditional Wasserstein Generative Adversarial Network with Gradient Penalty (cWGAN-GP), operating on log-mel spectrograms. To validate our method, we created a clinical audio dataset which was recorded in a real-world operating room during Total Hip Arthroplasty (THA) procedures and contains typical sounds which resemble the different phases of the intervention. We demonstrate the capability of the proposed method to generate realistic class-conditioned samples from the dataset distribution and show that training with the generated augmented samples outperforms classical audio augmentation methods in terms of classification accuracy. The performance was evaluated using a ResNet-18 classifier which shows a mean per-class accuracy improvement of 1.51% in a 5-fold cross validation experiment using the proposed augmentation method. Because clinical data is often expensive to acquire, the development of realistic and high-quality data augmentation methods is crucial to improve the robustness and generalization capabilities of learning-based algorithms which is especially important for safety-critical medical applications. Therefore, the proposed data augmentation method is an important step towards improving the data bottleneck for clinical audio-based machine learning systems. The code and dataset will be published upon acceptance.
Know your sensORs -- A Modality Study For Surgical Action Classification
Mar 22, 2022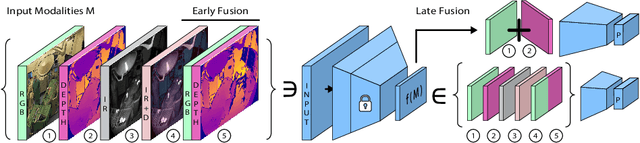
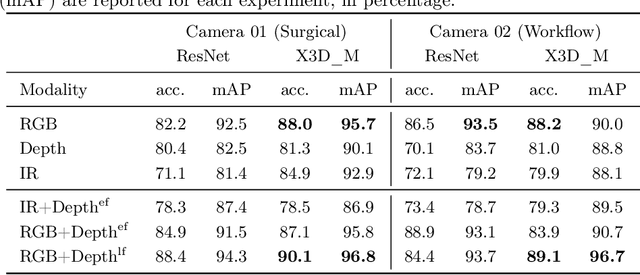
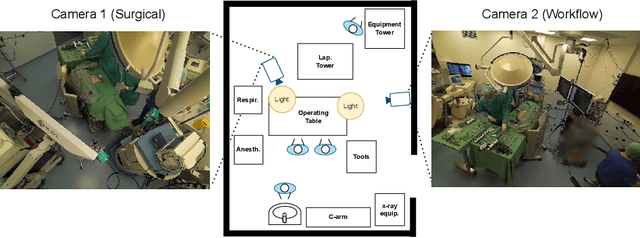
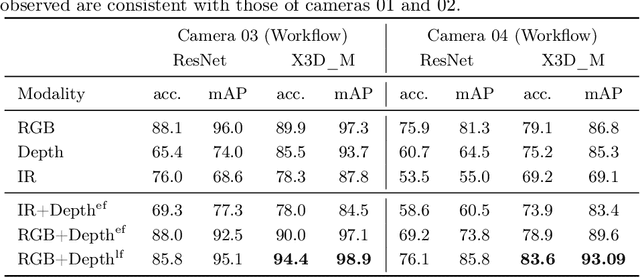
Abstract:The surgical operating room (OR) presents many opportunities for automation and optimization. Videos from various sources in the OR are becoming increasingly available. The medical community seeks to leverage this wealth of data to develop automated methods to advance interventional care, lower costs, and improve overall patient outcomes. Existing datasets from OR room cameras are thus far limited in size or modalities acquired, leaving it unclear which sensor modalities are best suited for tasks such as recognizing surgical action from videos. This study demonstrates that surgical action recognition performance can vary depending on the image modalities used. We perform a methodical analysis on several commonly available sensor modalities, presenting two fusion approaches that improve classification performance. The analyses are carried out on a set of multi-view RGB-D video recordings of 18 laparoscopic procedures.
Longitudinal Self-Supervision for COVID-19 Pathology Quantification
Mar 21, 2022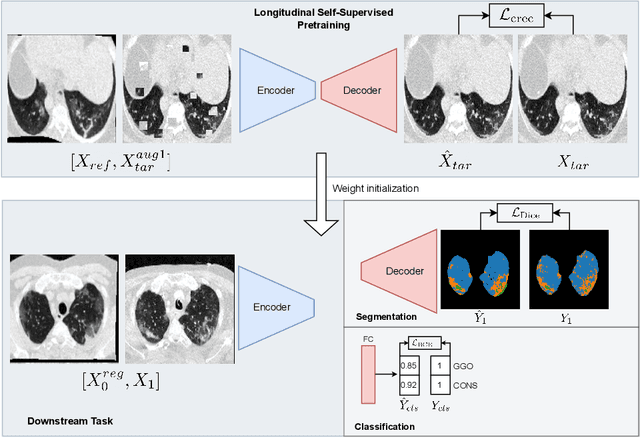
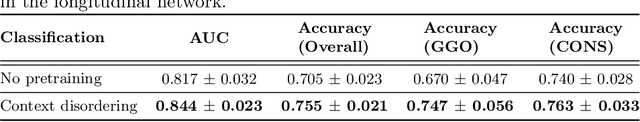
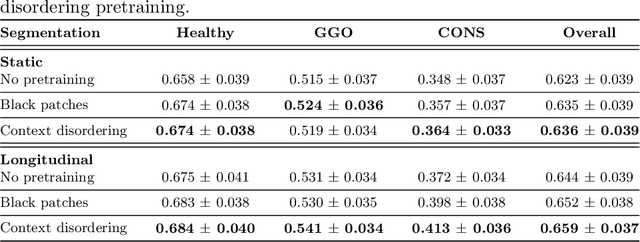
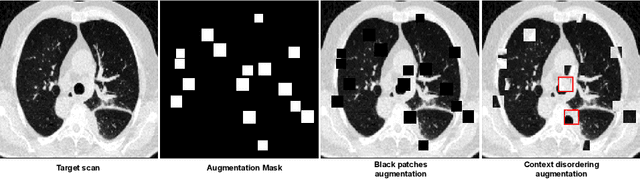

Abstract:Quantifying COVID-19 infection over time is an important task to manage the hospitalization of patients during a global pandemic. Recently, deep learning-based approaches have been proposed to help radiologists automatically quantify COVID-19 pathologies on longitudinal CT scans. However, the learning process of deep learning methods demands extensive training data to learn the complex characteristics of infected regions over longitudinal scans. It is challenging to collect a large-scale dataset, especially for longitudinal training. In this study, we want to address this problem by proposing a new self-supervised learning method to effectively train longitudinal networks for the quantification of COVID-19 infections. For this purpose, longitudinal self-supervision schemes are explored on clinical longitudinal COVID-19 CT scans. Experimental results show that the proposed method is effective, helping the model better exploit the semantics of longitudinal data and improve two COVID-19 quantification tasks.
GPV-Pose: Category-level Object Pose Estimation via Geometry-guided Point-wise Voting
Mar 17, 2022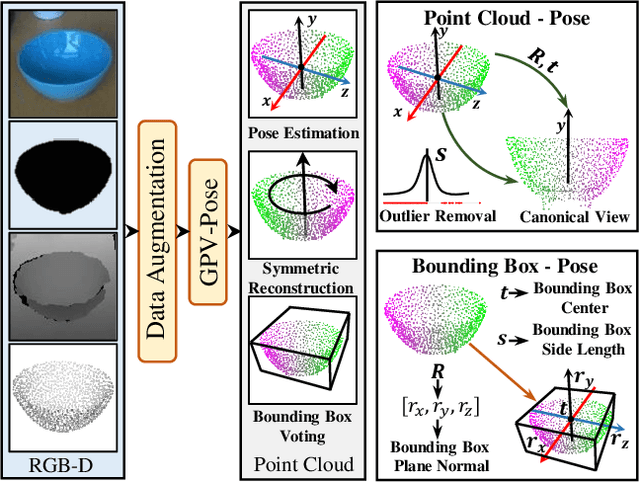
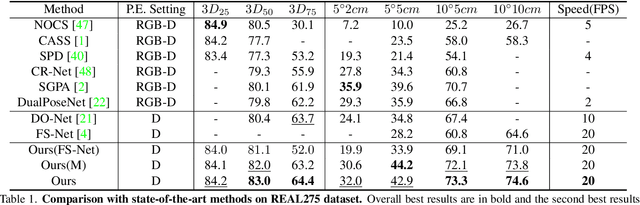
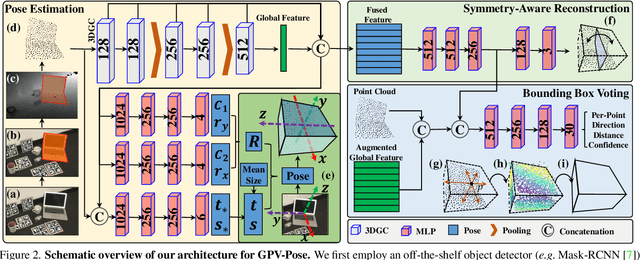
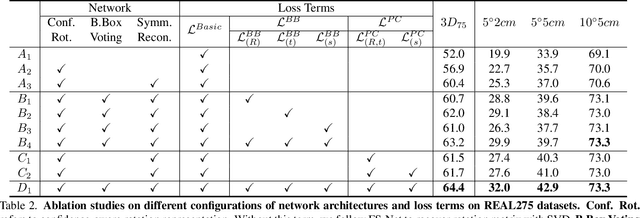
Abstract:While 6D object pose estimation has recently made a huge leap forward, most methods can still only handle a single or a handful of different objects, which limits their applications. To circumvent this problem, category-level object pose estimation has recently been revamped, which aims at predicting the 6D pose as well as the 3D metric size for previously unseen instances from a given set of object classes. This is, however, a much more challenging task due to severe intra-class shape variations. To address this issue, we propose GPV-Pose, a novel framework for robust category-level pose estimation, harnessing geometric insights to enhance the learning of category-level pose-sensitive features. First, we introduce a decoupled confidence-driven rotation representation, which allows geometry-aware recovery of the associated rotation matrix. Second, we propose a novel geometry-guided point-wise voting paradigm for robust retrieval of the 3D object bounding box. Finally, leveraging these different output streams, we can enforce several geometric consistency terms, further increasing performance, especially for non-symmetric categories. GPV-Pose produces superior results to state-of-the-art competitors on common public benchmarks, whilst almost achieving real-time inference speed at 20 FPS.
From 2D to 3D: Re-thinking Benchmarking of Monocular Depth Prediction
Mar 15, 2022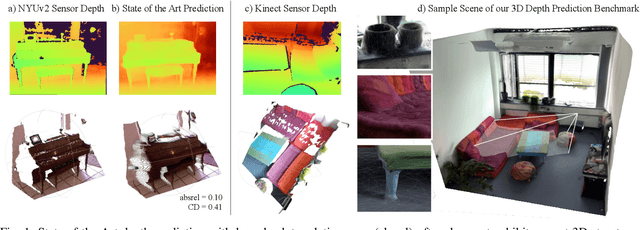
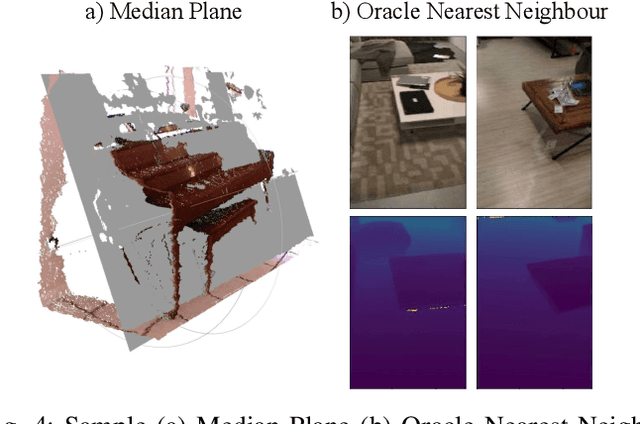
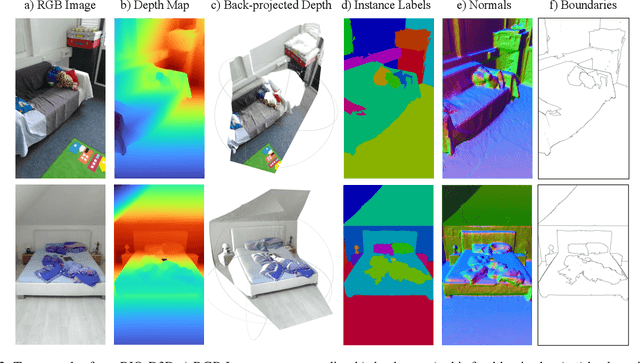
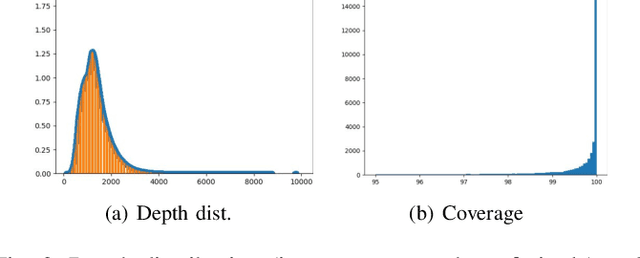

Abstract:There have been numerous recently proposed methods for monocular depth prediction (MDP) coupled with the equally rapid evolution of benchmarking tools. However, we argue that MDP is currently witnessing benchmark over-fitting and relying on metrics that are only partially helpful to gauge the usefulness of the predictions for 3D applications. This limits the design and development of novel methods that are truly aware of - and improving towards estimating - the 3D structure of the scene rather than optimizing 2D-based distances. In this work, we aim to bring structural awareness to MDP, an inherently 3D task, by exhibiting the limits of evaluation metrics towards assessing the quality of the 3D geometry. We propose a set of metrics well suited to evaluate the 3D geometry of MDP approaches and a novel indoor benchmark, RIO-D3D, crucial for the proposed evaluation methodology. Our benchmark is based on a real-world dataset featuring high-quality rendered depth maps obtained from RGB-D reconstructions. We further demonstrate this to help benchmark the closely-tied task of 3D scene completion.
Do Explanations Explain? Model Knows Best
Mar 04, 2022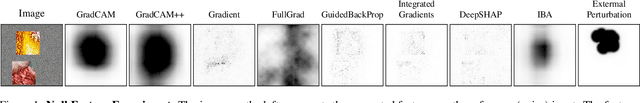
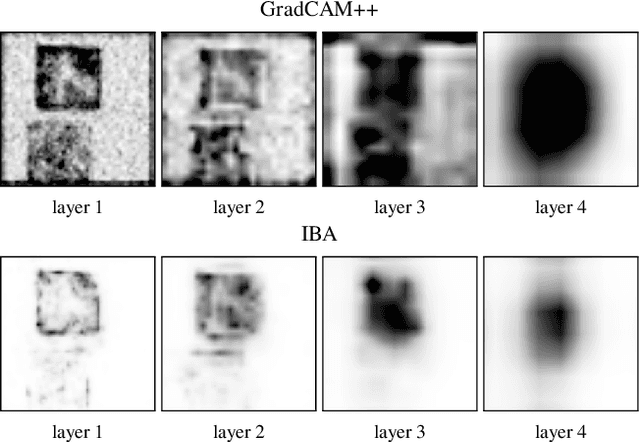
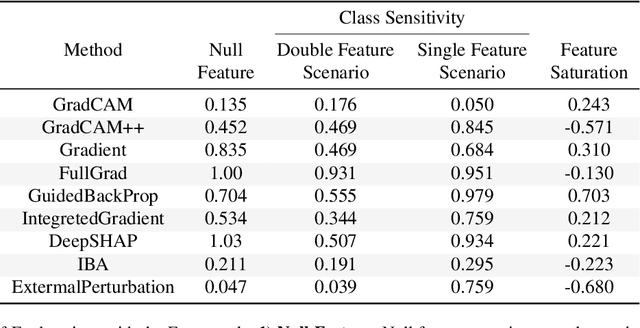
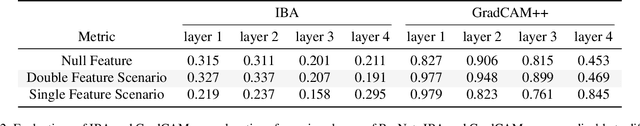

Abstract:It is a mystery which input features contribute to a neural network's output. Various explanation (feature attribution) methods are proposed in the literature to shed light on the problem. One peculiar observation is that these explanations (attributions) point to different features as being important. The phenomenon raises the question, which explanation to trust? We propose a framework for evaluating the explanations using the neural network model itself. The framework leverages the network to generate input features that impose a particular behavior on the output. Using the generated features, we devise controlled experimental setups to evaluate whether an explanation method conforms to an axiom. Thus we propose an empirical framework for axiomatic evaluation of explanation methods. We evaluate well-known and promising explanation solutions using the proposed framework. The framework provides a toolset to reveal properties and drawbacks within existing and future explanation solutions.
Bending Graphs: Hierarchical Shape Matching using Gated Optimal Transport
Feb 03, 2022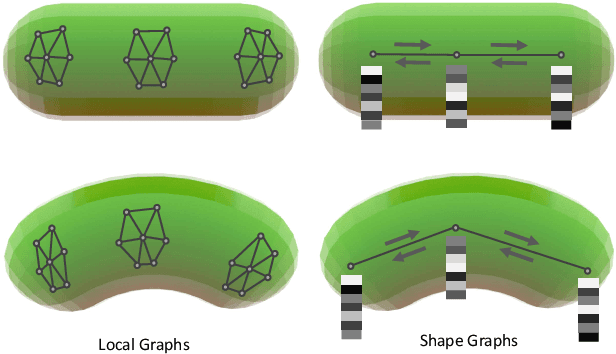
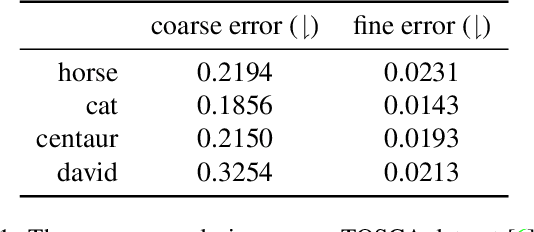

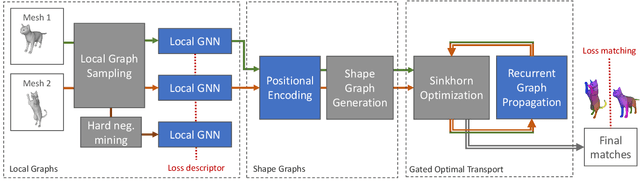
Abstract:Shape matching has been a long-studied problem for the computer graphics and vision community. The objective is to predict a dense correspondence between meshes that have a certain degree of deformation. Existing methods either consider the local description of sampled points or discover correspondences based on global shape information. In this work, we investigate a hierarchical learning design, to which we incorporate local patch-level information and global shape-level structures. This flexible representation enables correspondence prediction and provides rich features for the matching stage. Finally, we propose a novel optimal transport solver by recurrently updating features on non-confident nodes to learn globally consistent correspondences between the shapes. Our results on publicly available datasets suggest robust performance in presence of severe deformations without the need for extensive training or refinement.
Transformers in Action: Weakly Supervised Action Segmentation
Jan 20, 2022



Abstract:The video action segmentation task is regularly explored under weaker forms of supervision, such as transcript supervision, where a list of actions is easier to obtain than dense frame-wise labels. In this formulation, the task presents various challenges for sequence modeling approaches due to the emphasis on action transition points, long sequence lengths, and frame contextualization, making the task well-posed for transformers. Given developments enabling transformers to scale linearly, we demonstrate through our architecture how they can be applied to improve action alignment accuracy over the equivalent RNN-based models with the attention mechanism focusing around salient action transition regions. Additionally, given the recent focus on inference-time transcript selection, we propose a supplemental transcript embedding approach to select transcripts more quickly at inference-time. Furthermore, we subsequently demonstrate how this approach can also improve the overall segmentation performance. Finally, we evaluate our proposed methods across the benchmark datasets to better understand the applicability of transformers and the importance of transcript selection on this video-driven weakly-supervised task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge