Michael M. Bronstein
Graph Neural Networks as Gradient Flows
Jun 22, 2022
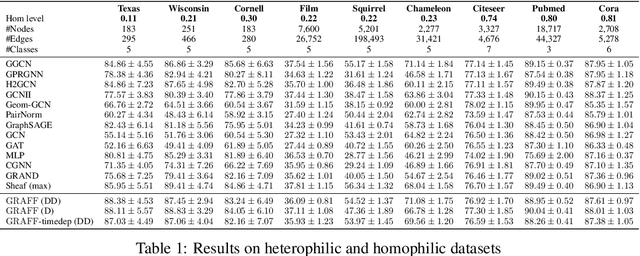
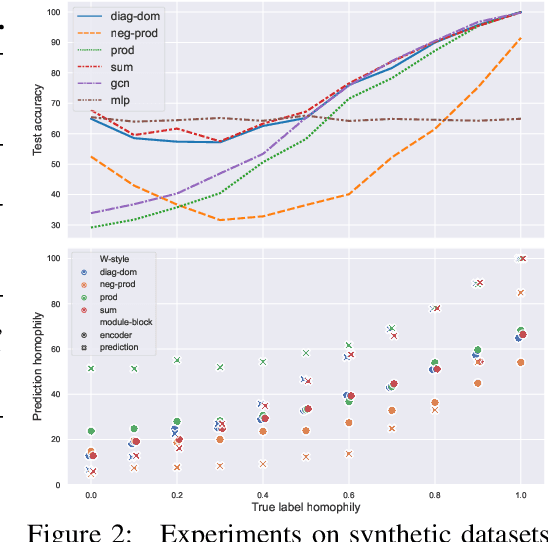
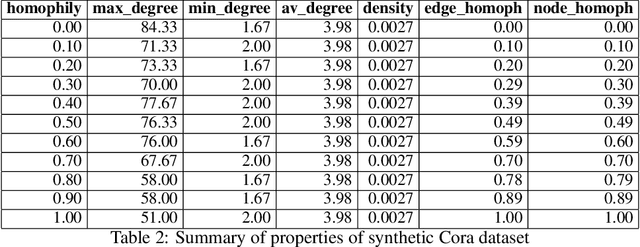
Abstract:Dynamical systems minimizing an energy are ubiquitous in geometry and physics. We propose a gradient flow framework for GNNs where the equations follow the direction of steepest descent of a learnable energy. This approach allows to explain the GNN evolution from a multi-particle perspective as learning attractive and repulsive forces in feature space via the positive and negative eigenvalues of a symmetric "channel-mixing" matrix. We perform spectral analysis of the solutions and conclude that gradient flow graph convolutional models can induce a dynamics dominated by the graph high frequencies which is desirable for heterophilic datasets. We also describe structural constraints on common GNN architectures allowing to interpret them as gradient flows. We perform thorough ablation studies corroborating our theoretical analysis and show competitive performance of simple and lightweight models on real-world homophilic and heterophilic datasets.
Learning to Infer Structures of Network Games
Jun 16, 2022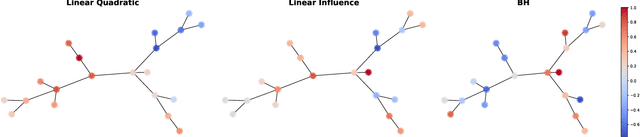
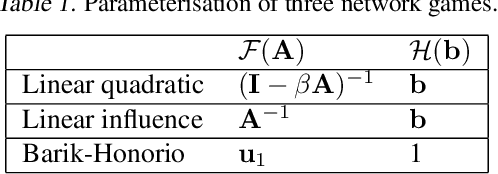

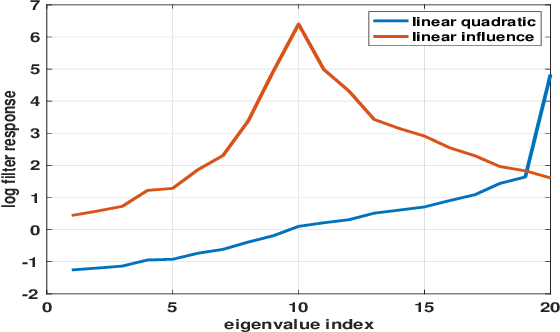
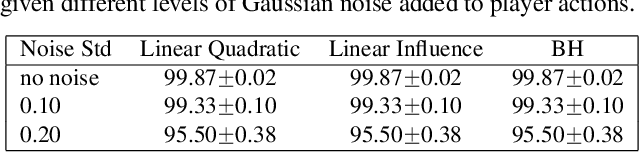
Abstract:Strategic interactions between a group of individuals or organisations can be modelled as games played on networks, where a player's payoff depends not only on their actions but also on those of their neighbours. Inferring the network structure from observed game outcomes (equilibrium actions) is an important problem with numerous potential applications in economics and social sciences. Existing methods mostly require the knowledge of the utility function associated with the game, which is often unrealistic to obtain in real-world scenarios. We adopt a transformer-like architecture which correctly accounts for the symmetries of the problem and learns a mapping from the equilibrium actions to the network structure of the game without explicit knowledge of the utility function. We test our method on three different types of network games using both synthetic and real-world data, and demonstrate its effectiveness in network structure inference and superior performance over existing methods.
Graph Anisotropic Diffusion
Apr 30, 2022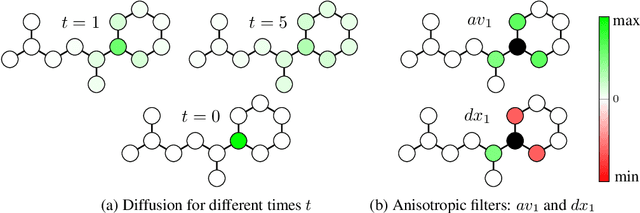
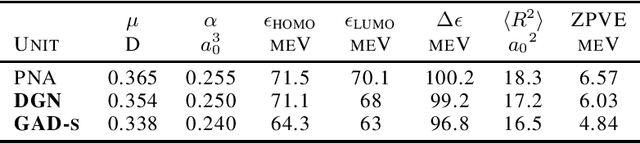
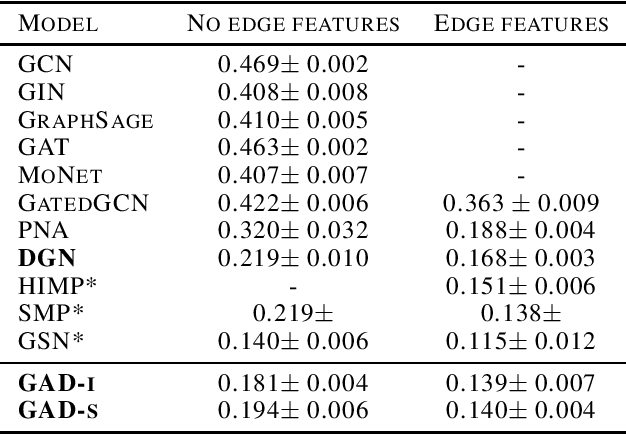
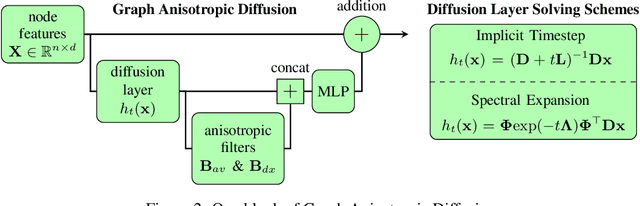

Abstract:Traditional Graph Neural Networks (GNNs) rely on message passing, which amounts to permutation-invariant local aggregation of neighbour features. Such a process is isotropic and there is no notion of `direction' on the graph. We present a new GNN architecture called Graph Anisotropic Diffusion. Our model alternates between linear diffusion, for which a closed-form solution is available, and local anisotropic filters to obtain efficient multi-hop anisotropic kernels. We test our model on two common molecular property prediction benchmarks (ZINC and QM9) and show its competitive performance.
Graph-in-Graph (GiG): Learning interpretable latent graphs in non-Euclidean domain for biological and healthcare applications
Apr 01, 2022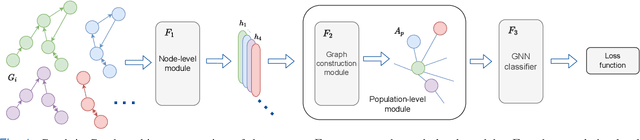
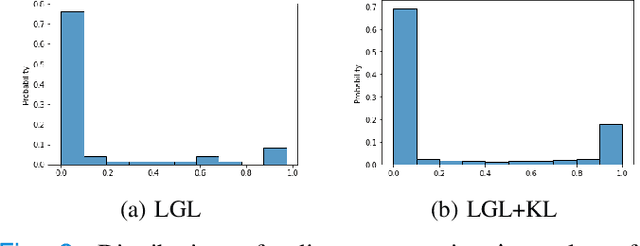
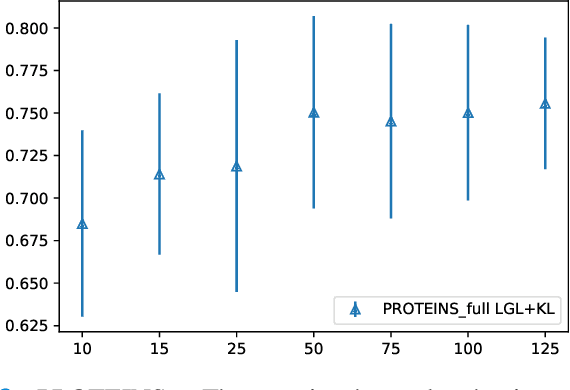
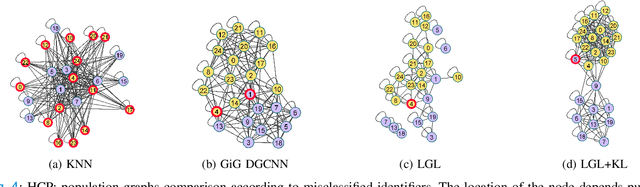
Abstract:Graphs are a powerful tool for representing and analyzing unstructured, non-Euclidean data ubiquitous in the healthcare domain. Two prominent examples are molecule property prediction and brain connectome analysis. Importantly, recent works have shown that considering relationships between input data samples have a positive regularizing effect for the downstream task in healthcare applications. These relationships are naturally modeled by a (possibly unknown) graph structure between input samples. In this work, we propose Graph-in-Graph (GiG), a neural network architecture for protein classification and brain imaging applications that exploits the graph representation of the input data samples and their latent relation. We assume an initially unknown latent-graph structure between graph-valued input data and propose to learn end-to-end a parametric model for message passing within and across input graph samples, along with the latent structure connecting the input graphs. Further, we introduce a degree distribution loss that helps regularize the predicted latent relationships structure. This regularization can significantly improve the downstream task. Moreover, the obtained latent graph can represent patient population models or networks of molecule clusters, providing a level of interpretability and knowledge discovery in the input domain of particular value in healthcare.
Neural Sheaf Diffusion: A Topological Perspective on Heterophily and Oversmoothing in GNNs
Feb 09, 2022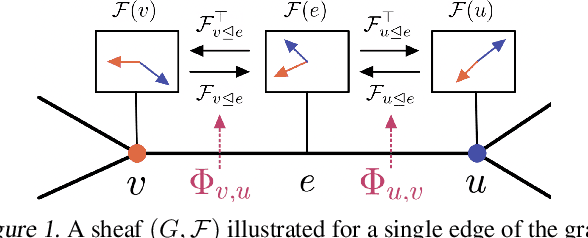
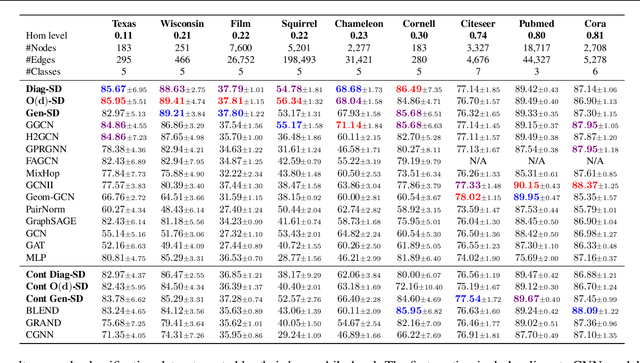
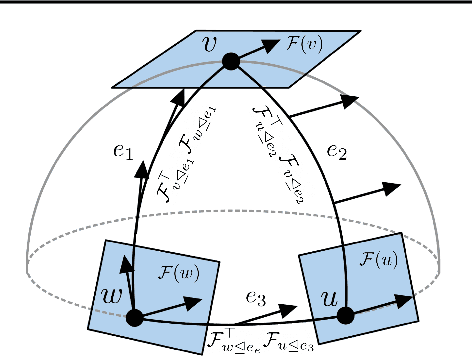
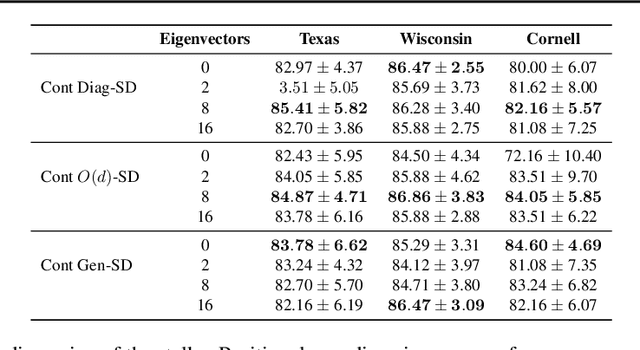
Abstract:Cellular sheaves equip graphs with "geometrical" structure by assigning vector spaces and linear maps to nodes and edges. Graph Neural Networks (GNNs) implicitly assume a graph with a trivial underlying sheaf. This choice is reflected in the structure of the graph Laplacian operator, the properties of the associated diffusion equation, and the characteristics of the convolutional models that discretise this equation. In this paper, we use cellular sheaf theory to show that the underlying geometry of the graph is deeply linked with the performance of GNNs in heterophilic settings and their oversmoothing behaviour. By considering a hierarchy of increasingly general sheaves, we study how the ability of the sheaf diffusion process to achieve linear separation of the classes in the infinite time limit expands. At the same time, we prove that when the sheaf is non-trivial, discretised parametric diffusion processes have greater control than GNNs over their asymptotic behaviour. On the practical side, we study how sheaves can be learned from data. The resulting sheaf diffusion models have many desirable properties that address the limitations of classical graph diffusion equations (and corresponding GNN models) and obtain state-of-the-art results in heterophilic settings. Overall, our work provides new connections between GNNs and algebraic topology and would be of interest to both fields.
RECOVER: sequential model optimization platform for combination drug repurposing identifies novel synergistic compounds in vitro
Feb 07, 2022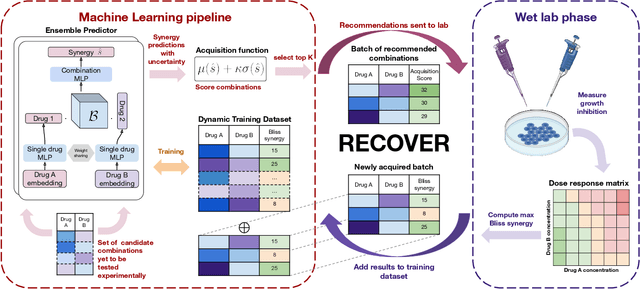
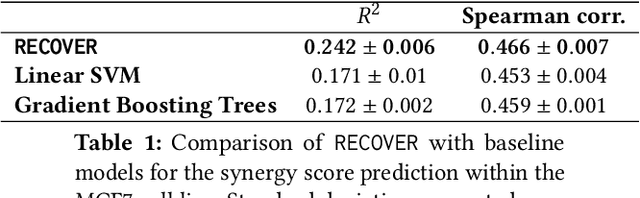

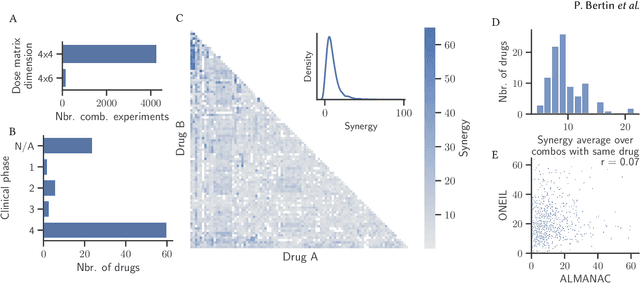
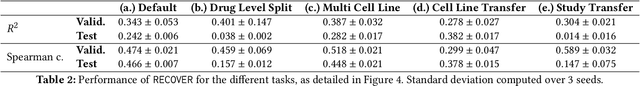
Abstract:Selecting optimal drug repurposing combinations for further preclinical development is a challenging technical feat. Due to the toxicity of many therapeutic agents (e.g., chemotherapy), practitioners have favoured selection of synergistic compounds whereby lower doses can be used whilst maintaining high efficacy. For a fixed small molecule library, an exhaustive combinatorial chemical screen becomes infeasible to perform for academic and industry laboratories alike. Deep learning models have achieved state-of-the-art results in silico for the prediction of synergy scores. However, databases of drug combinations are highly biased towards synergistic agents and these results do not necessarily generalise out of distribution. We employ a sequential model optimization search applied to a deep learning model to quickly discover highly synergistic drug combinations active against a cancer cell line, while requiring substantially less screening than an exhaustive evaluation. Through iteratively adapting the model to newly acquired data, after only 3 rounds of ML-guided experimentation (including a calibration round), we find that the set of combinations queried by our model is enriched for highly synergistic combinations. Remarkably, we rediscovered a synergistic drug combination that was later confirmed to be under study within clinical trials.
Graph-Coupled Oscillator Networks
Feb 04, 2022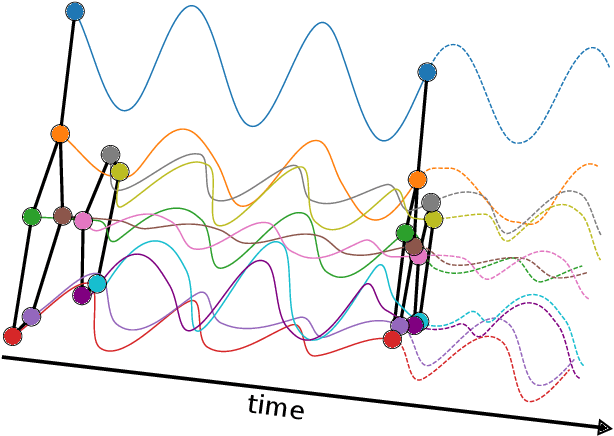
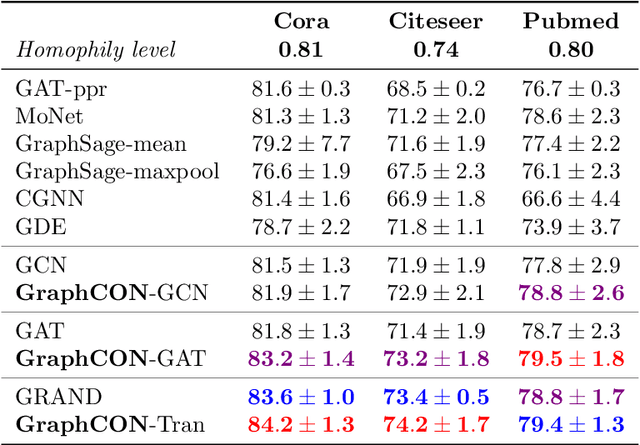
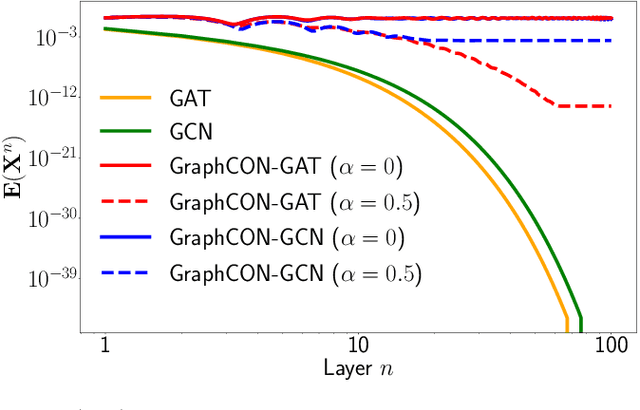
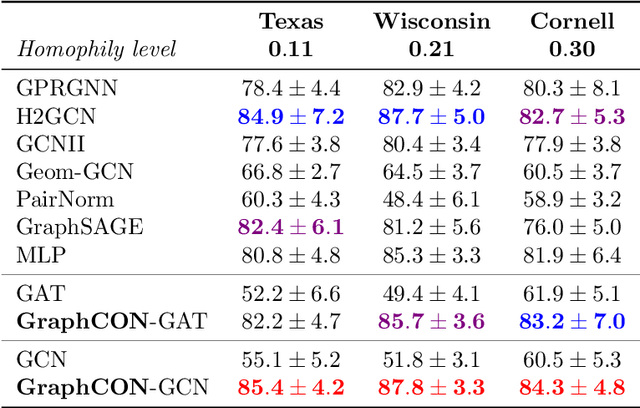
Abstract:We propose Graph-Coupled Oscillator Networks (GraphCON), a novel framework for deep learning on graphs. It is based on discretizations of a second-order system of ordinary differential equations (ODEs), which model a network of nonlinear forced and damped oscillators, coupled via the adjacency structure of the underlying graph. The flexibility of our framework permits any basic GNN layer (e.g. convolutional or attentional) as the coupling function, from which a multi-layer deep neural network is built up via the dynamics of the proposed ODEs. We relate the oversmoothing problem, commonly encountered in GNNs, to the stability of steady states of the underlying ODE and show that zero-Dirichlet energy steady states are not stable for our proposed ODEs. This demonstrates that the proposed framework mitigates the oversmoothing problem. Finally, we show that our approach offers competitive performance with respect to the state-of-the-art on a variety of graph-based learning tasks.
Understanding over-squashing and bottlenecks on graphs via curvature
Nov 29, 2021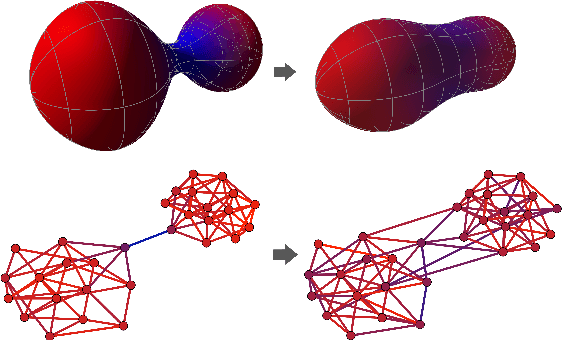
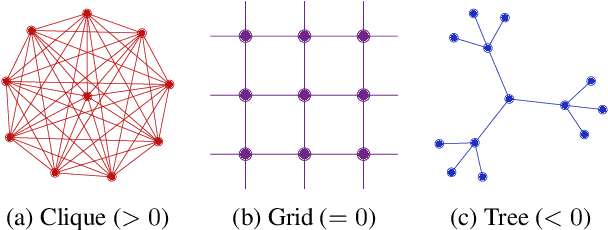
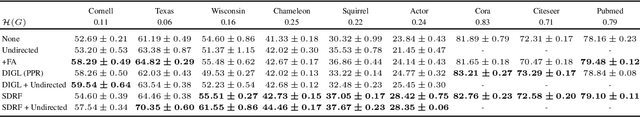
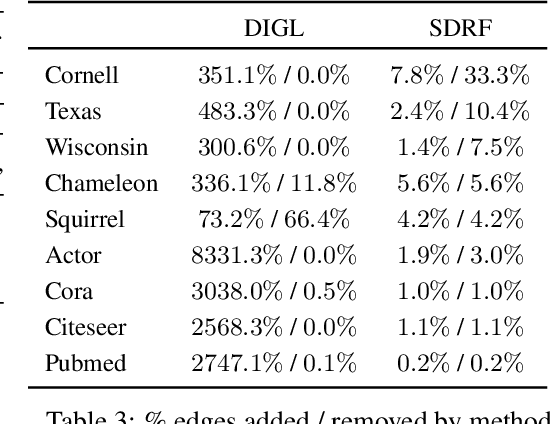
Abstract:Most graph neural networks (GNNs) use the message passing paradigm, in which node features are propagated on the input graph. Recent works pointed to the distortion of information flowing from distant nodes as a factor limiting the efficiency of message passing for tasks relying on long-distance interactions. This phenomenon, referred to as 'over-squashing', has been heuristically attributed to graph bottlenecks where the number of $k$-hop neighbors grows rapidly with $k$. We provide a precise description of the over-squashing phenomenon in GNNs and analyze how it arises from bottlenecks in the graph. For this purpose, we introduce a new edge-based combinatorial curvature and prove that negatively curved edges are responsible for the over-squashing issue. We also propose and experimentally test a curvature-based graph rewiring method to alleviate the over-squashing.
Equivariant Subgraph Aggregation Networks
Oct 06, 2021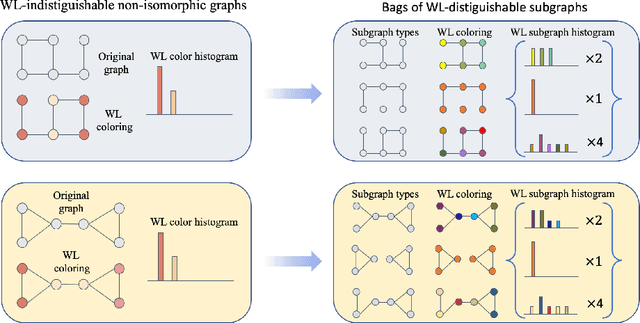
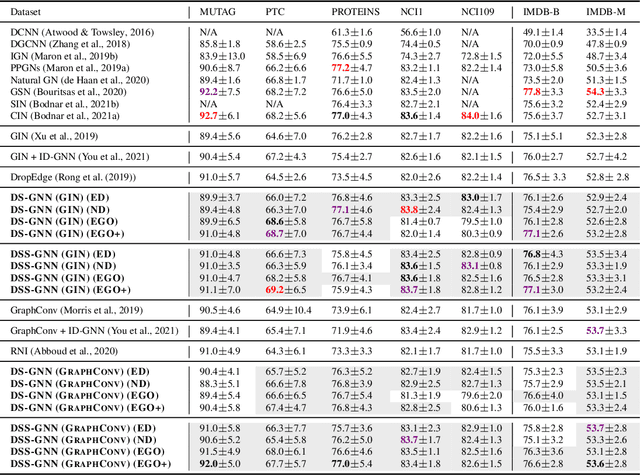
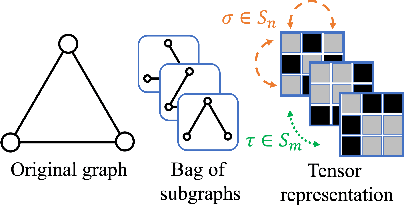
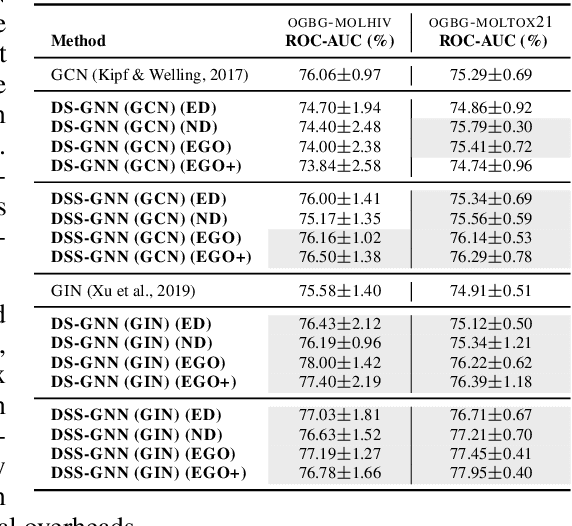
Abstract:Message-passing neural networks (MPNNs) are the leading architecture for deep learning on graph-structured data, in large part due to their simplicity and scalability. Unfortunately, it was shown that these architectures are limited in their expressive power. This paper proposes a novel framework called Equivariant Subgraph Aggregation Networks (ESAN) to address this issue. Our main observation is that while two graphs may not be distinguishable by an MPNN, they often contain distinguishable subgraphs. Thus, we propose to represent each graph as a set of subgraphs derived by some predefined policy, and to process it using a suitable equivariant architecture. We develop novel variants of the 1-dimensional Weisfeiler-Leman (1-WL) test for graph isomorphism, and prove lower bounds on the expressiveness of ESAN in terms of these new WL variants. We further prove that our approach increases the expressive power of both MPNNs and more expressive architectures. Moreover, we provide theoretical results that describe how design choices such as the subgraph selection policy and equivariant neural architecture affect our architecture's expressive power. To deal with the increased computational cost, we propose a subgraph sampling scheme, which can be viewed as a stochastic version of our framework. A comprehensive set of experiments on real and synthetic datasets demonstrates that our framework improves the expressive power and overall performance of popular GNN architectures.
Partition and Code: learning how to compress graphs
Jul 05, 2021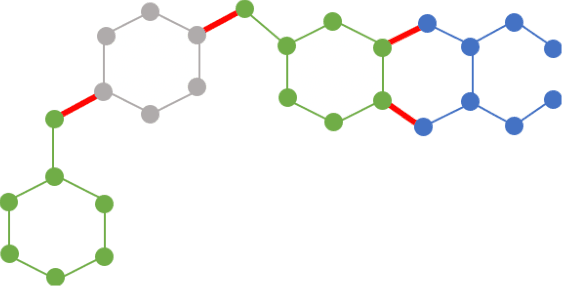
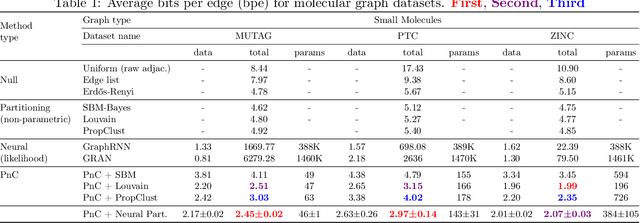
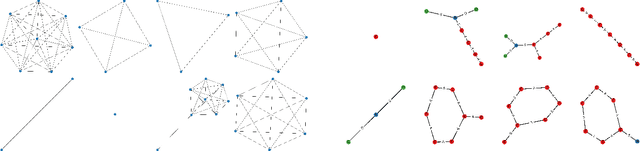
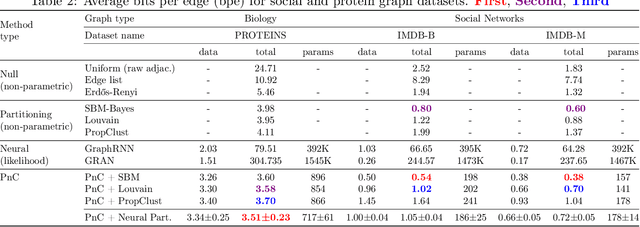
Abstract:Can we use machine learning to compress graph data? The absence of ordering in graphs poses a significant challenge to conventional compression algorithms, limiting their attainable gains as well as their ability to discover relevant patterns. On the other hand, most graph compression approaches rely on domain-dependent handcrafted representations and cannot adapt to different underlying graph distributions. This work aims to establish the necessary principles a lossless graph compression method should follow to approach the entropy storage lower bound. Instead of making rigid assumptions about the graph distribution, we formulate the compressor as a probabilistic model that can be learned from data and generalise to unseen instances. Our "Partition and Code" framework entails three steps: first, a partitioning algorithm decomposes the graph into elementary structures, then these are mapped to the elements of a small dictionary on which we learn a probability distribution, and finally, an entropy encoder translates the representation into bits. All three steps are parametric and can be trained with gradient descent. We theoretically compare the compression quality of several graph encodings and prove, under mild conditions, a total ordering of their expected description lengths. Moreover, we show that, under the same conditions, PnC achieves compression gains w.r.t. the baselines that grow either linearly or quadratically with the number of vertices. Our algorithms are quantitatively evaluated on diverse real-world networks obtaining significant performance improvements with respect to different families of non-parametric and parametric graph compressors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge