Maria Vakalopoulou
Université Paris-Saclay, CentraleSupélec, Mathématiques et Informatique pour la Complexité et les Systèmes, Inria Saclay
PVeRA: Probabilistic Vector-Based Random Matrix Adaptation
Dec 08, 2025



Abstract:Large foundation models have emerged in the last years and are pushing performance boundaries for a variety of tasks. Training or even finetuning such models demands vast datasets and computational resources, which are often scarce and costly. Adaptation methods provide a computationally efficient solution to address these limitations by allowing such models to be finetuned on small amounts of data and computing power. This is achieved by appending new trainable modules to frozen backbones with only a fraction of the trainable parameters and fitting only these modules on novel tasks. Recently, the VeRA adapter was shown to excel in parameter-efficient adaptations by utilizing a pair of frozen random low-rank matrices shared across all layers. In this paper, we propose PVeRA, a probabilistic version of the VeRA adapter, which modifies the low-rank matrices of VeRA in a probabilistic manner. This modification naturally allows handling inherent ambiguities in the input and allows for different sampling configurations during training and testing. A comprehensive evaluation was performed on the VTAB-1k benchmark and seven adapters, with PVeRA outperforming VeRA and other adapters. Our code for training models with PVeRA and benchmarking all adapters is available https://github.com/leofillioux/pvera.
Controllable Latent Space Augmentation for Digital Pathology
Aug 20, 2025Abstract:Whole slide image (WSI) analysis in digital pathology presents unique challenges due to the gigapixel resolution of WSIs and the scarcity of dense supervision signals. While Multiple Instance Learning (MIL) is a natural fit for slide-level tasks, training robust models requires large and diverse datasets. Even though image augmentation techniques could be utilized to increase data variability and reduce overfitting, implementing them effectively is not a trivial task. Traditional patch-level augmentation is prohibitively expensive due to the large number of patches extracted from each WSI, and existing feature-level augmentation methods lack control over transformation semantics. We introduce HistAug, a fast and efficient generative model for controllable augmentations in the latent space for digital pathology. By conditioning on explicit patch-level transformations (e.g., hue, erosion), HistAug generates realistic augmented embeddings while preserving initial semantic information. Our method allows the processing of a large number of patches in a single forward pass efficiently, while at the same time consistently improving MIL model performance. Experiments across multiple slide-level tasks and diverse organs show that HistAug outperforms existing methods, particularly in low-data regimes. Ablation studies confirm the benefits of learned transformations over noise-based perturbations and highlight the importance of uniform WSI-wise augmentation. Code is available at https://github.com/MICS-Lab/HistAug.
On the Risk of Misleading Reports: Diagnosing Textual Biases in Multimodal Clinical AI
Jul 31, 2025



Abstract:Clinical decision-making relies on the integrated analysis of medical images and the associated clinical reports. While Vision-Language Models (VLMs) can offer a unified framework for such tasks, they can exhibit strong biases toward one modality, frequently overlooking critical visual cues in favor of textual information. In this work, we introduce Selective Modality Shifting (SMS), a perturbation-based approach to quantify a model's reliance on each modality in binary classification tasks. By systematically swapping images or text between samples with opposing labels, we expose modality-specific biases. We assess six open-source VLMs-four generalist models and two fine-tuned for medical data-on two medical imaging datasets with distinct modalities: MIMIC-CXR (chest X-ray) and FairVLMed (scanning laser ophthalmoscopy). By assessing model performance and the calibration of every model in both unperturbed and perturbed settings, we reveal a marked dependency on text input, which persists despite the presence of complementary visual information. We also perform a qualitative attention-based analysis which further confirms that image content is often overshadowed by text details. Our findings highlight the importance of designing and evaluating multimodal medical models that genuinely integrate visual and textual cues, rather than relying on single-modality signals.
THUNDER: Tile-level Histopathology image UNDERstanding benchmark
Jul 10, 2025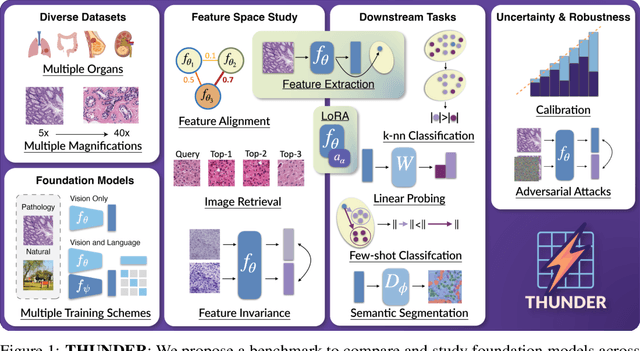
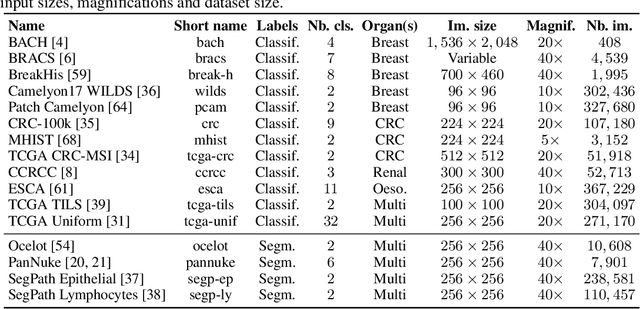
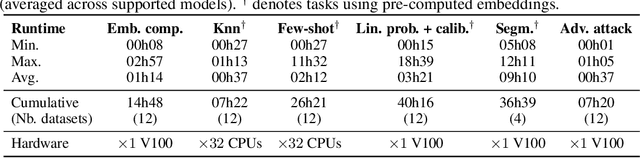
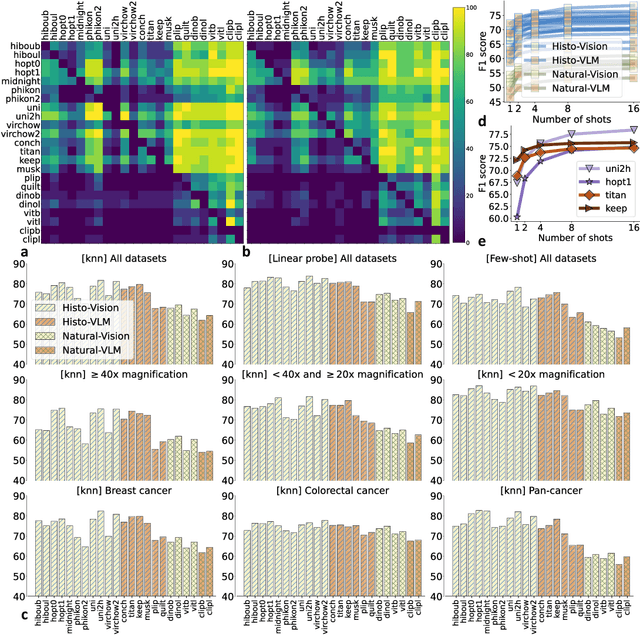
Abstract:Progress in a research field can be hard to assess, in particular when many concurrent methods are proposed in a short period of time. This is the case in digital pathology, where many foundation models have been released recently to serve as feature extractors for tile-level images, being used in a variety of downstream tasks, both for tile- and slide-level problems. Benchmarking available methods then becomes paramount to get a clearer view of the research landscape. In particular, in critical domains such as healthcare, a benchmark should not only focus on evaluating downstream performance, but also provide insights about the main differences between methods, and importantly, further consider uncertainty and robustness to ensure a reliable usage of proposed models. For these reasons, we introduce THUNDER, a tile-level benchmark for digital pathology foundation models, allowing for efficient comparison of many models on diverse datasets with a series of downstream tasks, studying their feature spaces and assessing the robustness and uncertainty of predictions informed by their embeddings. THUNDER is a fast, easy-to-use, dynamic benchmark that can already support a large variety of state-of-the-art foundation, as well as local user-defined models for direct tile-based comparison. In this paper, we provide a comprehensive comparison of 23 foundation models on 16 different datasets covering diverse tasks, feature analysis, and robustness. The code for THUNDER is publicly available at https://github.com/MICS-Lab/thunder.
Full Conformal Adaptation of Medical Vision-Language Models
Jun 06, 2025



Abstract:Vision-language models (VLMs) pre-trained at large scale have shown unprecedented transferability capabilities and are being progressively integrated into medical image analysis. Although its discriminative potential has been widely explored, its reliability aspect remains overlooked. This work investigates their behavior under the increasingly popular split conformal prediction (SCP) framework, which theoretically guarantees a given error level on output sets by leveraging a labeled calibration set. However, the zero-shot performance of VLMs is inherently limited, and common practice involves few-shot transfer learning pipelines, which cannot absorb the rigid exchangeability assumptions of SCP. To alleviate this issue, we propose full conformal adaptation, a novel setting for jointly adapting and conformalizing pre-trained foundation models, which operates transductively over each test data point using a few-shot adaptation set. Moreover, we complement this framework with SS-Text, a novel training-free linear probe solver for VLMs that alleviates the computational cost of such a transductive approach. We provide comprehensive experiments using 3 different modality-specialized medical VLMs and 9 adaptation tasks. Our framework requires exactly the same data as SCP, and provides consistent relative improvements of up to 27% on set efficiency while maintaining the same coverage guarantees.
BayesAdapter: enhanced uncertainty estimation in CLIP few-shot adaptation
Dec 12, 2024



Abstract:The emergence of large pre-trained vision-language models (VLMs) represents a paradigm shift in machine learning, with unprecedented results in a broad span of visual recognition tasks. CLIP, one of the most popular VLMs, has exhibited remarkable zero-shot and transfer learning capabilities in classification. To transfer CLIP to downstream tasks, adapters constitute a parameter-efficient approach that avoids backpropagation through the large model (unlike related prompt learning methods). However, CLIP adapters have been developed to target discriminative performance, and the quality of their uncertainty estimates has been overlooked. In this work we show that the discriminative performance of state-of-the-art CLIP adapters does not always correlate with their uncertainty estimation capabilities, which are essential for a safe deployment in real-world scenarios. We also demonstrate that one of such adapters is obtained through MAP inference from a more general probabilistic framework. Based on this observation we introduce BayesAdapter, which leverages Bayesian inference to estimate a full probability distribution instead of a single point, better capturing the variability inherent in the parameter space. In a comprehensive empirical evaluation we show that our approach obtains high quality uncertainty estimates in the predictions, standing out in calibration and selective classification. Our code is publicly available at: https://github.com/pablomorales92/BayesAdapter.
Are foundation models for computer vision good conformal predictors?
Dec 08, 2024



Abstract:Recent advances in self-supervision and constrastive learning have brought the performance of foundation models to unprecedented levels in a variety of tasks. Fueled by this progress, these models are becoming the prevailing approach for a wide array of real-world vision problems, including risk-sensitive and high-stakes applications. However, ensuring safe deployment in these scenarios requires a more comprehensive understanding of their uncertainty modeling capabilities, which has been barely explored. In this work, we delve into the behavior of vision and vision-language foundation models under Conformal Prediction (CP), a statistical framework that provides theoretical guarantees of marginal coverage of the true class. Across extensive experiments including popular vision classification benchmarks, well-known foundation vision models, and three CP methods, our findings reveal that foundation models are well-suited for conformalization procedures, particularly those integrating Vision Transformers. Furthermore, we show that calibrating the confidence predictions of these models leads to efficiency degradation of the conformal set on adaptive CP methods. In contrast, few-shot adaptation to downstream tasks generally enhances conformal scores, where we identify Adapters as a better conformable alternative compared to Prompt Learning strategies. Our empirical study identifies APS as particularly promising in the context of vision foundation models, as it does not violate the marginal coverage property across multiple challenging, yet realistic scenarios.
ADAPT: Multimodal Learning for Detecting Physiological Changes under Missing Modalities
Jul 04, 2024



Abstract:Multimodality has recently gained attention in the medical domain, where imaging or video modalities may be integrated with biomedical signals or health records. Yet, two challenges remain: balancing the contributions of modalities, especially in cases with a limited amount of data available, and tackling missing modalities. To address both issues, in this paper, we introduce the AnchoreD multimodAl Physiological Transformer (ADAPT), a multimodal, scalable framework with two key components: (i) aligning all modalities in the space of the strongest, richest modality (called anchor) to learn a joint embedding space, and (ii) a Masked Multimodal Transformer, leveraging both inter- and intra-modality correlations while handling missing modalities. We focus on detecting physiological changes in two real-life scenarios: stress in individuals induced by specific triggers and fighter pilots' loss of consciousness induced by $g$-forces. We validate the generalizability of ADAPT through extensive experiments on two datasets for these tasks, where we set the new state of the art while demonstrating its robustness across various modality scenarios and its high potential for real-life applications.
You Don't Need Data-Augmentation in Self-Supervised Learning
Jun 13, 2024



Abstract:Self-Supervised learning (SSL) with Joint-Embedding Architectures (JEA) has led to outstanding performances. All instantiations of this paradigm were trained using strong and well-established hand-crafted data augmentations, leading to the general belief that they are required for the proper training and performance of such models. On the other hand, generative reconstruction-based models such as BEIT and MAE or Joint-Embedding Predictive Architectures such as I-JEPA have shown strong performance without using data augmentations except masking. In this work, we challenge the importance of invariance and data-augmentation in JEAs at scale. By running a case-study on a recent SSL foundation model - DINOv2 - we show that strong image representations can be obtained with JEAs and only cropping without resizing provided the training data is large enough, reaching state-of-the-art results and using the least amount of augmentation in the literature. Through this study, we also discuss the impact of compute constraints on the outcomes of experimental deep learning research, showing that they can lead to very different conclusions.
Advancing human-centric AI for robust X-ray analysis through holistic self-supervised learning
May 02, 2024



Abstract:AI Foundation models are gaining traction in various applications, including medical fields like radiology. However, medical foundation models are often tested on limited tasks, leaving their generalisability and biases unexplored. We present RayDINO, a large visual encoder trained by self-supervision on 873k chest X-rays. We compare RayDINO to previous state-of-the-art models across nine radiology tasks, from classification and dense segmentation to text generation, and provide an in depth analysis of population, age and sex biases of our model. Our findings suggest that self-supervision allows patient-centric AI proving useful in clinical workflows and interpreting X-rays holistically. With RayDINO and small task-specific adapters, we reach state-of-the-art results and improve generalization to unseen populations while mitigating bias, illustrating the true promise of foundation models: versatility and robustness.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge