Jingyi Ren
Effective and Transparent RAG: Adaptive-Reward Reinforcement Learning for Decision Traceability
May 19, 2025Abstract:Retrieval-Augmented Generation (RAG) has significantly improved the performance of large language models (LLMs) on knowledge-intensive domains. However, although RAG achieved successes across distinct domains, there are still some unsolved challenges: 1) Effectiveness. Existing research mainly focuses on developing more powerful RAG retrievers, but how to enhance the generator's (LLM's) ability to utilize the retrieved information for reasoning and generation? 2) Transparency. Most RAG methods ignore which retrieved content actually contributes to the reasoning process, resulting in a lack of interpretability and visibility. To address this, we propose ARENA (Adaptive-Rewarded Evidence Navigation Agent), a transparent RAG generator framework trained via reinforcement learning (RL) with our proposed rewards. Based on the structured generation and adaptive reward calculation, our RL-based training enables the model to identify key evidence, perform structured reasoning, and generate answers with interpretable decision traces. Applied to Qwen2.5-7B-Instruct and Llama3.1-8B-Instruct, abundant experiments with various RAG baselines demonstrate that our model achieves 10-30% improvements on all multi-hop QA datasets, which is comparable with the SOTA Commercially-developed LLMs (e.g., OpenAI-o1, DeepSeek-R1). Further analyses show that ARENA has strong flexibility to be adopted on new datasets without extra training. Our models and codes are publicly released.
CodeRepoQA: A Large-scale Benchmark for Software Engineering Question Answering
Dec 19, 2024



Abstract:In this work, we introduce CodeRepoQA, a large-scale benchmark specifically designed for evaluating repository-level question-answering capabilities in the field of software engineering. CodeRepoQA encompasses five programming languages and covers a wide range of scenarios, enabling comprehensive evaluation of language models. To construct this dataset, we crawl data from 30 well-known repositories in GitHub, the largest platform for hosting and collaborating on code, and carefully filter raw data. In total, CodeRepoQA is a multi-turn question-answering benchmark with 585,687 entries, covering a diverse array of software engineering scenarios, with an average of 6.62 dialogue turns per entry. We evaluate ten popular large language models on our dataset and provide in-depth analysis. We find that LLMs still have limitations in question-answering capabilities in the field of software engineering, and medium-length contexts are more conducive to LLMs' performance. The entire benchmark is publicly available at https://github.com/kinesiatricssxilm14/CodeRepoQA.
DialogAgent: An Auto-engagement Agent for Code Question Answering Data Production
Dec 11, 2024



Abstract:Large Language Models (LLMs) have become increasingly integral to enhancing developer productivity, particularly in code generation, comprehension, and repair tasks. However, fine-tuning these models with high-quality, real-world data is challenging due to privacy concerns and the lack of accessible, labeled datasets. In this paper, we present DialogAgent, an automated tool for generating synthetic training data that closely mimics real developer interactions within Integrated Development Environments (IDEs). DialogAgent enables the production of diverse, high-fidelity query-response pairs by simulating multi-turn dialogues and contextual behaviors observed in real-world programming scenarios. The tool significantly reduces the reliance on manual data generation, increasing efficiency by 4.8 times compared to traditional methods. Our experiments and online deployment demonstrate substantial improvements in model performance for code-related question-answering tasks: the acceptance rate of responses generated by our in-house model is improved by 33%, after training on synthesized data generated by DialogAgent.
AcademicGPT: Empowering Academic Research
Nov 21, 2023



Abstract:Large Language Models (LLMs) have demonstrated exceptional capabilities across various natural language processing tasks. Yet, many of these advanced LLMs are tailored for broad, general-purpose applications. In this technical report, we introduce AcademicGPT, designed specifically to empower academic research. AcademicGPT is a continual training model derived from LLaMA2-70B. Our training corpus mainly consists of academic papers, thesis, content from some academic domain, high-quality Chinese data and others. While it may not be extensive in data scale, AcademicGPT marks our initial venture into a domain-specific GPT tailored for research area. We evaluate AcademicGPT on several established public benchmarks such as MMLU and CEval, as well as on some specialized academic benchmarks like PubMedQA, SCIEval, and our newly-created ComputerScienceQA, to demonstrate its ability from general knowledge ability, to Chinese ability, and to academic ability. Building upon AcademicGPT's foundation model, we also developed several applications catered to the academic area, including General Academic Question Answering, AI-assisted Paper Reading, Paper Review, and AI-assisted Title and Abstract Generation.
BIOS: An Algorithmically Generated Biomedical Knowledge Graph
Mar 18, 2022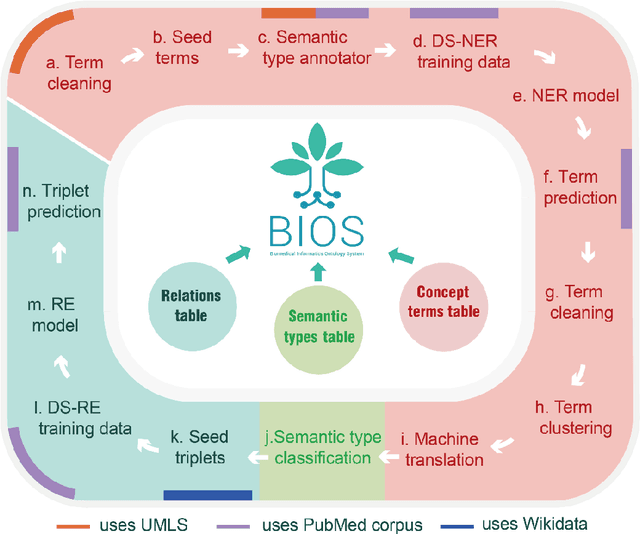
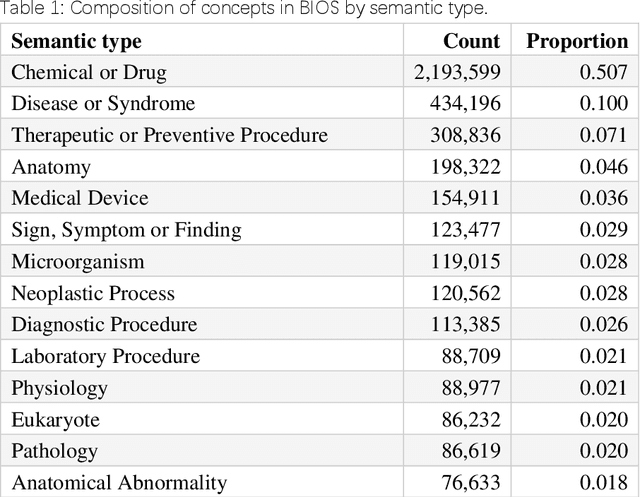
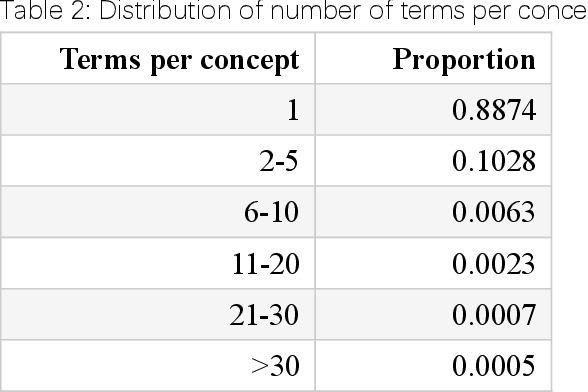

Abstract:Biomedical knowledge graphs (BioMedKGs) are essential infrastructures for biomedical and healthcare big data and artificial intelligence (AI), facilitating natural language processing, model development, and data exchange. For many decades, these knowledge graphs have been built via expert curation, which can no longer catch up with the speed of today's AI development, and a transition to algorithmically generated BioMedKGs is necessary. In this work, we introduce the Biomedical Informatics Ontology System (BIOS), the first large scale publicly available BioMedKG that is fully generated by machine learning algorithms. BIOS currently contains 4.1 million concepts, 7.4 million terms in two languages, and 7.3 million relation triplets. We introduce the methodology for developing BIOS, which covers curation of raw biomedical terms, computationally identifying synonymous terms and aggregating them to create concept nodes, semantic type classification of the concepts, relation identification, and biomedical machine translation. We provide statistics about the current content of BIOS and perform preliminary assessment for term quality, synonym grouping, and relation extraction. Results suggest that machine learning-based BioMedKG development is a totally viable solution for replacing traditional expert curation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge