Donglin Xie
GenomeQA: Benchmarking General Large Language Models for Genome Sequence Understanding
Apr 07, 2026Abstract:Large Language Models (LLMs) are increasingly adopted as conversational assistants in genomics, where they are mainly used to reason over biological knowledge, annotations, and analysis outputs through natural language interfaces. However, existing benchmarks either focus on specialized DNA models trained for sequence prediction or evaluate biological knowledge using text-only questions, leaving the behavior of general-purpose LLMs when directly exposed to raw genome sequences underexplored. We introduce GenomeQA, a benchmark designed to provide a controlled evaluation setting for general-purpose LLMs on sequence-based genome inference tasks. GenomeQA comprises 5,200 samples drawn from multiple biological databases, with sequence lengths ranging from 6 to 1,000 base pairs (bp), spanning six task families: Enhancer and Promoter Identification, Splice Site Identification, Taxonomic Classification, Histone Mark Prediction, Transcription Factor Binding Site Prediction, and TF Motif Prediction. Across six frontier LLMs, we find that models consistently outperform random baselines and can exploit local sequence signals such as GC content and short motifs, while performance degrades on tasks that require more indirect or multi-step inference over sequence patterns. GenomeQA establishes a diagnostic benchmark for studying and improving the use of general-purpose LLMs on raw genomic sequences.
mtslearn: Machine Learning in Python for Medical Time Series
Mar 31, 2026Abstract:Medical time-series data captures the dynamic progression of patient conditions, playing a vital role in modern clinical decision support systems. However, real-world clinical data is highly heterogeneous and inconsistently formatted. Furthermore, existing machine learning tools often have steep learning curves and fragmented workflows. Consequently, a significant gap remains between cutting-edge AI technologies and clinical application. To address this, we introduce mtslearn, an end-to-end integrated toolkit specifically designed for medical time-series data. First, the framework provides a unified data interface that automates the parsing and alignment of wide, long, and flat data formats. This design significantly reduces data cleaning overhead. Building on this, mtslearn provides a complete pipeline from data reading and feature engineering to model training and result visualization. Furthermore, it offers flexible interfaces for custom algorithms. Through a modular design, mtslearn simplifies complex data engineering tasks into a few lines of code. This significantly lowers the barrier to entry for clinicians with limited programming experience, empowering them to focus more on exploring medical hypotheses and accelerating the translation of advanced algorithms into real-world clinical practice. mtslearn is publicly available at https://github.com/PKUDigitalHealth/mtslearn.
Holter-to-Sleep: AI-Enabled Repurposing of Single-Lead ECG for Sleep Phenotyping
Mar 19, 2026Abstract:Sleep disturbances are tightly linked to cardiovascular risk, yet polysomnography (PSG)-the clinical reference standard-remains resource-intensive and poorly suited for multi-night, home-based, and large-scale screening. Single-lead electrocardiography (ECG), already ubiquitous in Holter and patch-based devices, enables comfortable long-term acquisition and encodes sleep-relevant physiology through autonomic modulation and cardiorespiratory coupling. Here, we present a proof-of-concept Holter-to-Sleep framework that, using single-lead ECG as the sole input, jointly supports overnight sleep phenotyping and Holter-grade cardiac phenotyping within the same recording, and further provides an explicit analytic pathway for scalable cardio-sleep association studies. The framework is developed and validated on a pooled multi-center PSG sample of 10,439 studies spanning four public cohorts, with independent external evaluation to assess cross-cohort generalizability, and additional real-world feasibility assessment using overnight patch-ECG recordings via objective-subjective consistency analysis. This integrated design enables robust extraction of clinically meaningful overnight sleep phenotypes under heterogeneous populations and acquisition conditions, and facilitates systematic linkage between ECG-derived sleep metrics and arrhythmia-related Holter phenotypes. Collectively, the Holter-to-Sleep paradigm offers a practical foundation for low-burden, home-deployable, and scalable cardio-sleep monitoring and research beyond traditional PSG-centric workflows.
Artificial intelligence-enabled single-lead ECG for non-invasive hyperkalemia detection: development, multicenter validation, and proof-of-concept deployment
Mar 17, 2026Abstract:Hyperkalemia is a life-threatening electrolyte disorder that is common in patients with chronic kidney disease and heart failure, yet frequent monitoring remains difficult outside hospital settings. We developed and validated Pocket-K, a single-lead AI-ECG system initialized from the ECGFounder foundation model for non-invasive hyperkalemia screening and handheld deployment. In this multicentre observational study using routinely collected clinical ECG and laboratory data, 34,439 patients contributed 62,290 ECG--potassium pairs. Lead I data were used to fine-tune the model. Data from Peking University People's Hospital were divided into development and temporal validation sets, and data from The Second Hospital of Tianjin Medical University served as an independent external validation set. Hyperkalemia was defined as venous serum potassium > 5.5 mmol/L. Pocket-K achieved AUROCs of 0.936 in internal testing, 0.858 in temporal validation, and 0.808 in external validation. For KDIGO-defined moderate-to-severe hyperkalemia (serum potassium >= 6.0 mmol/L), AUROCs increased to 0.940 and 0.861 in the temporal and external sets, respectively. External negative predictive value exceeded 99.3%. Model-predicted high risk below the hyperkalemia threshold was more common in patients with chronic kidney disease and heart failure. A handheld prototype enabled near-real-time inference, supporting future prospective evaluation in native handheld and wearable settings.
sleep2vec: Unified Cross-Modal Alignment for Heterogeneous Nocturnal Biosignals
Feb 14, 2026Abstract:Tasks ranging from sleep staging to clinical diagnosis traditionally rely on standard polysomnography (PSG) devices, bedside monitors and wearable devices, which capture diverse nocturnal biosignals (e.g., EEG, EOG, ECG, SpO$_2$). However, heterogeneity across devices and frequent sensor dropout pose significant challenges for unified modelling of these multimodal signals. We present \texttt{sleep2vec}, a foundation model for diverse and incomplete nocturnal biosignals that learns a shared representation via cross-modal alignment. \texttt{sleep2vec} is contrastively pre-trained on 42,249 overnight recordings spanning nine modalities using a \textit{Demography, Age, Site \& History-aware InfoNCE} objective that incorporates physiological and acquisition metadata (\textit{e.g.}, age, gender, recording site) to dynamically weight negatives and mitigate cohort-specific shortcuts. On downstream sleep staging and clinical outcome assessment, \texttt{sleep2vec} consistently outperforms strong baselines and remains robust to any subset of available modalities and sensor dropout. We further characterize, to our knowledge for the first time, scaling laws for nocturnal biosignals with respect to modality diversity and model capacity. Together, these results show that unified cross-modal alignment, coupled with principled scaling, enables label-efficient, general-purpose modelling of real-world nocturnal biosignals.
Simultaneous Polysomnography and Cardiotocography Reveal Temporal Correlation Between Maternal Obstructive Sleep Apnea and Fetal Hypoxia
Apr 17, 2025Abstract:Background: Obstructive sleep apnea syndrome (OSAS) during pregnancy is common and can negatively affect fetal outcomes. However, studies on the immediate effects of maternal hypoxia on fetal heart rate (FHR) changes are lacking. Methods: We used time-synchronized polysomnography (PSG) and cardiotocography (CTG) data from two cohorts to analyze the correlation between maternal hypoxia and FHR changes (accelerations or decelerations). Maternal hypoxic event characteristics were analyzed using generalized linear modeling (GLM) to assess their associations with different FHR changes. Results: A total of 118 pregnant women participated. FHR changes were significantly associated with maternal hypoxia, primarily characterized by accelerations. A longer hypoxic duration correlated with more significant FHR accelerations (P < 0.05), while prolonged hypoxia and greater SpO2 drop were linked to FHR decelerations (P < 0.05). Both cohorts showed a transient increase in FHR during maternal hypoxia, which returned to baseline after the event resolved. Conclusion: Maternal hypoxia significantly affects FHR, suggesting that maternal OSAS may contribute to fetal hypoxia. These findings highlight the importance of maternal-fetal interactions and provide insights for future interventions.
Federated Selective Aggregation for Knowledge Amalgamation
Jul 27, 2022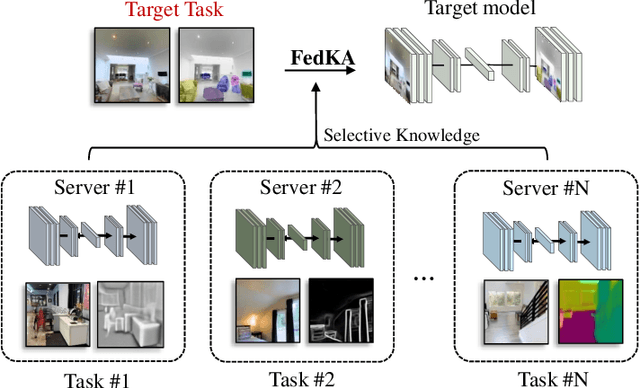
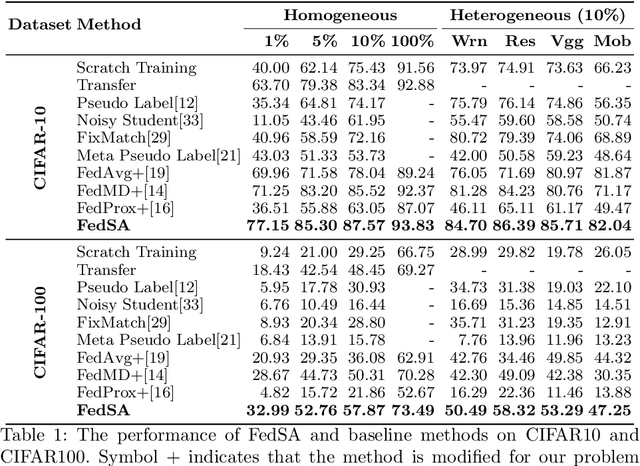
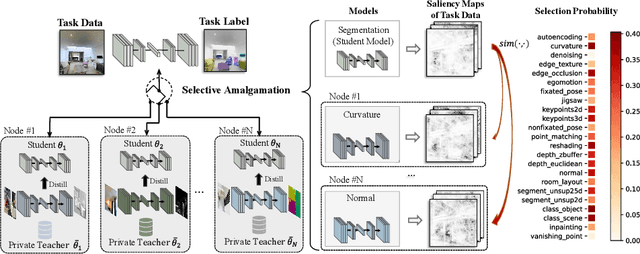
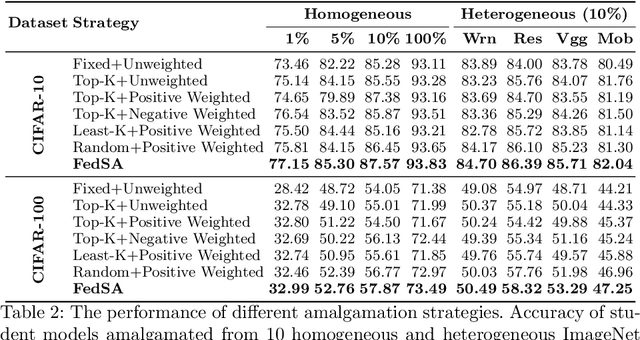
Abstract:In this paper, we explore a new knowledge-amalgamation problem, termed Federated Selective Aggregation (FedSA). The goal of FedSA is to train a student model for a new task with the help of several decentralized teachers, whose pre-training tasks and data are different and agnostic. Our motivation for investigating such a problem setup stems from a recent dilemma of model sharing. Many researchers or institutes have spent enormous resources on training large and competent networks. Due to the privacy, security, or intellectual property issues, they are, however, not able to share their own pre-trained models, even if they wish to contribute to the community. The proposed FedSA offers a solution to this dilemma and makes it one step further since, again, the learned student may specialize in a new task different from all of the teachers. To this end, we proposed a dedicated strategy for handling FedSA. Specifically, our student-training process is driven by a novel saliency-based approach that adaptively selects teachers as the participants and integrates their representative capabilities into the student. To evaluate the effectiveness of FedSA, we conduct experiments on both single-task and multi-task settings. Experimental results demonstrate that FedSA effectively amalgamates knowledge from decentralized models and achieves competitive performance to centralized baselines.
CADG: A Model Based on Cross Attention for Domain Generalization
Apr 07, 2022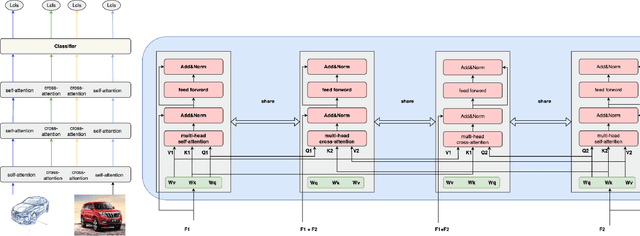
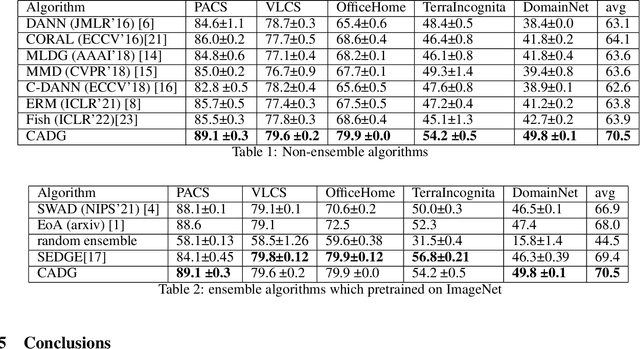
Abstract:In Domain Generalization (DG) tasks, models are trained by using only training data from the source domains to achieve generalization on an unseen target domain, this will suffer from the distribution shift problem. So it's important to learn a classifier to focus on the common representation which can be used to classify on multi-domains, so that this classifier can achieve a high performance on an unseen target domain as well. With the success of cross attention in various cross-modal tasks, we find that cross attention is a powerful mechanism to align the features come from different distributions. So we design a model named CADG (cross attention for domain generalization), wherein cross attention plays a important role, to address distribution shift problem. Such design makes the classifier can be adopted on multi-domains, so the classifier will generalize well on an unseen domain. Experiments show that our proposed method achieves state-of-the-art performance on a variety of domain generalization benchmarks compared with other single model and can even achieve a better performance than some ensemble-based methods.
Mosaicking to Distill: Knowledge Distillation from Out-of-Domain Data
Oct 27, 2021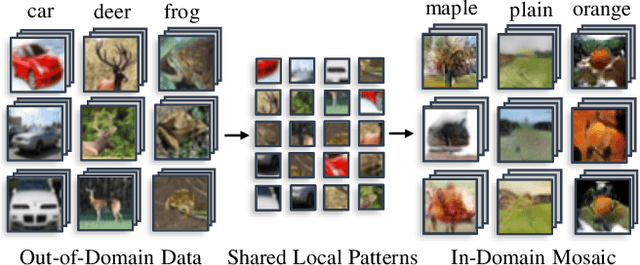
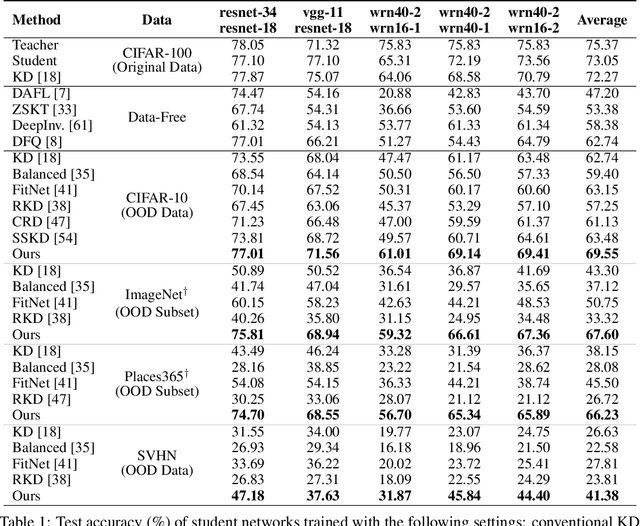
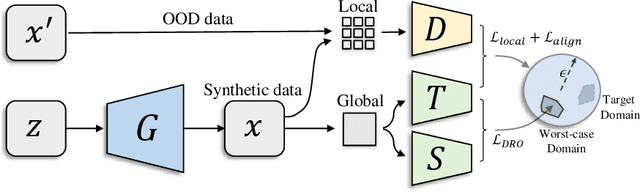
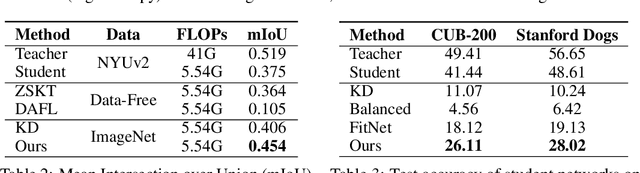
Abstract:Knowledge distillation~(KD) aims to craft a compact student model that imitates the behavior of a pre-trained teacher in a target domain. Prior KD approaches, despite their gratifying results, have largely relied on the premise that \emph{in-domain} data is available to carry out the knowledge transfer. Such an assumption, unfortunately, in many cases violates the practical setting, since the original training data or even the data domain is often unreachable due to privacy or copyright reasons. In this paper, we attempt to tackle an ambitious task, termed as \emph{out-of-domain} knowledge distillation~(OOD-KD), which allows us to conduct KD using only OOD data that can be readily obtained at a very low cost. Admittedly, OOD-KD is by nature a highly challenging task due to the agnostic domain gap. To this end, we introduce a handy yet surprisingly efficacious approach, dubbed as~\textit{MosaicKD}. The key insight behind MosaicKD lies in that, samples from various domains share common local patterns, even though their global semantic may vary significantly; these shared local patterns, in turn, can be re-assembled analogous to mosaic tiling, to approximate the in-domain data and to further alleviating the domain discrepancy. In MosaicKD, this is achieved through a four-player min-max game, in which a generator, a discriminator, a student network, are collectively trained in an adversarial manner, partially under the guidance of a pre-trained teacher. We validate MosaicKD over {classification and semantic segmentation tasks} across various benchmarks, and demonstrate that it yields results much superior to the state-of-the-art counterparts on OOD data. Our code is available at \url{https://github.com/zju-vipa/MosaicKD}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge