Dong Wei
RECIST Weakly Supervised Lesion Segmentation via Label-Space Co-Training
Mar 01, 2023



Abstract:As an essential indicator for cancer progression and treatment response, tumor size is often measured following the response evaluation criteria in solid tumors (RECIST) guideline in CT slices. By marking each lesion with its longest axis and the longest perpendicular one, laborious pixel-wise manual annotation can be avoided. However, such a coarse substitute cannot provide a rich and accurate base to allow versatile quantitative analysis of lesions. To this end, we propose a novel weakly supervised framework to exploit the existing rich RECIST annotations for pixel-wise lesion segmentation. Specifically, a pair of under- and over-segmenting masks are constructed for each lesion based on its RECIST annotation and served as the label for co-training a pair of subnets, respectively, along with the proposed label-space perturbation induced consistency loss to bridge the gap between the two subnets and enable effective co-training. Extensive experiments are conducted on a public dataset to demonstrate the superiority of the proposed framework regarding the RECIST-based weakly supervised segmentation task and its universal applicability to various backbone networks.
MADAv2: Advanced Multi-Anchor Based Active Domain Adaptation Segmentation
Jan 18, 2023Abstract:Unsupervised domain adaption has been widely adopted in tasks with scarce annotated data. Unfortunately, mapping the target-domain distribution to the source-domain unconditionally may distort the essential structural information of the target-domain data, leading to inferior performance. To address this issue, we firstly propose to introduce active sample selection to assist domain adaptation regarding the semantic segmentation task. By innovatively adopting multiple anchors instead of a single centroid, both source and target domains can be better characterized as multimodal distributions, in which way more complementary and informative samples are selected from the target domain. With only a little workload to manually annotate these active samples, the distortion of the target-domain distribution can be effectively alleviated, achieving a large performance gain. In addition, a powerful semi-supervised domain adaptation strategy is proposed to alleviate the long-tail distribution problem and further improve the segmentation performance. Extensive experiments are conducted on public datasets, and the results demonstrate that the proposed approach outperforms state-of-the-art methods by large margins and achieves similar performance to the fully-supervised upperbound, i.e., 71.4% mIoU on GTA5 and 71.8% mIoU on SYNTHIA. The effectiveness of each component is also verified by thorough ablation studies.
Lesion Guided Explainable Few Weak-shot Medical Report Generation
Nov 17, 2022Abstract:Medical images are widely used in clinical practice for diagnosis. Automatically generating interpretable medical reports can reduce radiologists' burden and facilitate timely care. However, most existing approaches to automatic report generation require sufficient labeled data for training. In addition, the learned model can only generate reports for the training classes, lacking the ability to adapt to previously unseen novel diseases. To this end, we propose a lesion guided explainable few weak-shot medical report generation framework that learns correlation between seen and novel classes through visual and semantic feature alignment, aiming to generate medical reports for diseases not observed in training. It integrates a lesion-centric feature extractor and a Transformer-based report generation module. Concretely, the lesion-centric feature extractor detects the abnormal regions and learns correlations between seen and novel classes with multi-view (visual and lexical) embeddings. Then, features of the detected regions and corresponding embeddings are concatenated as multi-view input to the report generation module for explainable report generation, including text descriptions and corresponding abnormal regions detected in the images. We conduct experiments on FFA-IR, a dataset providing explainable annotations, showing that our framework outperforms others on report generation for novel diseases.
Human Joint Kinematics Diffusion-Refinement for Stochastic Motion Prediction
Oct 12, 2022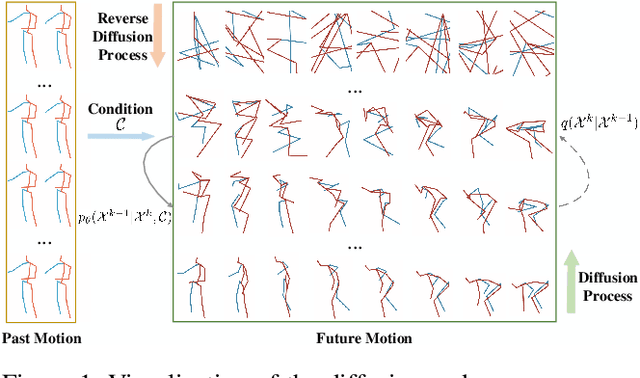
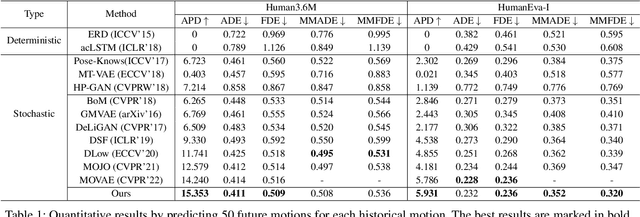
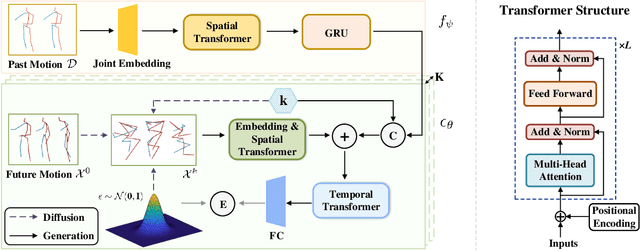
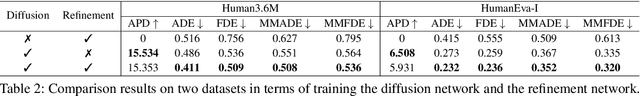
Abstract:Stochastic human motion prediction aims to forecast multiple plausible future motions given a single pose sequence from the past. Most previous works focus on designing elaborate losses to improve the accuracy, while the diversity is typically characterized by randomly sampling a set of latent variables from the latent prior, which is then decoded into possible motions. This joint training of sampling and decoding, however, suffers from posterior collapse as the learned latent variables tend to be ignored by a strong decoder, leading to limited diversity. Alternatively, inspired by the diffusion process in nonequilibrium thermodynamics, we propose MotionDiff, a diffusion probabilistic model to treat the kinematics of human joints as heated particles, which will diffuse from original states to a noise distribution. This process offers a natural way to obtain the "whitened" latents without any trainable parameters, and human motion prediction can be regarded as the reverse diffusion process that converts the noise distribution into realistic future motions conditioned on the observed sequence. Specifically, MotionDiff consists of two parts: a spatial-temporal transformer-based diffusion network to generate diverse yet plausible motions, and a graph convolutional network to further refine the outputs. Experimental results on two datasets demonstrate that our model yields the competitive performance in terms of both accuracy and diversity.
A Benchmark for Weakly Semi-Supervised Abnormality Localization in Chest X-Rays
Sep 05, 2022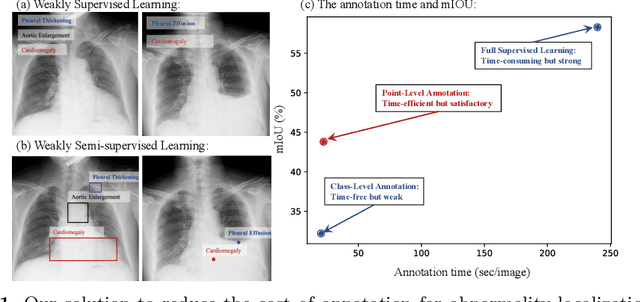
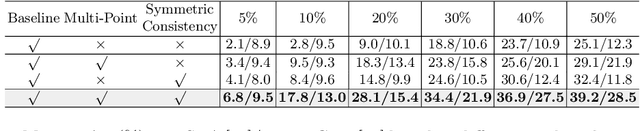
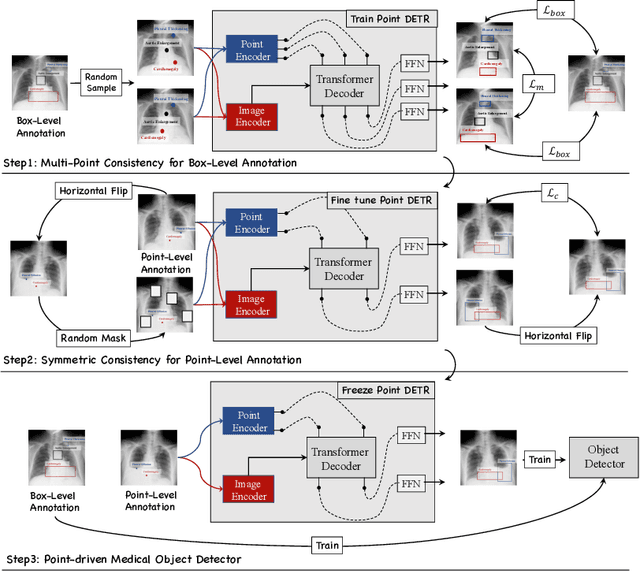
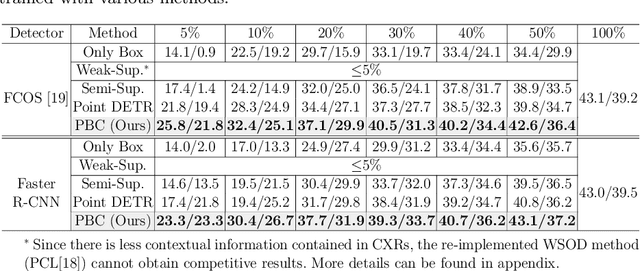
Abstract:Accurate abnormality localization in chest X-rays (CXR) can benefit the clinical diagnosis of various thoracic diseases. However, the lesion-level annotation can only be performed by experienced radiologists, and it is tedious and time-consuming, thus difficult to acquire. Such a situation results in a difficulty to develop a fully-supervised abnormality localization system for CXR. In this regard, we propose to train the CXR abnormality localization framework via a weakly semi-supervised strategy, termed Point Beyond Class (PBC), which utilizes a small number of fully annotated CXRs with lesion-level bounding boxes and extensive weakly annotated samples by points. Such a point annotation setting can provide weakly instance-level information for abnormality localization with a marginal annotation cost. Particularly, the core idea behind our PBC is to learn a robust and accurate mapping from the point annotations to the bounding boxes against the variance of annotated points. To achieve that, a regularization term, namely multi-point consistency, is proposed, which drives the model to generate the consistent bounding box from different point annotations inside the same abnormality. Furthermore, a self-supervision, termed symmetric consistency, is also proposed to deeply exploit the useful information from the weakly annotated data for abnormality localization. Experimental results on RSNA and VinDr-CXR datasets justify the effectiveness of the proposed method. When less than 20% box-level labels are used for training, an improvement of ~5 in mAP can be achieved by our PBC, compared to the current state-of-the-art method (i.e., Point DETR). Code is available at https://github.com/HaozheLiu-ST/Point-Beyond-Class.
Dense Cross-Query-and-Support Attention Weighted Mask Aggregation for Few-Shot Segmentation
Jul 18, 2022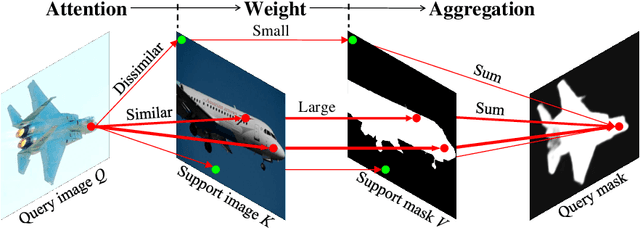
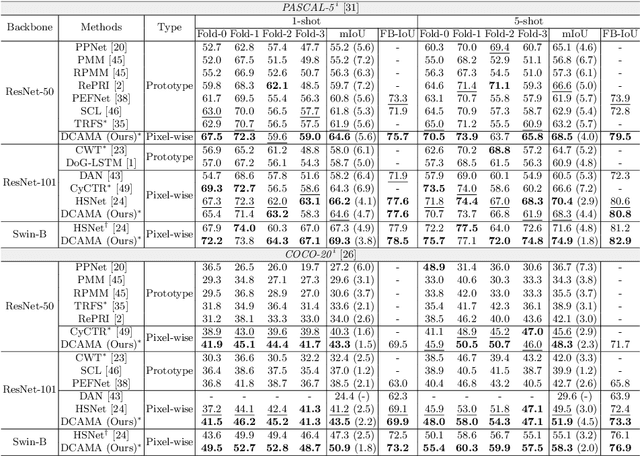
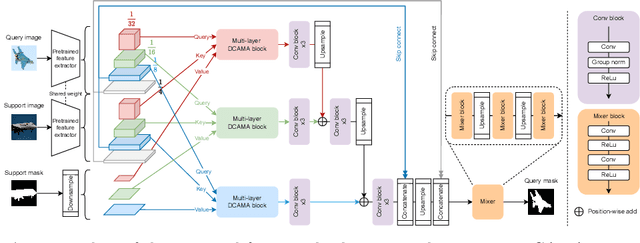
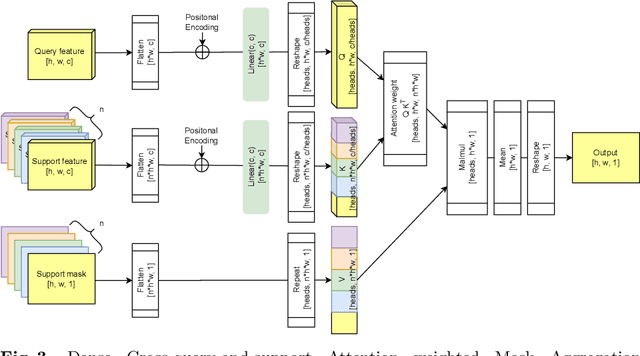
Abstract:Research into Few-shot Semantic Segmentation (FSS) has attracted great attention, with the goal to segment target objects in a query image given only a few annotated support images of the target class. A key to this challenging task is to fully utilize the information in the support images by exploiting fine-grained correlations between the query and support images. However, most existing approaches either compressed the support information into a few class-wise prototypes, or used partial support information (e.g., only foreground) at the pixel level, causing non-negligible information loss. In this paper, we propose Dense pixel-wise Cross-query-and-support Attention weighted Mask Aggregation (DCAMA), where both foreground and background support information are fully exploited via multi-level pixel-wise correlations between paired query and support features. Implemented with the scaled dot-product attention in the Transformer architecture, DCAMA treats every query pixel as a token, computes its similarities with all support pixels, and predicts its segmentation label as an additive aggregation of all the support pixels' labels -- weighted by the similarities. Based on the unique formulation of DCAMA, we further propose efficient and effective one-pass inference for n-shot segmentation, where pixels of all support images are collected for the mask aggregation at once. Experiments show that our DCAMA significantly advances the state of the art on standard FSS benchmarks of PASCAL-5i, COCO-20i, and FSS-1000, e.g., with 3.1%, 9.7%, and 3.6% absolute improvements in 1-shot mIoU over previous best records. Ablative studies also verify the design DCAMA.
Deformer: Towards Displacement Field Learning for Unsupervised Medical Image Registration
Jul 07, 2022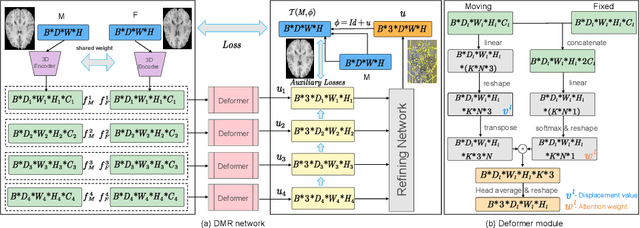
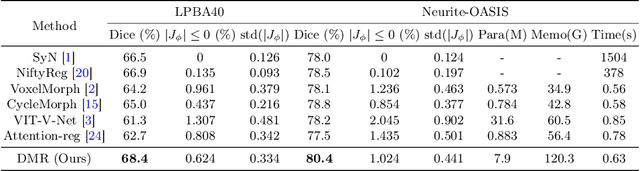
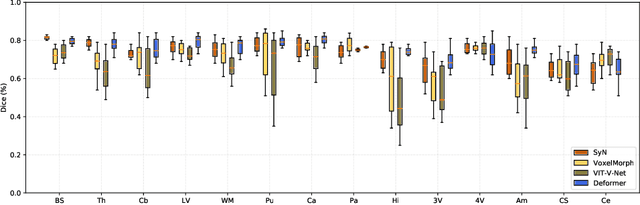
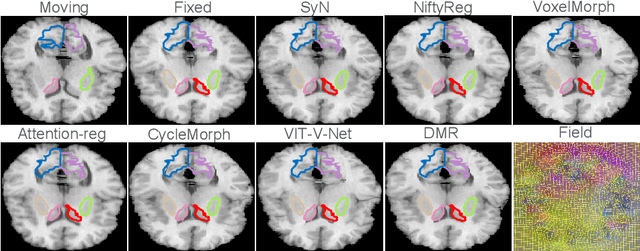
Abstract:Recently, deep-learning-based approaches have been widely studied for deformable image registration task. However, most efforts directly map the composite image representation to spatial transformation through the convolutional neural network, ignoring its limited ability to capture spatial correspondence. On the other hand, Transformer can better characterize the spatial relationship with attention mechanism, its long-range dependency may be harmful to the registration task, where voxels with too large distances are unlikely to be corresponding pairs. In this study, we propose a novel Deformer module along with a multi-scale framework for the deformable image registration task. The Deformer module is designed to facilitate the mapping from image representation to spatial transformation by formulating the displacement vector prediction as the weighted summation of several bases. With the multi-scale framework to predict the displacement fields in a coarse-to-fine manner, superior performance can be achieved compared with traditional and learning-based approaches. Comprehensive experiments on two public datasets are conducted to demonstrate the effectiveness of the proposed Deformer module as well as the multi-scale framework.
mmFormer: Multimodal Medical Transformer for Incomplete Multimodal Learning of Brain Tumor Segmentation
Jun 06, 2022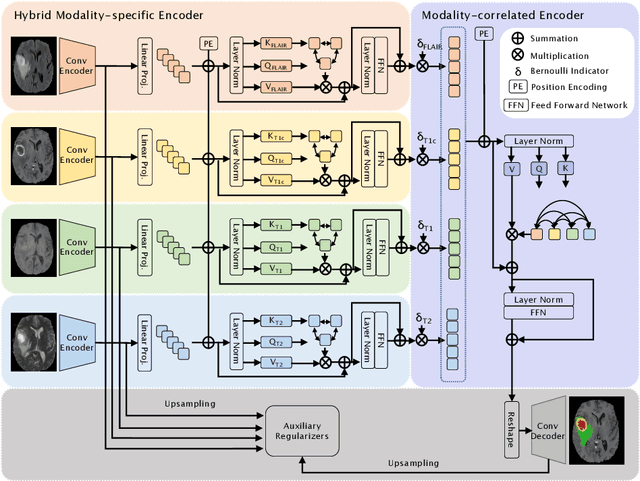
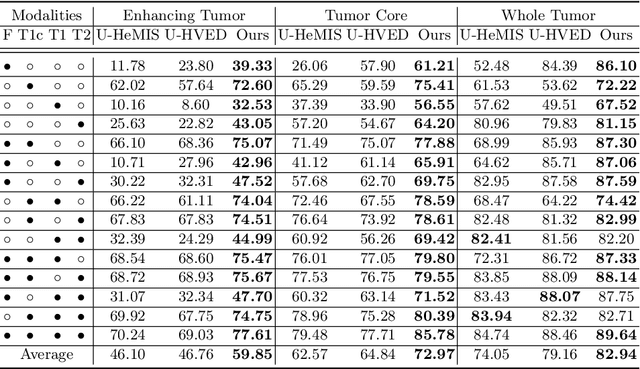

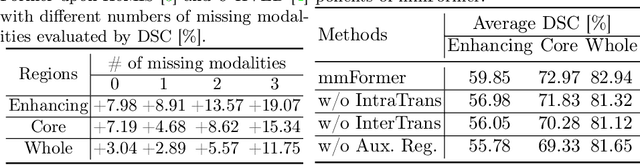
Abstract:Accurate brain tumor segmentation from Magnetic Resonance Imaging (MRI) is desirable to joint learning of multimodal images. However, in clinical practice, it is not always possible to acquire a complete set of MRIs, and the problem of missing modalities causes severe performance degradation in existing multimodal segmentation methods. In this work, we present the first attempt to exploit the Transformer for multimodal brain tumor segmentation that is robust to any combinatorial subset of available modalities. Concretely, we propose a novel multimodal Medical Transformer (mmFormer) for incomplete multimodal learning with three main components: the hybrid modality-specific encoders that bridge a convolutional encoder and an intra-modal Transformer for both local and global context modeling within each modality; an inter-modal Transformer to build and align the long-range correlations across modalities for modality-invariant features with global semantics corresponding to tumor region; a decoder that performs a progressive up-sampling and fusion with the modality-invariant features to generate robust segmentation. Besides, auxiliary regularizers are introduced in both encoder and decoder to further enhance the model's robustness to incomplete modalities. We conduct extensive experiments on the public BraTS $2018$ dataset for brain tumor segmentation. The results demonstrate that the proposed mmFormer outperforms the state-of-the-art methods for incomplete multimodal brain tumor segmentation on almost all subsets of incomplete modalities, especially by an average 19.07% improvement of Dice on tumor segmentation with only one available modality. The code is available at https://github.com/YaoZhang93/mmFormer.
Myocardial Segmentation of Late Gadolinium Enhanced MR Images by Propagation of Contours from Cine MR Images
May 21, 2022Abstract:Automatic segmentation of myocardium in Late Gadolinium Enhanced (LGE) Cardiac MR (CMR) images is often difficult due to the intensity heterogeneity resulting from accumulation of contrast agent in infarcted areas. In this paper, we propose an automatic segmentation framework that fully utilizes shared information between corresponding cine and LGE images of a same patient. Given myocardial contours in cine CMR images, the proposed framework achieves accurate segmentation of LGE CMR images in a coarse-to-fine manner. Affine registration is first performed between the corresponding cine and LGE image pair, followed by nonrigid registration, and finally local deformation of myocardial contours driven by forces derived from local features of the LGE image. Experimental results on real patient data with expert outlined ground truth show that the proposed framework can generate accurate and reliable results for myocardial segmentation of LGE CMR images.
A Comprehensive 3-D Framework for Automatic Quantification of Late Gadolinium Enhanced Cardiac Magnetic Resonance Images
May 21, 2022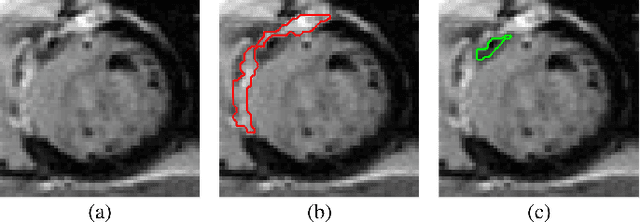
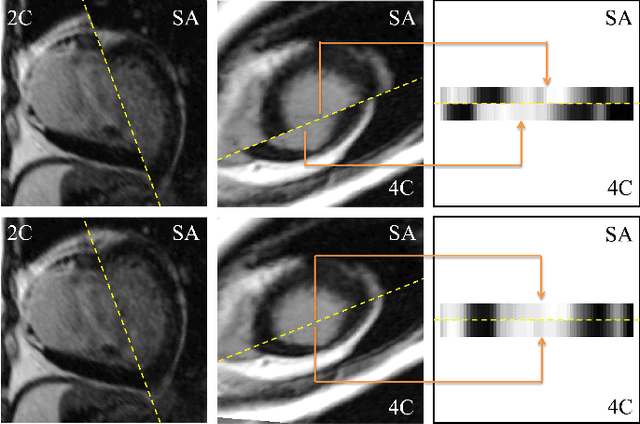
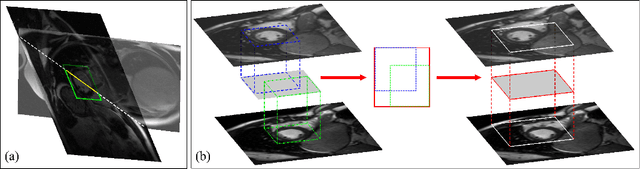
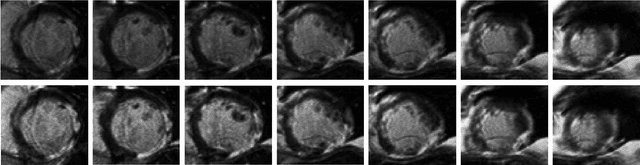
Abstract:Late gadolinium enhanced (LGE) cardiac magnetic resonance (CMR) can directly visualize nonviable myocardium with hyperenhanced intensities with respect to normal myocardium. For heart attack patients, it is crucial to facilitate the decision of appropriate therapy by analyzing and quantifying their LGE CMR images. To achieve accurate quantification, LGE CMR images need to be processed in two steps: segmentation of the myocardium followed by classification of infarcts within the segmented myocardium. However, automatic segmentation is difficult usually due to the intensity heterogeneity of the myocardium and intensity similarity between the infarcts and blood pool. Besides, the slices of an LGE CMR dataset often suffer from spatial and intensity distortions, causing further difficulties in segmentation and classification. In this paper, we present a comprehensive 3-D framework for automatic quantification of LGE CMR images. In this framework, myocardium is segmented with a novel method that deforms coupled endocardial and epicardial meshes and combines information in both short- and long-axis slices, while infarcts are classified with a graph-cut algorithm incorporating intensity and spatial information. Moreover, both spatial and intensity distortions are effectively corrected with specially designed countermeasures. Experiments with 20 sets of real patient data show visually good segmentation and classification results that are quantitatively in strong agreement with those manually obtained by experts.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge