Da Zheng
DistTGL: Distributed Memory-Based Temporal Graph Neural Network Training
Jul 14, 2023



Abstract:Memory-based Temporal Graph Neural Networks are powerful tools in dynamic graph representation learning and have demonstrated superior performance in many real-world applications. However, their node memory favors smaller batch sizes to capture more dependencies in graph events and needs to be maintained synchronously across all trainers. As a result, existing frameworks suffer from accuracy loss when scaling to multiple GPUs. Evenworse, the tremendous overhead to synchronize the node memory make it impractical to be deployed to distributed GPU clusters. In this work, we propose DistTGL -- an efficient and scalable solution to train memory-based TGNNs on distributed GPU clusters. DistTGL has three improvements over existing solutions: an enhanced TGNN model, a novel training algorithm, and an optimized system. In experiments, DistTGL achieves near-linear convergence speedup, outperforming state-of-the-art single-machine method by 14.5% in accuracy and 10.17x in training throughput.
Graph-Aware Language Model Pre-Training on a Large Graph Corpus Can Help Multiple Graph Applications
Jun 05, 2023



Abstract:Model pre-training on large text corpora has been demonstrated effective for various downstream applications in the NLP domain. In the graph mining domain, a similar analogy can be drawn for pre-training graph models on large graphs in the hope of benefiting downstream graph applications, which has also been explored by several recent studies. However, no existing study has ever investigated the pre-training of text plus graph models on large heterogeneous graphs with abundant textual information (a.k.a. large graph corpora) and then fine-tuning the model on different related downstream applications with different graph schemas. To address this problem, we propose a framework of graph-aware language model pre-training (GALM) on a large graph corpus, which incorporates large language models and graph neural networks, and a variety of fine-tuning methods on downstream applications. We conduct extensive experiments on Amazon's real internal datasets and large public datasets. Comprehensive empirical results and in-depth analysis demonstrate the effectiveness of our proposed methods along with lessons learned.
Train Your Own GNN Teacher: Graph-Aware Distillation on Textual Graphs
Apr 20, 2023



Abstract:How can we learn effective node representations on textual graphs? Graph Neural Networks (GNNs) that use Language Models (LMs) to encode textual information of graphs achieve state-of-the-art performance in many node classification tasks. Yet, combining GNNs with LMs has not been widely explored for practical deployments due to its scalability issues. In this work, we tackle this challenge by developing a Graph-Aware Distillation framework (GRAD) to encode graph structures into an LM for graph-free, fast inference. Different from conventional knowledge distillation, GRAD jointly optimizes a GNN teacher and a graph-free student over the graph's nodes via a shared LM. This encourages the graph-free student to exploit graph information encoded by the GNN teacher while at the same time, enables the GNN teacher to better leverage textual information from unlabeled nodes. As a result, the teacher and the student models learn from each other to improve their overall performance. Experiments in eight node classification benchmarks in both transductive and inductive settings showcase GRAD's superiority over existing distillation approaches for textual graphs.
PaGE-Link: Path-based Graph Neural Network Explanation for Heterogeneous Link Prediction
Mar 21, 2023Abstract:Transparency and accountability have become major concerns for black-box machine learning (ML) models. Proper explanations for the model behavior increase model transparency and help researchers develop more accountable models. Graph neural networks (GNN) have recently shown superior performance in many graph ML problems than traditional methods, and explaining them has attracted increased interest. However, GNN explanation for link prediction (LP) is lacking in the literature. LP is an essential GNN task and corresponds to web applications like recommendation and sponsored search on web. Given existing GNN explanation methods only address node/graph-level tasks, we propose Path-based GNN Explanation for heterogeneous Link prediction (PaGE-Link) that generates explanations with connection interpretability, enjoys model scalability, and handles graph heterogeneity. Qualitatively, PaGE-Link can generate explanations as paths connecting a node pair, which naturally captures connections between the two nodes and easily transfer to human-interpretable explanations. Quantitatively, explanations generated by PaGE-Link improve AUC for recommendation on citation and user-item graphs by 9 - 35% and are chosen as better by 78.79% of responses in human evaluation.
OrthoReg: Improving Graph-regularized MLPs via Orthogonality Regularization
Jan 31, 2023



Abstract:Graph Neural Networks (GNNs) are currently dominating in modeling graph-structure data, while their high reliance on graph structure for inference significantly impedes them from widespread applications. By contrast, Graph-regularized MLPs (GR-MLPs) implicitly inject the graph structure information into model weights, while their performance can hardly match that of GNNs in most tasks. This motivates us to study the causes of the limited performance of GR-MLPs. In this paper, we first demonstrate that node embeddings learned from conventional GR-MLPs suffer from dimensional collapse, a phenomenon in which the largest a few eigenvalues dominate the embedding space, through empirical observations and theoretical analysis. As a result, the expressive power of the learned node representations is constrained. We further propose OrthoReg, a novel GR-MLP model to mitigate the dimensional collapse issue. Through a soft regularization loss on the correlation matrix of node embeddings, OrthoReg explicitly encourages orthogonal node representations and thus can naturally avoid dimensionally collapsed representations. Experiments on traditional transductive semi-supervised classification tasks and inductive node classification for cold-start scenarios demonstrate its effectiveness and superiority.
PIGEON: Optimizing CUDA Code Generator for End-to-End Training and Inference of Relational Graph Neural Networks
Jan 16, 2023Abstract:Relational graph neural networks (RGNNs) are graph neural networks (GNNs) with dedicated structures for modeling the different types of nodes and/or edges in heterogeneous graphs. While RGNNs have been increasingly adopted in many real-world applications due to their versatility and accuracy, they pose performance and system design challenges due to their inherent computation patterns, gap between the programming interface and kernel APIs, and heavy programming efforts in optimizing kernels caused by their coupling with data layout and heterogeneity. To systematically address these challenges, we propose Pigeon, a novel two-level intermediate representation (IR) and its code generator framework, that (a) represents the key properties of the RGNN models to bridge the gap between the programming interface and kernel APIs, (b) decouples model semantics, data layout, and operators-specific optimization from each other to reduce programming efforts, (c) expresses and leverages optimization opportunities in inter-operator transforms, data layout, and operator-specific schedules. By building on one general matrix multiply (GEMM) template and a node/edge traversal template, Pigeon achieves up to 7.8x speed-up in inference and 5.6x speed-up in training compared with the state-of-the-art public systems in select models, i.e., RGCN, RGAT, HGT, when running heterogeneous graphs provided by Deep Graph Library (DGL) and Open Graph Benchmark (OGB). Pigeon also triggers fewer out-of-memory (OOM) errors. In addition, we propose linear operator fusion and compact materialization to further accelerate the system by up to 2.2x.
From Local to Global: Spectral-Inspired Graph Neural Networks
Sep 24, 2022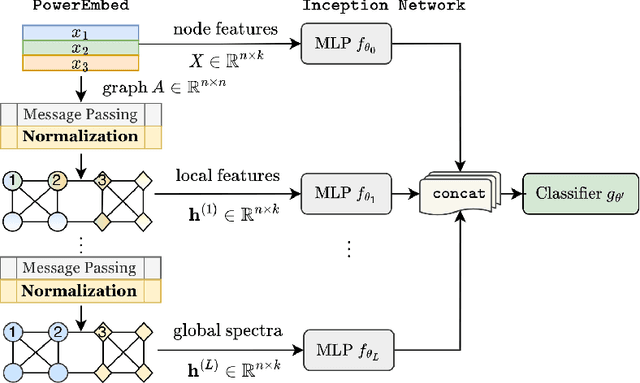
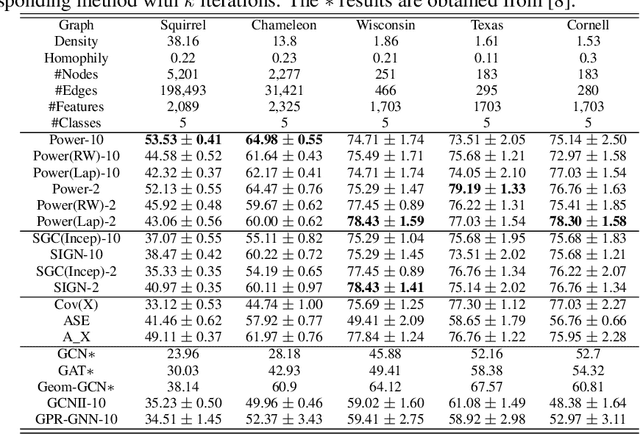
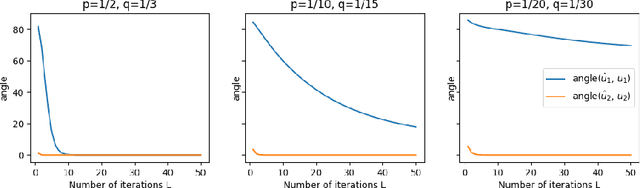
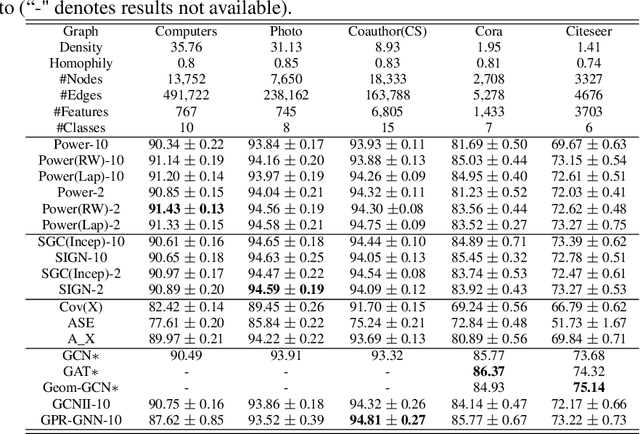
Abstract:Graph Neural Networks (GNNs) are powerful deep learning methods for Non-Euclidean data. Popular GNNs are message-passing algorithms (MPNNs) that aggregate and combine signals in a local graph neighborhood. However, shallow MPNNs tend to miss long-range signals and perform poorly on some heterophilous graphs, while deep MPNNs can suffer from issues like over-smoothing or over-squashing. To mitigate such issues, existing works typically borrow normalization techniques from training neural networks on Euclidean data or modify the graph structures. Yet these approaches are not well-understood theoretically and could increase the overall computational complexity. In this work, we draw inspirations from spectral graph embedding and propose $\texttt{PowerEmbed}$ -- a simple layer-wise normalization technique to boost MPNNs. We show $\texttt{PowerEmbed}$ can provably express the top-$k$ leading eigenvectors of the graph operator, which prevents over-smoothing and is agnostic to the graph topology; meanwhile, it produces a list of representations ranging from local features to global signals, which avoids over-squashing. We apply $\texttt{PowerEmbed}$ in a wide range of simulated and real graphs and demonstrate its competitive performance, particularly for heterophilous graphs.
Efficient and effective training of language and graph neural network models
Jun 22, 2022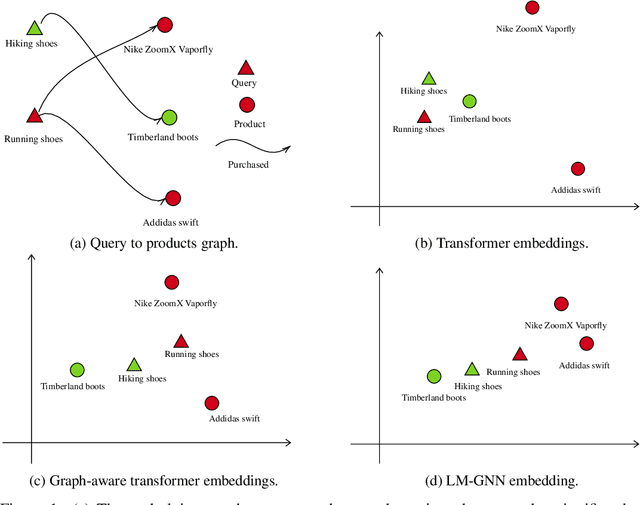
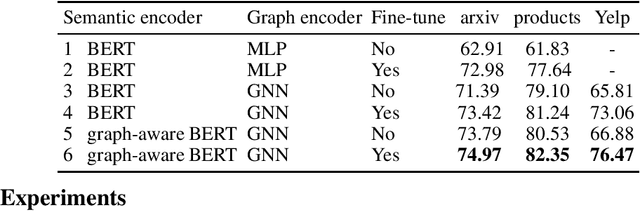
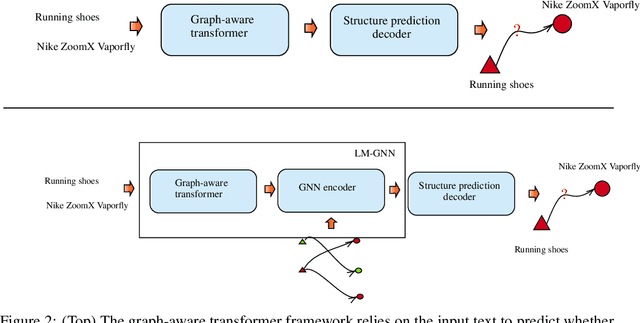
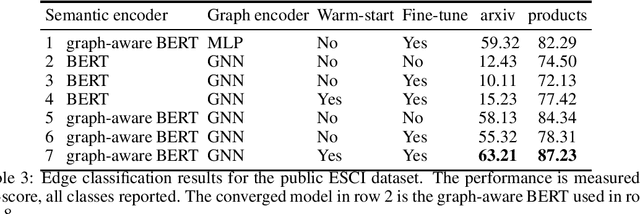
Abstract:Can we combine heterogenous graph structure with text to learn high-quality semantic and behavioural representations? Graph neural networks (GNN)s encode numerical node attributes and graph structure to achieve impressive performance in a variety of supervised learning tasks. Current GNN approaches are challenged by textual features, which typically need to be encoded to a numerical vector before provided to the GNN that may incur some information loss. In this paper, we put forth an efficient and effective framework termed language model GNN (LM-GNN) to jointly train large-scale language models and graph neural networks. The effectiveness in our framework is achieved by applying stage-wise fine-tuning of the BERT model first with heterogenous graph information and then with a GNN model. Several system and design optimizations are proposed to enable scalable and efficient training. LM-GNN accommodates node and edge classification as well as link prediction tasks. We evaluate the LM-GNN framework in different datasets performance and showcase the effectiveness of the proposed approach. LM-GNN provides competitive results in an Amazon query-purchase-product application.
Nimble GNN Embedding with Tensor-Train Decomposition
Jun 21, 2022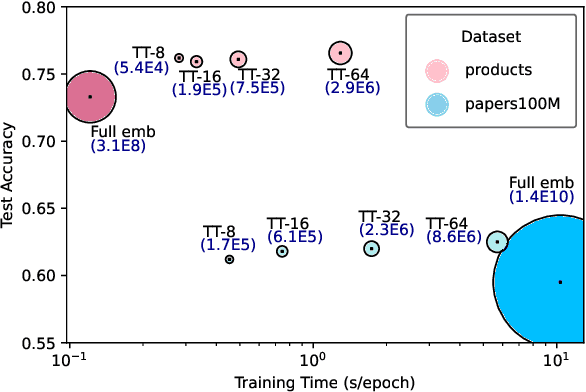
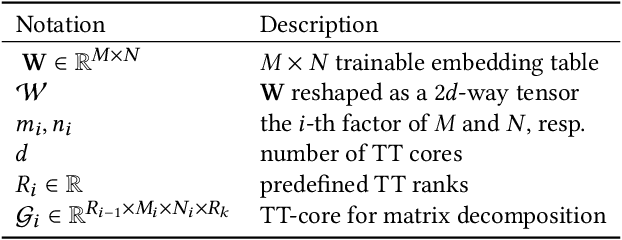

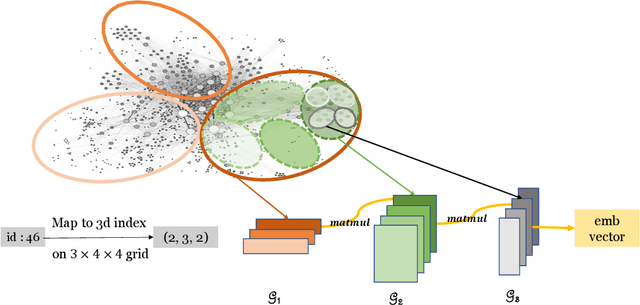
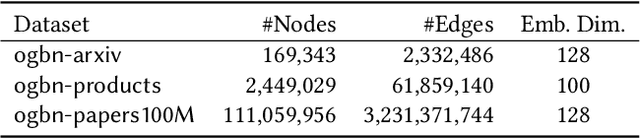
Abstract:This paper describes a new method for representing embedding tables of graph neural networks (GNNs) more compactly via tensor-train (TT) decomposition. We consider the scenario where (a) the graph data that lack node features, thereby requiring the learning of embeddings during training; and (b) we wish to exploit GPU platforms, where smaller tables are needed to reduce host-to-GPU communication even for large-memory GPUs. The use of TT enables a compact parameterization of the embedding, rendering it small enough to fit entirely on modern GPUs even for massive graphs. When combined with judicious schemes for initialization and hierarchical graph partitioning, this approach can reduce the size of node embedding vectors by 1,659 times to 81,362 times on large publicly available benchmark datasets, achieving comparable or better accuracy and significant speedups on multi-GPU systems. In some cases, our model without explicit node features on input can even match the accuracy of models that use node features.
TGL: A General Framework for Temporal GNN Training on Billion-Scale Graphs
Mar 28, 2022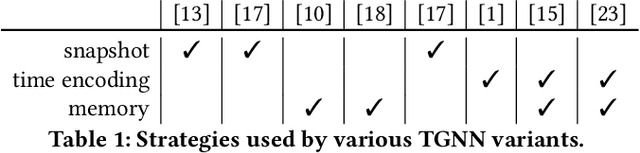
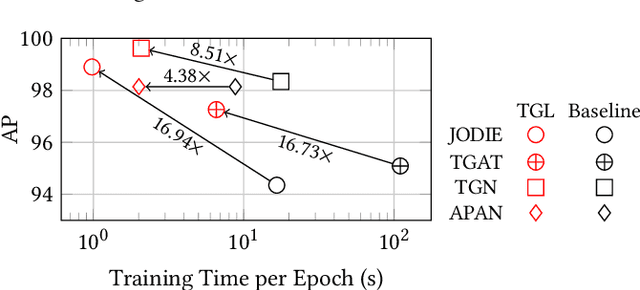
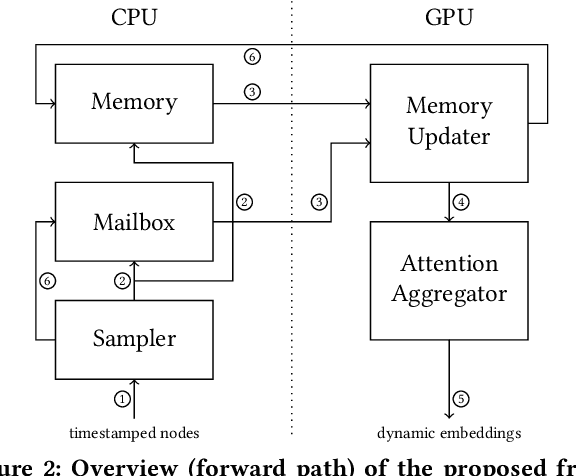
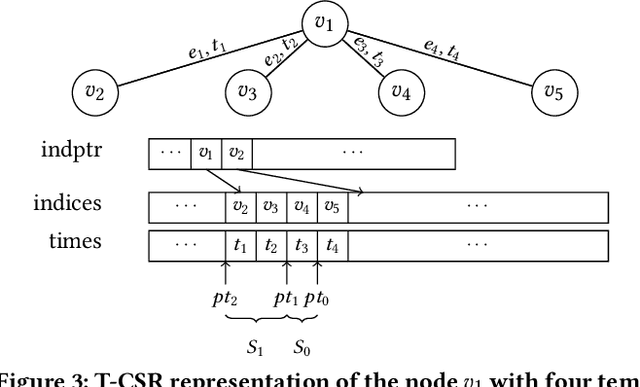
Abstract:Many real world graphs contain time domain information. Temporal Graph Neural Networks capture temporal information as well as structural and contextual information in the generated dynamic node embeddings. Researchers have shown that these embeddings achieve state-of-the-art performance in many different tasks. In this work, we propose TGL, a unified framework for large-scale offline Temporal Graph Neural Network training where users can compose various Temporal Graph Neural Networks with simple configuration files. TGL comprises five main components, a temporal sampler, a mailbox, a node memory module, a memory updater, and a message passing engine. We design a Temporal-CSR data structure and a parallel sampler to efficiently sample temporal neighbors to formtraining mini-batches. We propose a novel random chunk scheduling technique that mitigates the problem of obsolete node memory when training with a large batch size. To address the limitations of current TGNNs only being evaluated on small-scale datasets, we introduce two large-scale real-world datasets with 0.2 and 1.3 billion temporal edges. We evaluate the performance of TGL on four small-scale datasets with a single GPU and the two large datasets with multiple GPUs for both link prediction and node classification tasks. We compare TGL with the open-sourced code of five methods and show that TGL achieves similar or better accuracy with an average of 13x speedup. Our temporal parallel sampler achieves an average of 173x speedup on a multi-core CPU compared with the baselines. On a 4-GPU machine, TGL can train one epoch of more than one billion temporal edges within 1-10 hours. To the best of our knowledge, this is the first work that proposes a general framework for large-scale Temporal Graph Neural Networks training on multiple GPUs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge