"Information Extraction": models, code, and papers
Calibrated Seq2seq Models for Efficient and Generalizable Ultra-fine Entity Typing
Nov 01, 2023Ultra-fine entity typing plays a crucial role in information extraction by predicting fine-grained semantic types for entity mentions in text. However, this task poses significant challenges due to the massive number of entity types in the output space. The current state-of-the-art approaches, based on standard multi-label classifiers or cross-encoder models, suffer from poor generalization performance or inefficient inference. In this paper, we present CASENT, a seq2seq model designed for ultra-fine entity typing that predicts ultra-fine types with calibrated confidence scores. Our model takes an entity mention as input and employs constrained beam search to generate multiple types autoregressively. The raw sequence probabilities associated with the predicted types are then transformed into confidence scores using a novel calibration method. We conduct extensive experiments on the UFET dataset which contains over 10k types. Our method outperforms the previous state-of-the-art in terms of F1 score and calibration error, while achieving an inference speedup of over 50 times. Additionally, we demonstrate the generalization capabilities of our model by evaluating it in zero-shot and few-shot settings on five specialized domain entity typing datasets that are unseen during training. Remarkably, our model outperforms large language models with 10 times more parameters in the zero-shot setting, and when fine-tuned on 50 examples, it significantly outperforms ChatGPT on all datasets. Our code, models and demo are available at https://github.com/yanlinf/CASENT.
Joint Entity and Relation Extraction with Span Pruning and Hypergraph Neural Networks
Oct 26, 2023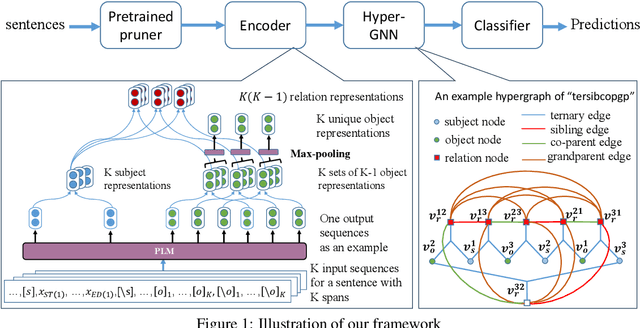
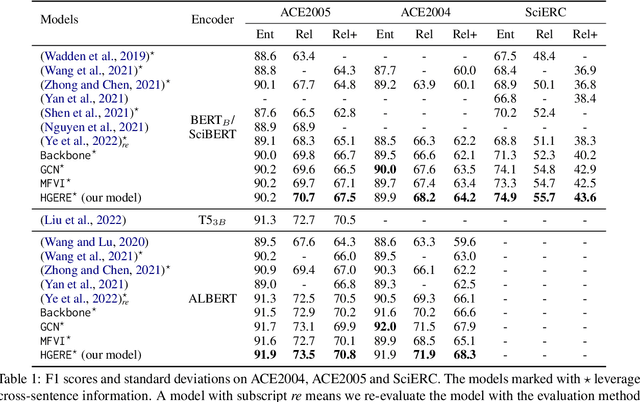
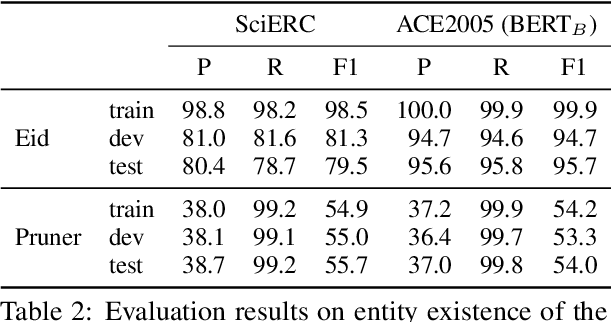
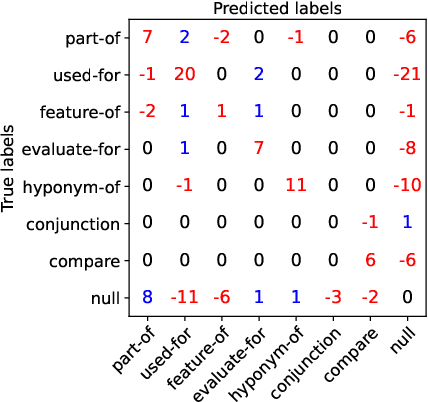
Entity and Relation Extraction (ERE) is an important task in information extraction. Recent marker-based pipeline models achieve state-of-the-art performance, but still suffer from the error propagation issue. Also, most of current ERE models do not take into account higher-order interactions between multiple entities and relations, while higher-order modeling could be beneficial.In this work, we propose HyperGraph neural network for ERE ($\hgnn{}$), which is built upon the PL-marker (a state-of-the-art marker-based pipleline model). To alleviate error propagation,we use a high-recall pruner mechanism to transfer the burden of entity identification and labeling from the NER module to the joint module of our model. For higher-order modeling, we build a hypergraph, where nodes are entities (provided by the span pruner) and relations thereof, and hyperedges encode interactions between two different relations or between a relation and its associated subject and object entities. We then run a hypergraph neural network for higher-order inference by applying message passing over the built hypergraph. Experiments on three widely used benchmarks (\acef{}, \ace{} and \scierc{}) for ERE task show significant improvements over the previous state-of-the-art PL-marker.
PV2TEA: Patching Visual Modality to Textual-Established Information Extraction
Jun 01, 2023
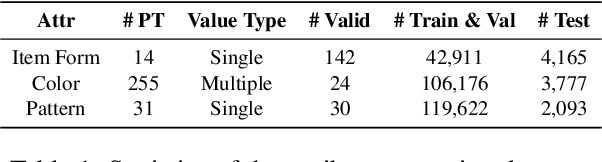

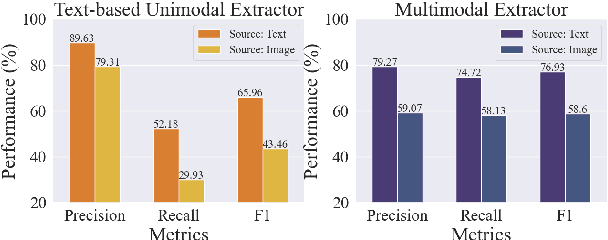
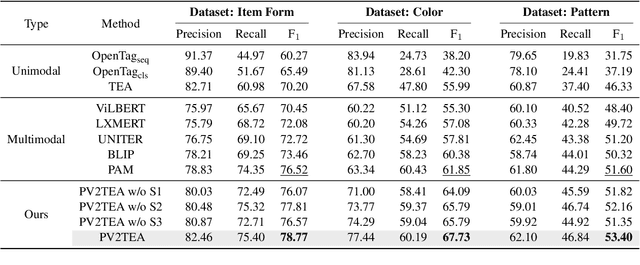
Information extraction, e.g., attribute value extraction, has been extensively studied and formulated based only on text. However, many attributes can benefit from image-based extraction, like color, shape, pattern, among others. The visual modality has long been underutilized, mainly due to multimodal annotation difficulty. In this paper, we aim to patch the visual modality to the textual-established attribute information extractor. The cross-modality integration faces several unique challenges: (C1) images and textual descriptions are loosely paired intra-sample and inter-samples; (C2) images usually contain rich backgrounds that can mislead the prediction; (C3) weakly supervised labels from textual-established extractors are biased for multimodal training. We present PV2TEA, an encoder-decoder architecture equipped with three bias reduction schemes: (S1) Augmented label-smoothed contrast to improve the cross-modality alignment for loosely-paired image and text; (S2) Attention-pruning that adaptively distinguishes the visual foreground; (S3) Two-level neighborhood regularization that mitigates the label textual bias via reliability estimation. Empirical results on real-world e-Commerce datasets demonstrate up to 11.74% absolute (20.97% relatively) F1 increase over unimodal baselines.
Data-Driven Information Extraction and Enrichment of Molecular Profiling Data for Cancer Cell Lines
Jul 03, 2023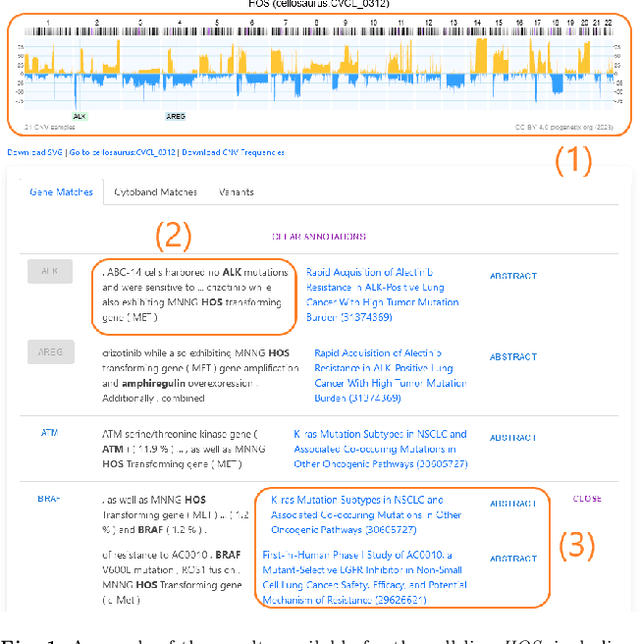
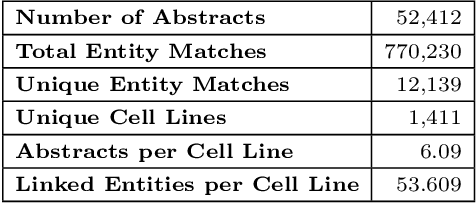

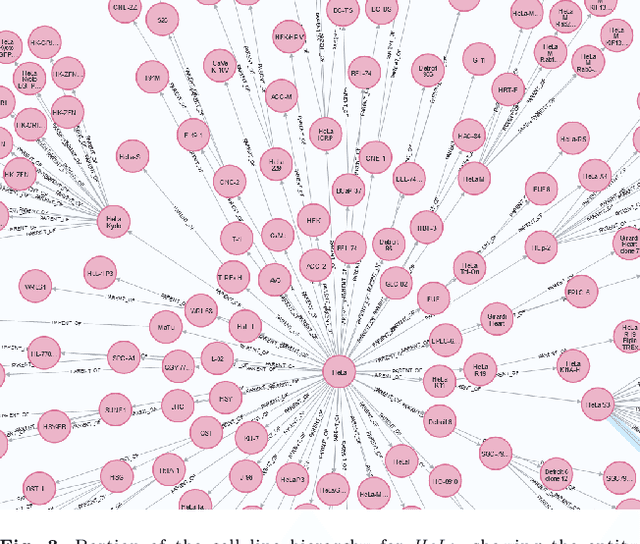
With the proliferation of research means and computational methodologies, published biomedical literature is growing exponentially in numbers and volume. As a consequence, in the fields of biological, medical and clinical research, domain experts have to sift through massive amounts of scientific text to find relevant information. However, this process is extremely tedious and slow to be performed by humans. Hence, novel computational information extraction and correlation mechanisms are required to boost meaningful knowledge extraction. In this work, we present the design, implementation and application of a novel data extraction and exploration system. This system extracts deep semantic relations between textual entities from scientific literature to enrich existing structured clinical data in the domain of cancer cell lines. We introduce a new public data exploration portal, which enables automatic linking of genomic copy number variants plots with ranked, related entities such as affected genes. Each relation is accompanied by literature-derived evidences, allowing for deep, yet rapid, literature search, using existing structured data as a springboard. Our system is publicly available on the web at https://cancercelllines.org
Vision-Enhanced Semantic Entity Recognition in Document Images via Visually-Asymmetric Consistency Learning
Oct 23, 2023Extracting meaningful entities belonging to predefined categories from Visually-rich Form-like Documents (VFDs) is a challenging task. Visual and layout features such as font, background, color, and bounding box location and size provide important cues for identifying entities of the same type. However, existing models commonly train a visual encoder with weak cross-modal supervision signals, resulting in a limited capacity to capture these non-textual features and suboptimal performance. In this paper, we propose a novel \textbf{V}isually-\textbf{A}symmetric co\textbf{N}sisten\textbf{C}y \textbf{L}earning (\textsc{Vancl}) approach that addresses the above limitation by enhancing the model's ability to capture fine-grained visual and layout features through the incorporation of color priors. Experimental results on benchmark datasets show that our approach substantially outperforms the strong LayoutLM series baseline, demonstrating the effectiveness of our approach. Additionally, we investigate the effects of different color schemes on our approach, providing insights for optimizing model performance. We believe our work will inspire future research on multimodal information extraction.
Augmented Kinesthetic Teaching: Enhancing Task Execution Efficiency through Intuitive Human Instructions
Dec 01, 2023In this paper, we present a complete and efficient implementation of a knowledge-sharing augmented kinesthetic teaching approach for efficient task execution in robotics. Our augmented kinesthetic teaching method integrates intuitive human feedback, including verbal, gesture, gaze, and physical guidance, to facilitate the extraction of multiple layers of task information including control type, attention direction, input and output type, action state change trigger, etc., enhancing the adaptability and autonomy of robots during task execution. We propose an efficient Programming by Demonstration (PbD) framework for users with limited technical experience to teach the robot in an intuitive manner. The proposed framework provides an interface for such users to teach customized tasks using high-level commands, with the goal of achieving a smoother teaching experience and task execution. This is demonstrated with the sample task of pouring water.
Semi-supervised Medical Image Segmentation via Query Distribution Consistency
Nov 21, 2023Semi-supervised learning is increasingly popular in medical image segmentation due to its ability to leverage large amounts of unlabeled data to extract additional information. However, most existing semi-supervised segmentation methods focus only on extracting information from unlabeled data. In this paper, we propose a novel Dual KMax UX-Net framework that leverages labeled data to guide the extraction of information from unlabeled data. Our approach is based on a mutual learning strategy that incorporates two modules: 3D UX-Net as our backbone meta-architecture and KMax decoder to enhance the segmentation performance. Extensive experiments on the Atrial Segmentation Challenge dataset have shown that our method can significantly improve performance by merging unlabeled data. Meanwhile, our framework outperforms state-of-the-art semi-supervised learning methods on 10\% and 20\% labeled settings. Code located at: https://github.com/Rows21/DK-UXNet.
Tracking electricity losses and their perceived causes using nighttime light and social media
Oct 18, 2023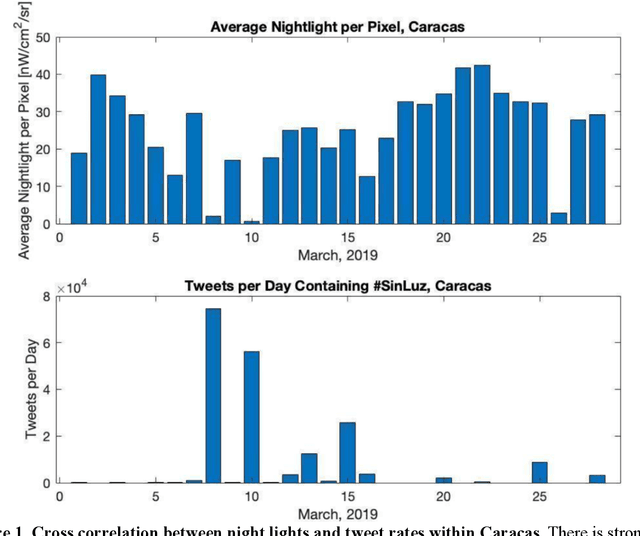
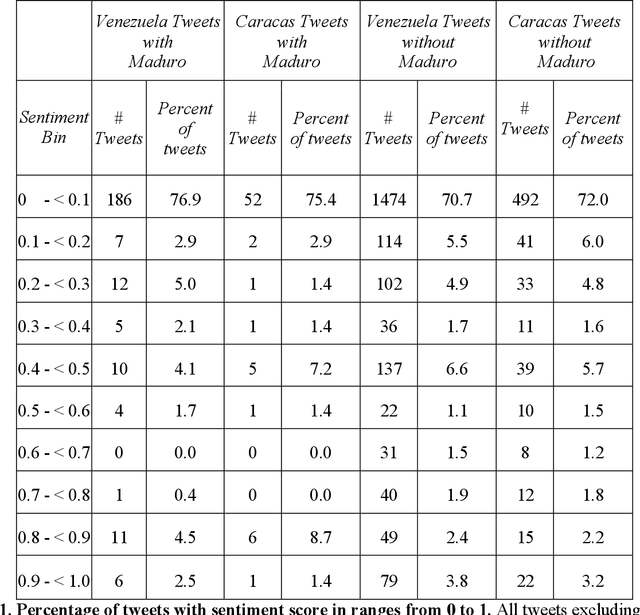

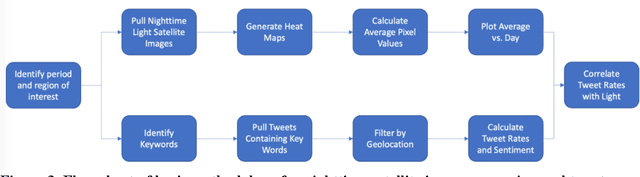
Urban environments are intricate systems where the breakdown of critical infrastructure can impact both the economic and social well-being of communities. Electricity systems hold particular significance, as they are essential for other infrastructure, and disruptions can trigger widespread consequences. Typically, assessing electricity availability requires ground-level data, a challenge in conflict zones and regions with limited access. This study shows how satellite imagery, social media, and information extraction can monitor blackouts and their perceived causes. Night-time light data (in March 2019 for Caracas, Venezuela) is used to indicate blackout regions. Twitter data is used to determine sentiment and topic trends, while statistical analysis and topic modeling delved into public perceptions regarding blackout causes. The findings show an inverse relationship between nighttime light intensity. Tweets mentioning the Venezuelan President displayed heightened negativity and a greater prevalence of blame-related terms, suggesting a perception of government accountability for the outages.
InteractiveIE: Towards Assessing the Strength of Human-AI Collaboration in Improving the Performance of Information Extraction
May 24, 2023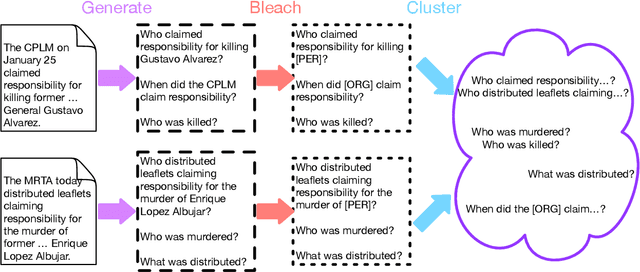
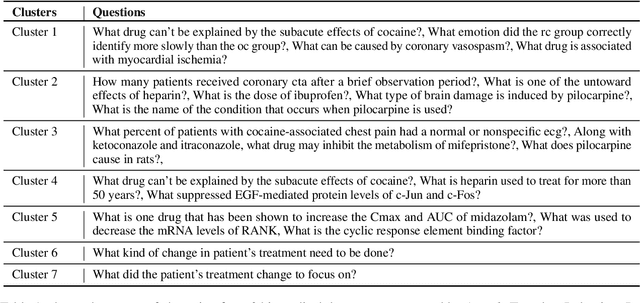

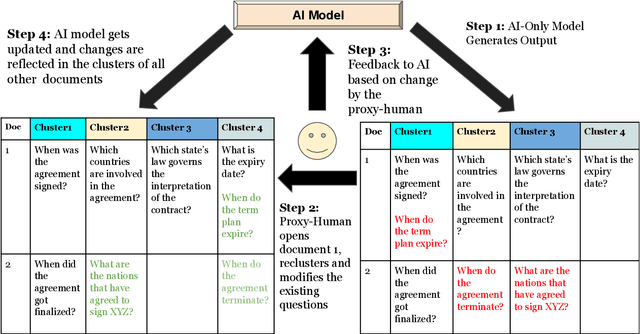
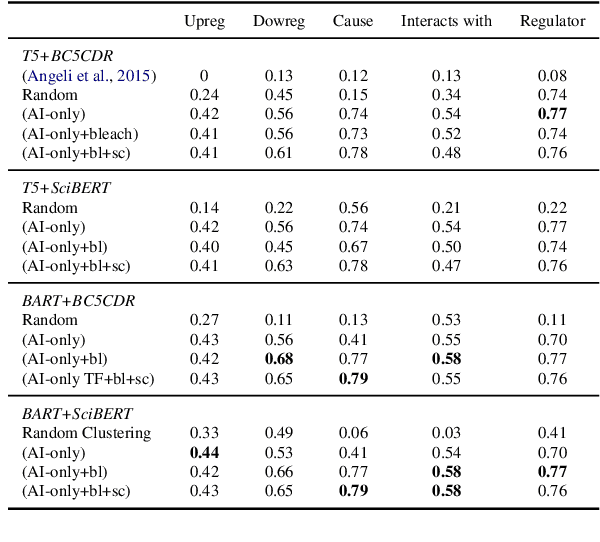
Learning template based information extraction from documents is a crucial yet difficult task. Prior template-based IE approaches assume foreknowledge of the domain templates; however, real-world IE do not have pre-defined schemas and it is a figure-out-as you go phenomena. To quickly bootstrap templates in a real-world setting, we need to induce template slots from documents with zero or minimal supervision. Since the purpose of question answering intersect with the goal of information extraction, we use automatic question generation to induce template slots from the documents and investigate how a tiny amount of a proxy human-supervision on-the-fly (termed as InteractiveIE) can further boost the performance. Extensive experiments on biomedical and legal documents, where obtaining training data is expensive, reveal encouraging trends of performance improvement using InteractiveIE over AI-only baseline.
Leveraging Knowledge Graphs for Orphan Entity Allocation in Resume Processing
Oct 21, 2023Significant challenges are posed in talent acquisition and recruitment by processing and analyzing unstructured data, particularly resumes. This research presents a novel approach for orphan entity allocation in resume processing using knowledge graphs. Techniques of association mining, concept extraction, external knowledge linking, named entity recognition, and knowledge graph construction are integrated into our pipeline. By leveraging these techniques, the aim is to automate and enhance the efficiency of the job screening process by successfully bucketing orphan entities within resumes. This allows for more effective matching between candidates and job positions, streamlining the resume screening process, and enhancing the accuracy of candidate-job matching. The approach's exceptional effectiveness and resilience are highlighted through extensive experimentation and evaluation, ensuring that alternative measures can be relied upon for seamless processing and orphan entity allocation in case of any component failure. The capabilities of knowledge graphs in generating valuable insights through intelligent information extraction and representation, specifically in the domain of categorizing orphan entities, are highlighted by the results of our research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge