"Image": models, code, and papers
Decoupled Gradient Harmonized Detector for Partial Annotation: Application to Signet Ring Cell Detection
Apr 09, 2020
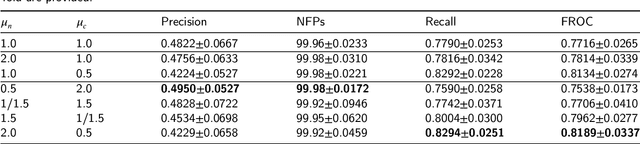
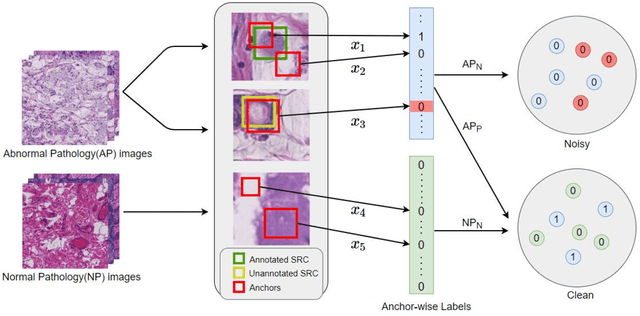
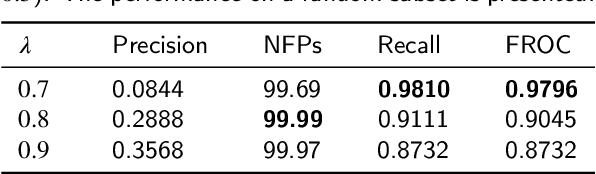
Early diagnosis of signet ring cell carcinoma dramatically improves the survival rate of patients. Due to lack of public dataset and expert-level annotations, automatic detection on signet ring cell (SRC) has not been thoroughly investigated. In MICCAI DigestPath2019 challenge, apart from foreground (SRC region)-background (normal tissue area) class imbalance, SRCs are partially annotated due to costly medical image annotation, which introduces extra label noise. To address the issues simultaneously, we propose Decoupled Gradient Harmonizing Mechanism (DGHM) and embed it into classification loss, denoted as DGHM-C loss. Specifically, besides positive (SRCs) and negative (normal tissues) examples, we further decouple noisy examples from clean examples and harmonize the corresponding gradient distributions in classification respectively. Without whistles and bells, we achieved the 2nd place in the challenge. Ablation studies and controlled label missing rate experiments demonstrate that DGHM-C loss can bring substantial improvement in partially annotated object detection.
Visualization of Convolutional Neural Networks for Monocular Depth Estimation
Apr 06, 2019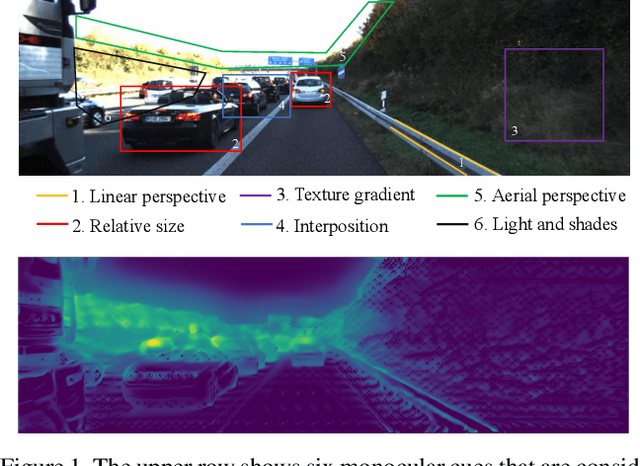
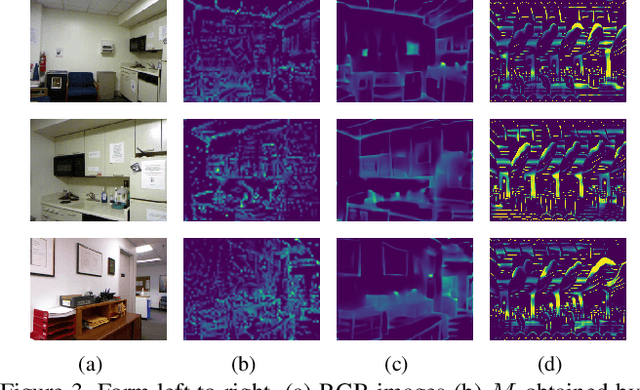
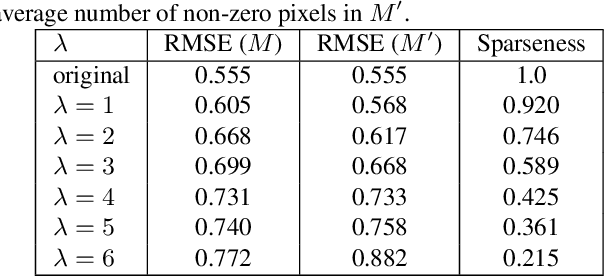
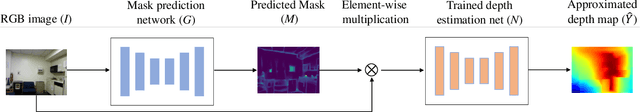

Recently, convolutional neural networks (CNNs) have shown great success on the task of monocular depth estimation. A fundamental yet unanswered question is: how CNNs can infer depth from a single image. Toward answering this question, we consider visualization of inference of a CNN by identifying relevant pixels of an input image to depth estimation. We formulate it as an optimization problem of identifying the smallest number of image pixels from which the CNN can estimate a depth map with the minimum difference from the estimate from the entire image. To cope with a difficulty with optimization through a deep CNN, we propose to use another network to predict those relevant image pixels in a forward computation. In our experiments, we first show the effectiveness of this approach, and then apply it to different depth estimation networks on indoor and outdoor scene datasets. The results provide several findings that help exploration of the above question.
Multi-Stage Variational Auto-Encoders for Coarse-to-Fine Image Generation
May 19, 2017
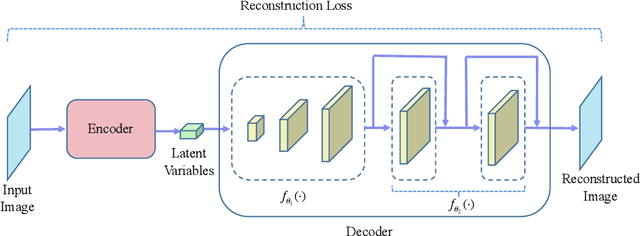
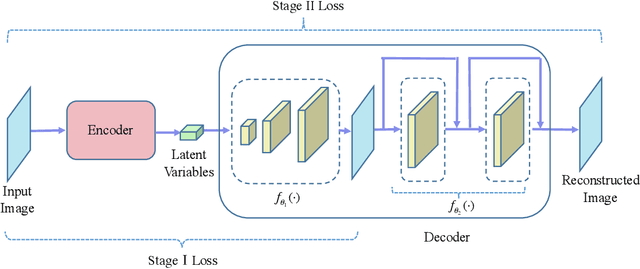

Variational auto-encoder (VAE) is a powerful unsupervised learning framework for image generation. One drawback of VAE is that it generates blurry images due to its Gaussianity assumption and thus L2 loss. To allow the generation of high quality images by VAE, we increase the capacity of decoder network by employing residual blocks and skip connections, which also enable efficient optimization. To overcome the limitation of L2 loss, we propose to generate images in a multi-stage manner from coarse to fine. In the simplest case, the proposed multi-stage VAE divides the decoder into two components in which the second component generates refined images based on the course images generated by the first component. Since the second component is independent of the VAE model, it can employ other loss functions beyond the L2 loss and different model architectures. The proposed framework can be easily generalized to contain more than two components. Experiment results on the MNIST and CelebA datasets demonstrate that the proposed multi-stage VAE can generate sharper images as compared to those from the original VAE.
Generative Smoke Removal
Feb 01, 2019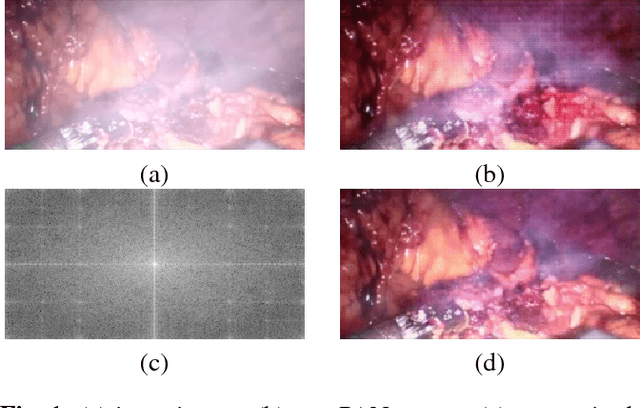
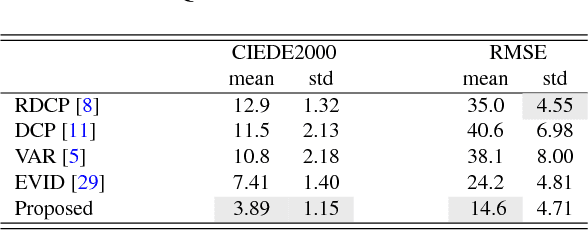
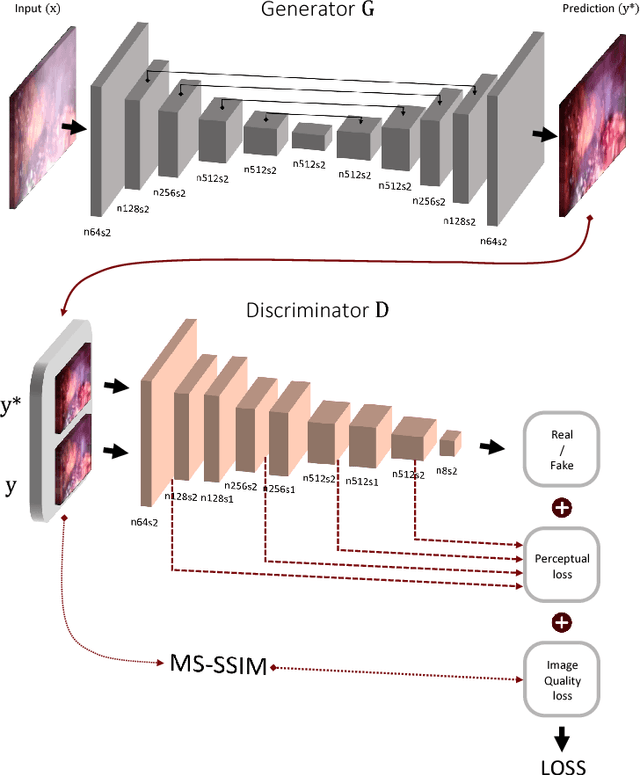
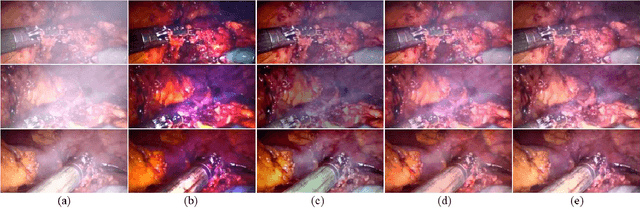
In minimally invasive surgery, the use of tissue dissection tools causes smoke, which inevitably degrades the image quality. This could reduce the visibility of the operation field for surgeons and introduces errors for the computer vision algorithms used in surgical navigation systems. In this paper, we propose a novel approach for computational smoke removal using supervised image-to-image translation. We demonstrate that straightforward application of existing generative algorithms allows removing smoke but decreases image quality and introduces synthetic noise (grid-structure). Thus, we propose to solve this issue by modification of GAN's architecture and adding perceptual image quality metric to the loss function. Obtained results demonstrate that proposed method efficiently removes smoke as well as preserves perceptually sufficient image quality.
Sketch-BERT: Learning Sketch Bidirectional Encoder Representation from Transformers by Self-supervised Learning of Sketch Gestalt
May 19, 2020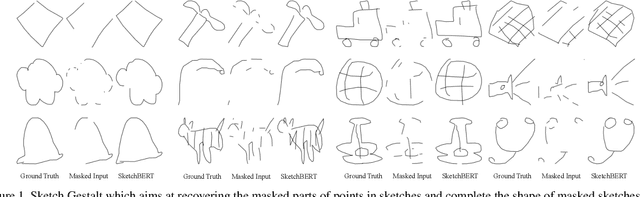
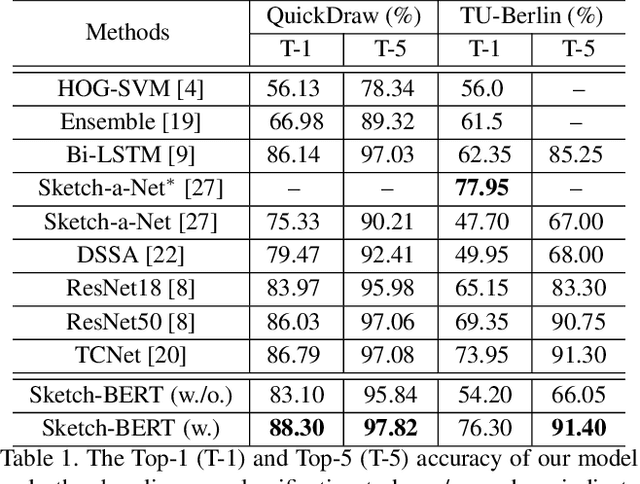
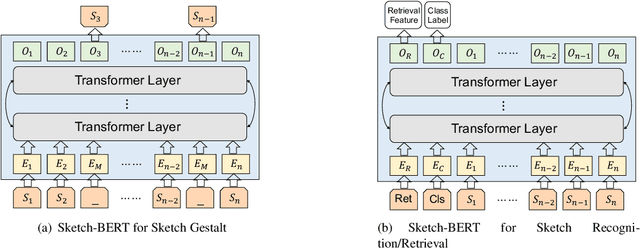
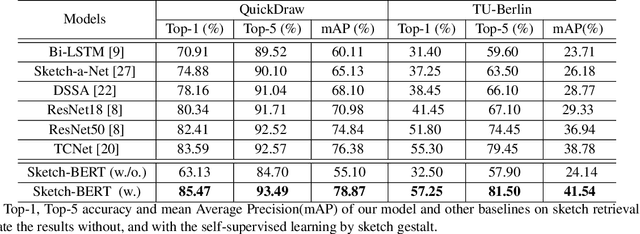
Previous researches of sketches often considered sketches in pixel format and leveraged CNN based models in the sketch understanding. Fundamentally, a sketch is stored as a sequence of data points, a vector format representation, rather than the photo-realistic image of pixels. SketchRNN studied a generative neural representation for sketches of vector format by Long Short Term Memory networks (LSTM). Unfortunately, the representation learned by SketchRNN is primarily for the generation tasks, rather than the other tasks of recognition and retrieval of sketches. To this end and inspired by the recent BERT model, we present a model of learning Sketch Bidirectional Encoder Representation from Transformer (Sketch-BERT). We generalize BERT to sketch domain, with the novel proposed components and pre-training algorithms, including the newly designed sketch embedding networks, and the self-supervised learning of sketch gestalt. Particularly, towards the pre-training task, we present a novel Sketch Gestalt Model (SGM) to help train the Sketch-BERT. Experimentally, we show that the learned representation of Sketch-BERT can help and improve the performance of the downstream tasks of sketch recognition, sketch retrieval, and sketch gestalt.
Learned reconstructions for practical mask-based lensless imaging
Aug 30, 2019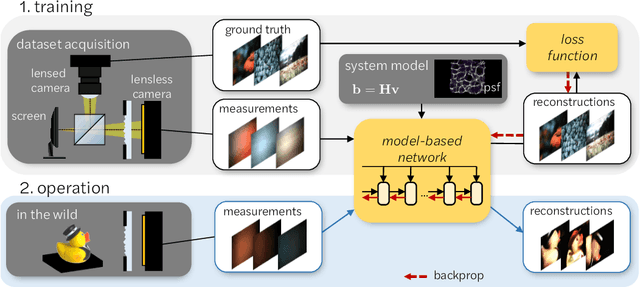
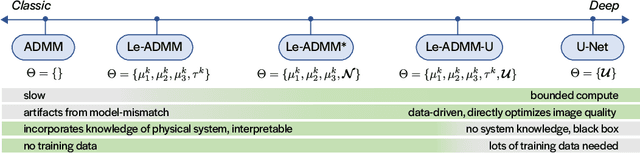
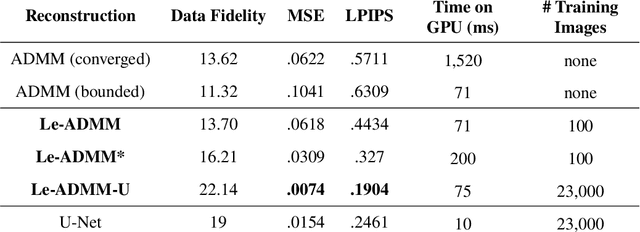
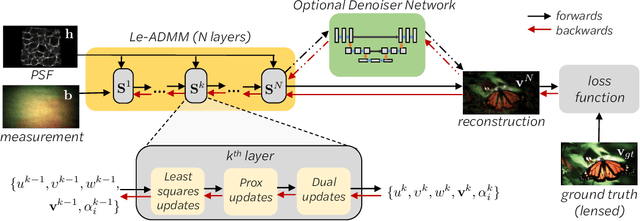
Mask-based lensless imagers are smaller and lighter than traditional lensed cameras. In these imagers, the sensor does not directly record an image of the scene; rather, a computational algorithm reconstructs it. Typically, mask-based lensless imagers use a model-based reconstruction approach that suffers from long compute times and a heavy reliance on both system calibration and heuristically chosen denoisers. In this work, we address these limitations using a bounded-compute, trainable neural network to reconstruct the image. We leverage our knowledge of the physical system by unrolling a traditional model-based optimization algorithm, whose parameters we optimize using experimentally gathered ground-truth data. Optionally, images produced by the unrolled network are then fed into a jointly-trained denoiser. As compared to traditional methods, our architecture achieves better perceptual image quality and runs 20x faster, enabling interactive previewing of the scene. We explore a spectrum between model-based and deep learning methods, showing the benefits of using an intermediate approach. Finally, we test our network on images taken in the wild with a prototype mask-based camera, demonstrating that our network generalizes to natural images.
Medical Image Fusion: A survey of the state of the art
Dec 31, 2013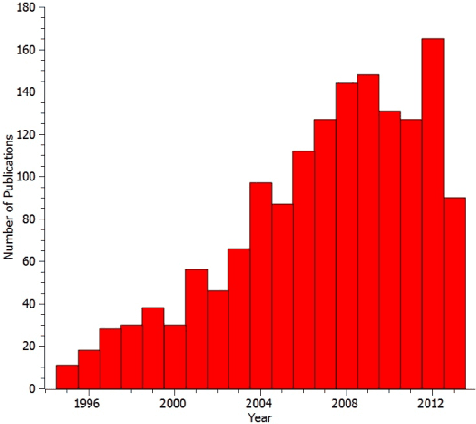
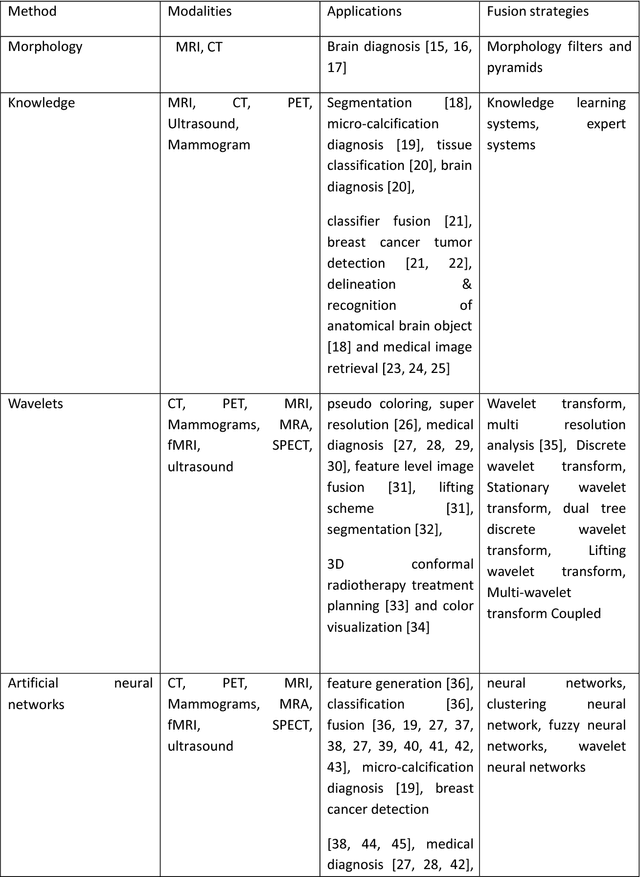
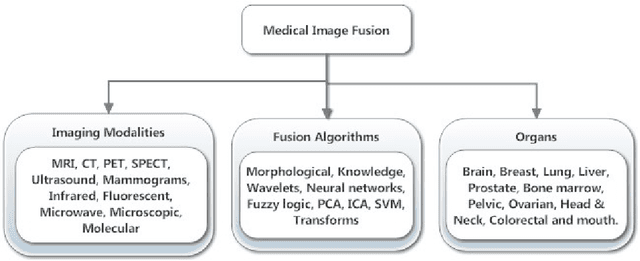
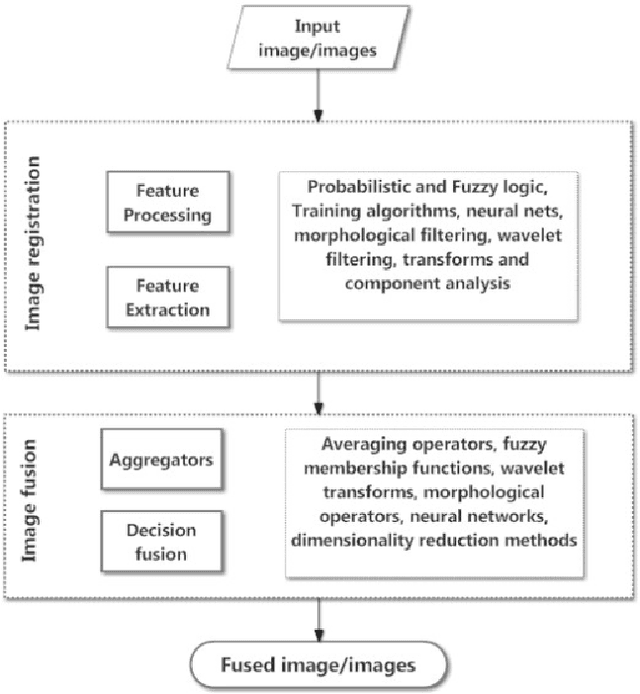
Medical image fusion is the process of registering and combining multiple images from single or multiple imaging modalities to improve the imaging quality and reduce randomness and redundancy in order to increase the clinical applicability of medical images for diagnosis and assessment of medical problems. Multi-modal medical image fusion algorithms and devices have shown notable achievements in improving clinical accuracy of decisions based on medical images. This review article provides a factual listing of methods and summarizes the broad scientific challenges faced in the field of medical image fusion. We characterize the medical image fusion research based on (1) the widely used image fusion methods, (2) imaging modalities, and (3) imaging of organs that are under study. This review concludes that even though there exists several open ended technological and scientific challenges, the fusion of medical images has proved to be useful for advancing the clinical reliability of using medical imaging for medical diagnostics and analysis, and is a scientific discipline that has the potential to significantly grow in the coming years.
Feedback Recurrent Autoencoder for Video Compression
Apr 09, 2020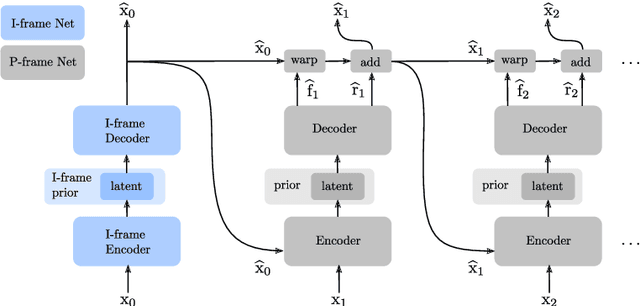
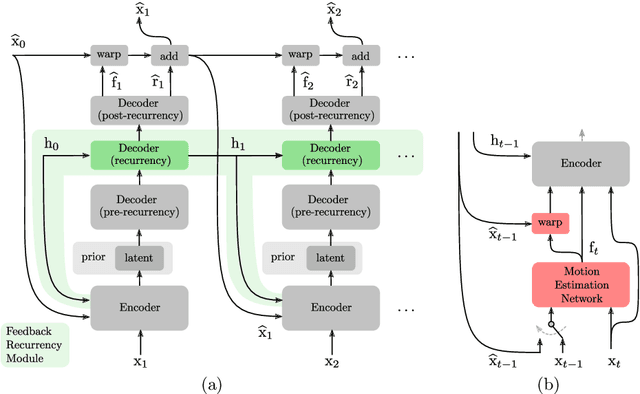
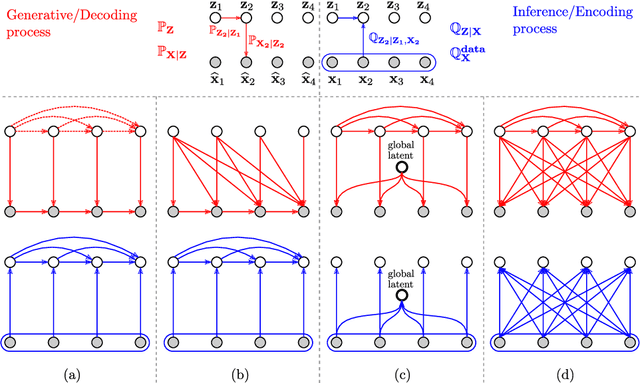
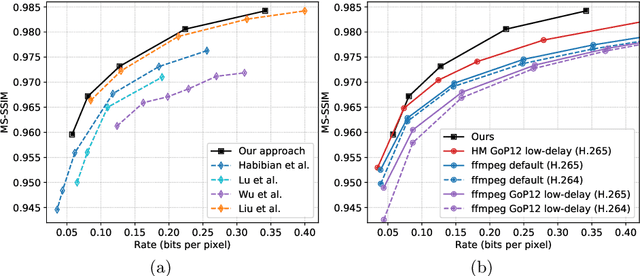
Recent advances in deep generative modeling have enabled efficient modeling of high dimensional data distributions and opened up a new horizon for solving data compression problems. Specifically, autoencoder based learned image or video compression solutions are emerging as strong competitors to traditional approaches. In this work, We propose a new network architecture, based on common and well studied components, for learned video compression operating in low latency mode. Our method yields state of the art MS-SSIM/rate performance on the high-resolution UVG dataset, among both learned video compression approaches and classical video compression methods (H.265 and H.264) in the rate range of interest for streaming applications. Additionally, we provide an analysis of existing approaches through the lens of their underlying probabilistic graphical models. Finally, we point out issues with temporal consistency and color shift observed in empirical evaluation, and suggest directions forward to alleviate those.
Domain Adaptive Relational Reasoning for 3D Multi-Organ Segmentation
May 18, 2020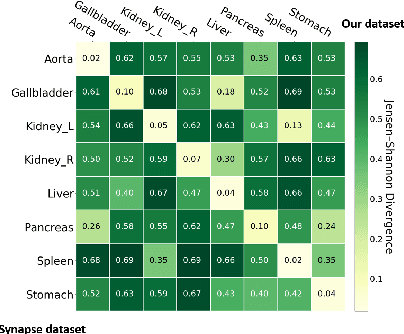
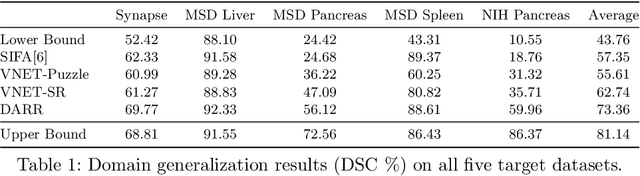
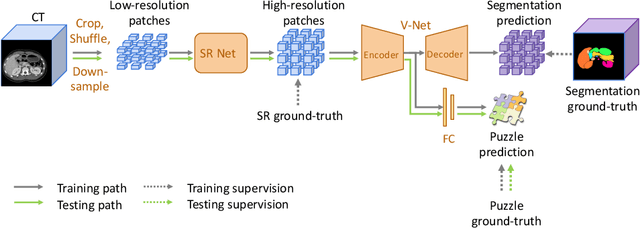
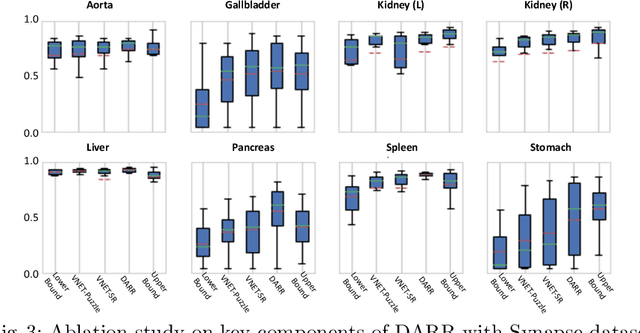
In this paper, we present a novel unsupervised domain adaptation (UDA) method, named Domain Adaptive Relational Reasoning (DARR), to generalize 3D multi-organ segmentation models to medical data collected from different scanners and/or protocols (domains). Our method is inspired by the fact that the spatial relationship between internal structures in medical images is relatively fixed, e.g., a spleen is always located at the tail of a pancreas, which serves as a latent variable to transfer the knowledge shared across multiple domains. We formulate the spatial relationship by solving a jigsaw puzzle task, i.e., recovering a CT scan from its shuffled patches, and jointly train it with the organ segmentation task. To guarantee the transferability of the learned spatial relationship to multiple domains, we additionally introduce two schemes: 1) Employing a super-resolution network also jointly trained with the segmentation model to standardize medical images from different domain to a certain spatial resolution; 2) Adapting the spatial relationship for a test image by test-time jigsaw puzzle training. Experimental results show that our method improves the performance by 29.60\% DSC on target datasets on average without using any data from the target domain during training.
Two-View Fine-grained Classification of Plant Species
May 18, 2020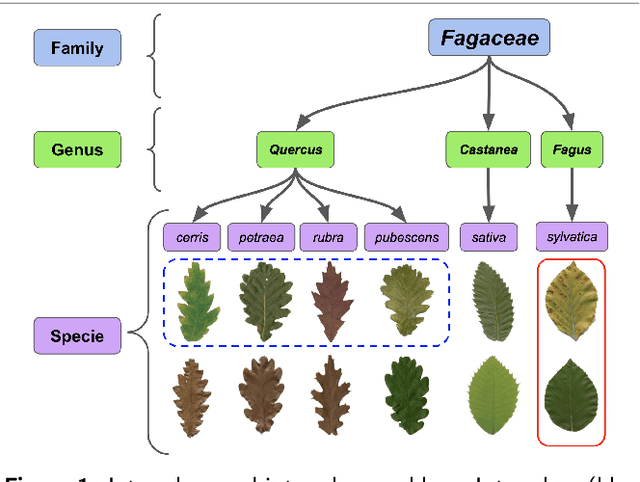
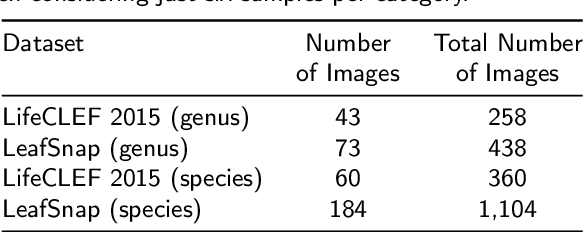
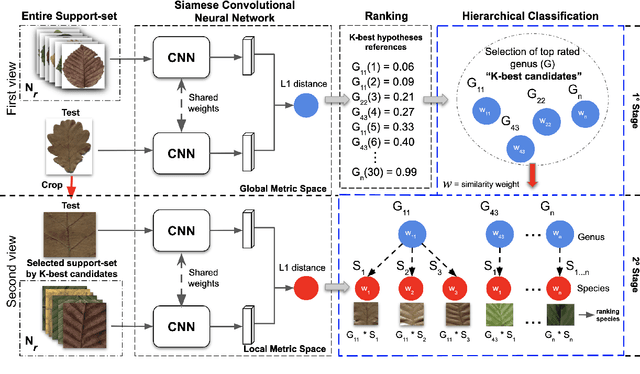
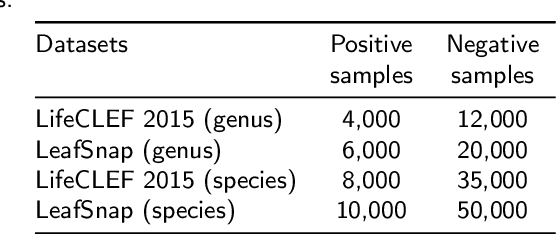
Automatic plant classification is a challenging problem due to the wide biodiversity of the existing plant species in a fine-grained scenario. Powerful deep learning architectures have been used to improve the classification performance in such a fine-grained problem, but usually building models that are highly dependent on a large training dataset and which are not scalable. In this paper, we propose a novel method based on a two-view leaf image representation and a hierarchical classification strategy for fine-grained recognition of plant species. It uses the botanical taxonomy as a basis for a coarse-to-fine strategy applied to identify the plant genus and species. The two-view representation provides complementary global and local features of leaf images. A deep metric based on Siamese convolutional neural networks is used to reduce the dependence on a large number of training samples and make the method scalable to new plant species. The experimental results on two challenging fine-grained datasets of leaf images (i.e. LifeCLEF 2015 and LeafSnap) have shown the effectiveness of the proposed method, which achieved recognition accuracy of 0.87 and 0.96 respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge