Kristina Monakhova
System- and Sample-agnostic Isotropic 3D Microscopy by Weakly Physics-informed, Domain-shift-resistant Axial Deblurring
Jun 10, 2024



Abstract:Three-dimensional (3D) subcellular imaging is essential for biomedical research, but the diffraction limit of optical microscopy compromises axial resolution, hindering accurate 3D structural analysis. This challenge is particularly pronounced in label-free imaging of thick, heterogeneous tissues, where assumptions about data distribution (e.g. sparsity, label-specific distribution, and lateral-axial similarity) and system priors (e.g. independent and identically distributed (i.i.d.) noise and linear shift-invariant (LSI) point-spread functions (PSFs)) are often invalid. Here, we introduce SSAI-3D, a weakly physics-informed, domain-shift-resistant framework for robust isotropic 3D imaging. SSAI-3D enables robust axial deblurring by generating a PSF-flexible, noise-resilient, sample-informed training dataset and sparsely fine-tuning a large pre-trained blind deblurring network. SSAI-3D was applied to label-free nonlinear imaging of living organoids, freshly excised human endometrium tissue, and mouse whisker pads, and further validated in publicly available ground-truth-paired experimental datasets of 3D heterogeneous biological tissues with unknown blurring and noise across different microscopy systems.
Learned, Uncertainty-driven Adaptive Acquisition for Photon-Efficient Multiphoton Microscopy
Oct 24, 2023Abstract:Multiphoton microscopy (MPM) is a powerful imaging tool that has been a critical enabler for live tissue imaging. However, since most multiphoton microscopy platforms rely on point scanning, there is an inherent trade-off between acquisition time, field of view (FOV), phototoxicity, and image quality, often resulting in noisy measurements when fast, large FOV, and/or gentle imaging is needed. Deep learning could be used to denoise multiphoton microscopy measurements, but these algorithms can be prone to hallucination, which can be disastrous for medical and scientific applications. We propose a method to simultaneously denoise and predict pixel-wise uncertainty for multiphoton imaging measurements, improving algorithm trustworthiness and providing statistical guarantees for the deep learning predictions. Furthermore, we propose to leverage this learned, pixel-wise uncertainty to drive an adaptive acquisition technique that rescans only the most uncertain regions of a sample. We demonstrate our method on experimental noisy MPM measurements of human endometrium tissues, showing that we can maintain fine features and outperform other denoising methods while predicting uncertainty at each pixel. Finally, with our adaptive acquisition technique, we demonstrate a 120X reduction in acquisition time and total light dose while successfully recovering fine features in the sample. We are the first to demonstrate distribution-free uncertainty quantification for a denoising task with real experimental data and the first to propose adaptive acquisition based on reconstruction uncertainty
Roadmap on Deep Learning for Microscopy
Mar 07, 2023



Abstract:Through digital imaging, microscopy has evolved from primarily being a means for visual observation of life at the micro- and nano-scale, to a quantitative tool with ever-increasing resolution and throughput. Artificial intelligence, deep neural networks, and machine learning are all niche terms describing computational methods that have gained a pivotal role in microscopy-based research over the past decade. This Roadmap is written collectively by prominent researchers and encompasses selected aspects of how machine learning is applied to microscopy image data, with the aim of gaining scientific knowledge by improved image quality, automated detection, segmentation, classification and tracking of objects, and efficient merging of information from multiple imaging modalities. We aim to give the reader an overview of the key developments and an understanding of possibilities and limitations of machine learning for microscopy. It will be of interest to a wide cross-disciplinary audience in the physical sciences and life sciences.
Dancing under the stars: video denoising in starlight
Apr 08, 2022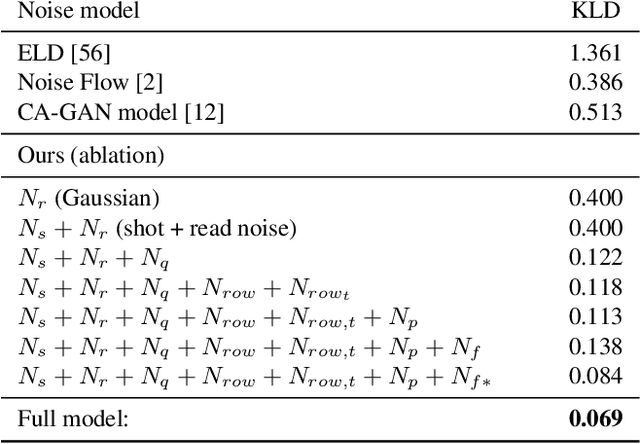
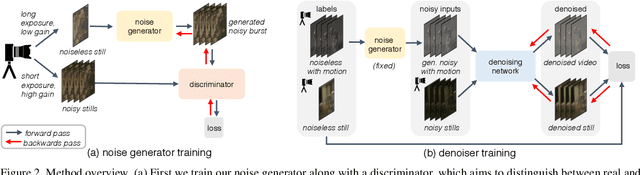
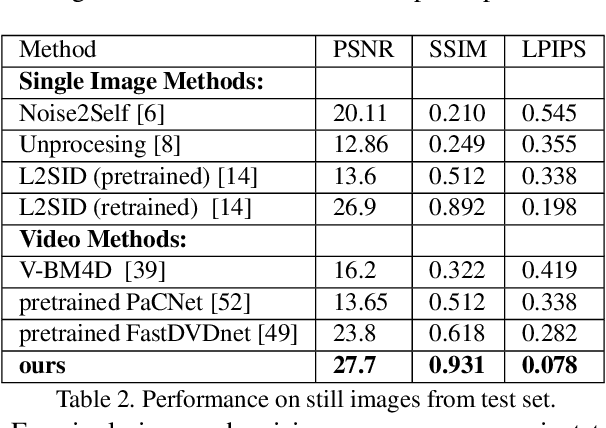
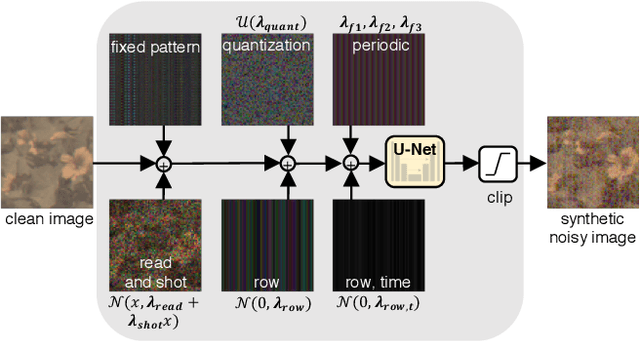
Abstract:Imaging in low light is extremely challenging due to low photon counts. Using sensitive CMOS cameras, it is currently possible to take videos at night under moonlight (0.05-0.3 lux illumination). In this paper, we demonstrate photorealistic video under starlight (no moon present, $<$0.001 lux) for the first time. To enable this, we develop a GAN-tuned physics-based noise model to more accurately represent camera noise at the lowest light levels. Using this noise model, we train a video denoiser using a combination of simulated noisy video clips and real noisy still images. We capture a 5-10 fps video dataset with significant motion at approximately 0.6-0.7 millilux with no active illumination. Comparing against alternative methods, we achieve improved video quality at the lowest light levels, demonstrating photorealistic video denoising in starlight for the first time.
Untrained networks for compressive lensless photography
Mar 13, 2021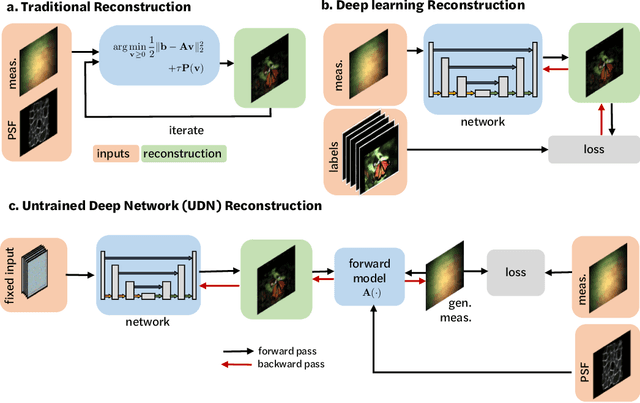
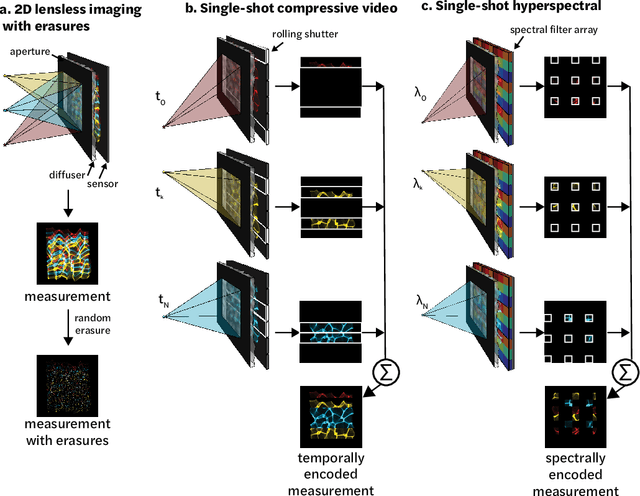
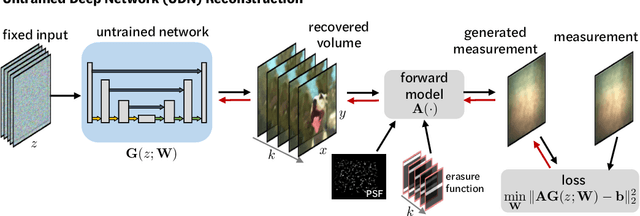
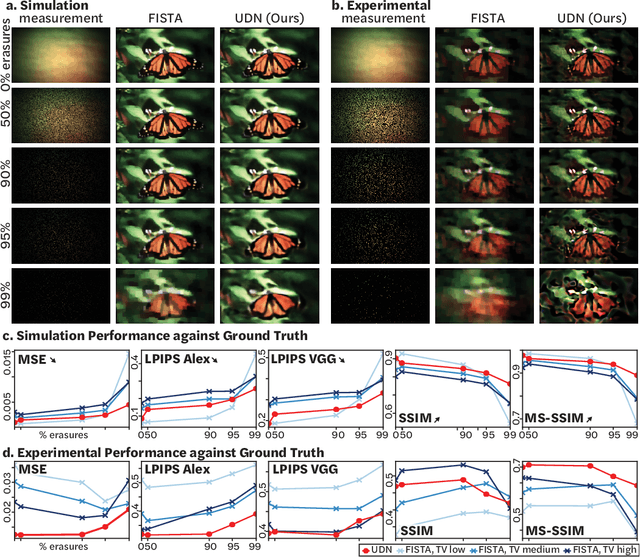
Abstract:Compressive lensless imagers enable novel applications in an extremely compact device, requiring only a phase or amplitude mask placed close to the sensor. They have been demonstrated for 2D and 3D microscopy, single-shot video, and single-shot hyperspectral imaging; in each of these cases, a compressive-sensing-based inverse problem is solved in order to recover a 3D data-cube from a 2D measurement. Typically, this is accomplished using convex optimization and hand-picked priors. Alternatively, deep learning-based reconstruction methods offer the promise of better priors, but require many thousands of ground truth training pairs, which can be difficult or impossible to acquire. In this work, we propose the use of untrained networks for compressive image recovery. Our approach does not require any labeled training data, but instead uses the measurement itself to update the network weights. We demonstrate our untrained approach on lensless compressive 2D imaging as well as single-shot high-speed video recovery using the camera's rolling shutter, and single-shot hyperspectral imaging. We provide simulation and experimental verification, showing that our method results in improved image quality over existing methods.
Miniscope3D: optimized single-shot miniature 3D fluorescence microscopy
Oct 12, 2020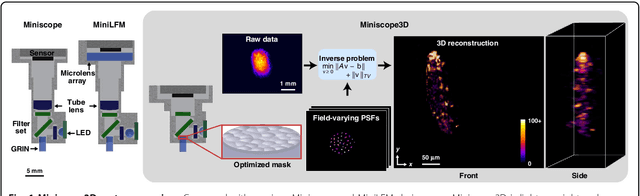
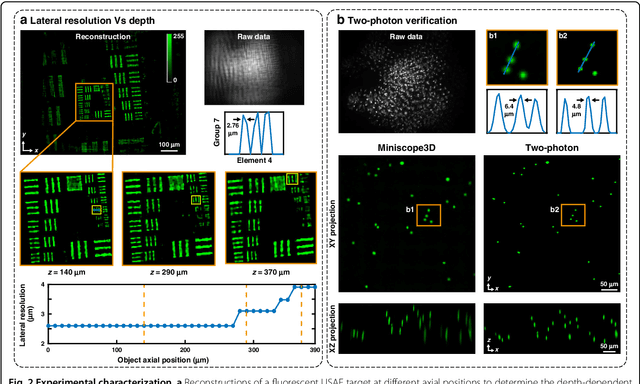
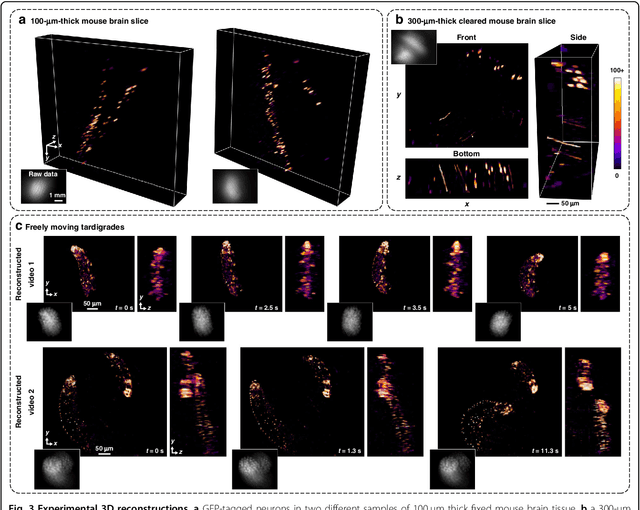
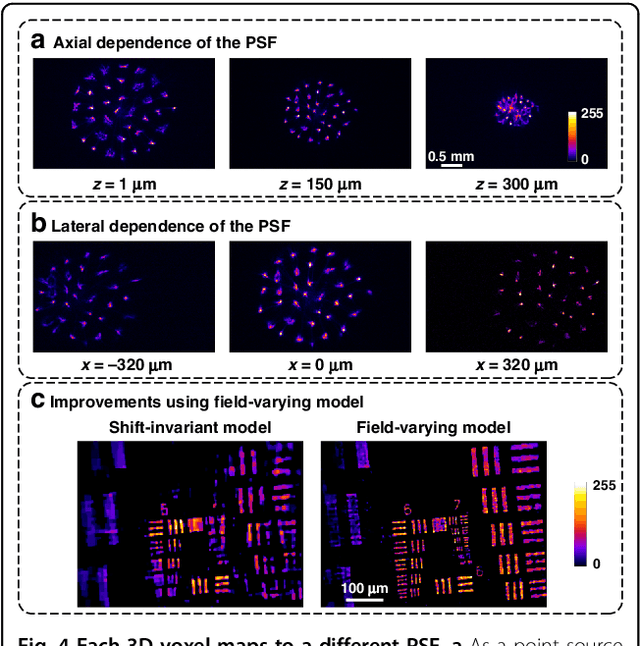
Abstract:Miniature fluorescence microscopes are a standard tool in systems biology. However, widefield miniature microscopes capture only 2D information, and modifications that enable 3D capabilities increase the size and weight and have poor resolution outside a narrow depth range. Here, we achieve the 3D capability by replacing the tube lens of a conventional 2D Miniscope with an optimized multifocal phase mask at the objective's aperture stop. Placing the phase mask at the aperture stop significantly reduces the size of the device, and varying the focal lengths enables a uniform resolution across a wide depth range. The phase mask encodes the 3D fluorescence intensity into a single 2D measurement, and the 3D volume is recovered by solving a sparsity-constrained inverse problem. We provide methods for designing and fabricating the phase mask and an efficient forward model that accounts for the field-varying aberrations in miniature objectives. We demonstrate a prototype that is 17 mm tall and weighs 2.5 grams, achieving 2.76 $\mu$m lateral, and 15 $\mu$m axial resolution across most of the 900x700x390 $\mu m^3$ volume at 40 volumes per second. The performance is validated experimentally on resolution targets, dynamic biological samples, and mouse brain tissue. Compared with existing miniature single-shot volume-capture implementations, our system is smaller and lighter and achieves a more than 2x better lateral and axial resolution throughout a 10x larger usable depth range. Our microscope design provides single-shot 3D imaging for applications where a compact platform matters, such as volumetric neural imaging in freely moving animals and 3D motion studies of dynamic samples in incubators and lab-on-a-chip devices.
* Published with Nature Springer in Light: Science and Applications
Spectral DiffuserCam: lensless snapshot hyperspectral imaging with a spectral filter array
Jun 15, 2020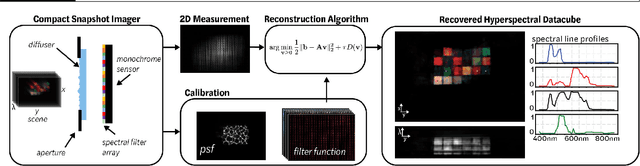
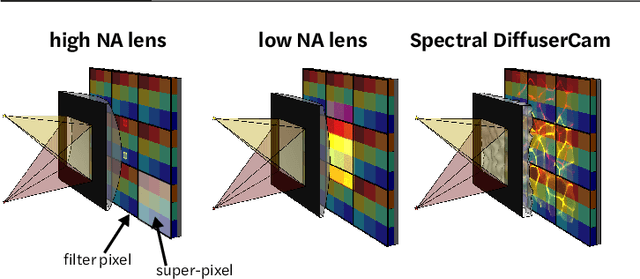
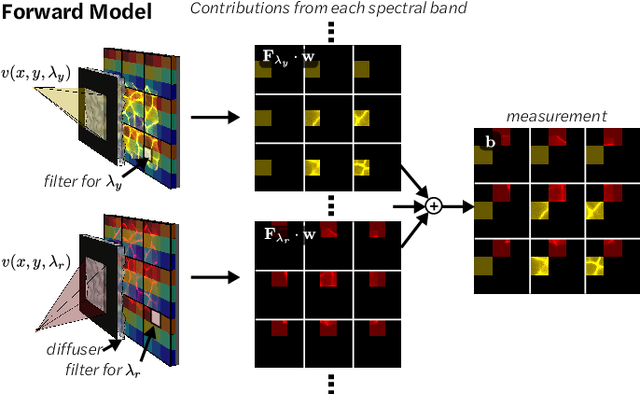
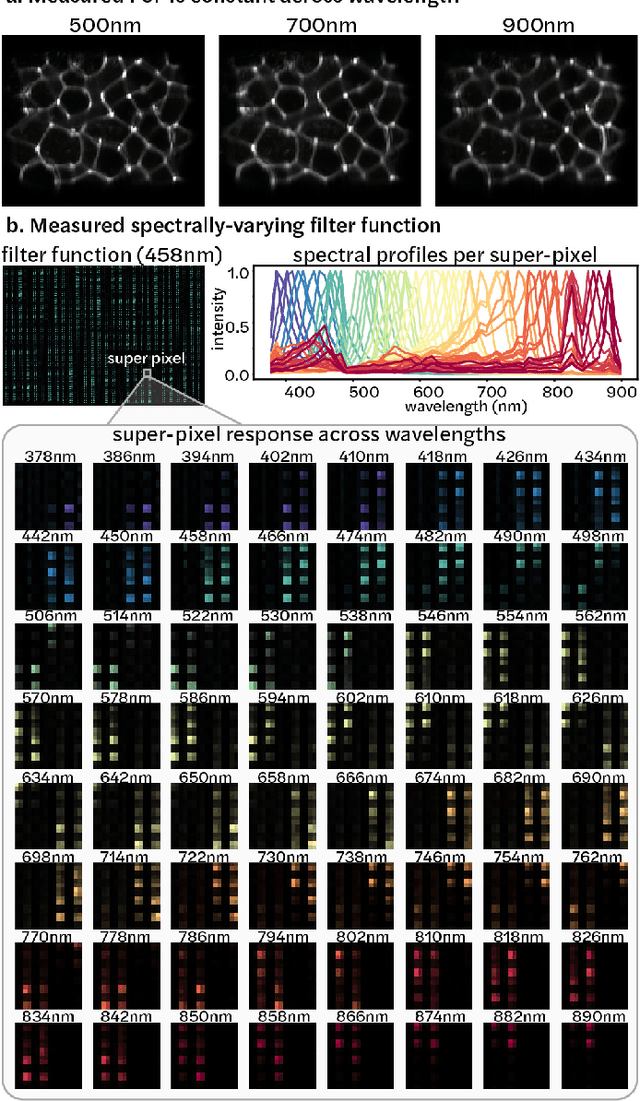
Abstract:Hyperspectral imaging is useful for applications ranging from medical diagnostics to crop monitoring; however, traditional scanning hyperspectral imagers are prohibitively slow and expensive for widespread adoption. Snapshot techniques exist but are often confined to bulky benchtop setups or have low spatio-spectral resolution. In this paper, we propose a novel, compact, and inexpensive computational camera for snapshot hyperspectral imaging. Our system consists of a repeated spectral filter array placed directly on the image sensor and a diffuser placed close to the sensor. Each point in the world maps to a unique pseudorandom pattern on the spectral filter array, which encodes multiplexed spatio-spectral information. A sparsity-constrained inverse problem solver then recovers the hyperspectral volume with good spatio-spectral resolution. By using a spectral filter array, our hyperspectral imaging framework is flexible and can be designed with contiguous or non-contiguous spectral filters that can be chosen for a given application. We provide theory for system design, demonstrate a prototype device, and present experimental results with high spatio-spectral resolution.
Learned reconstructions for practical mask-based lensless imaging
Aug 30, 2019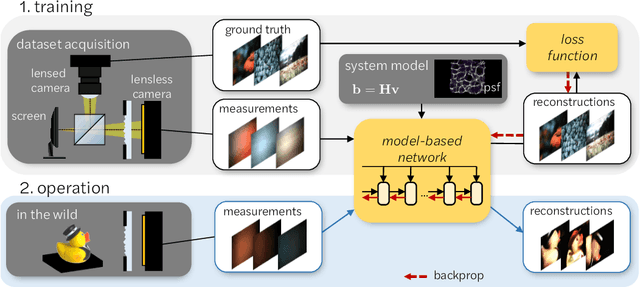
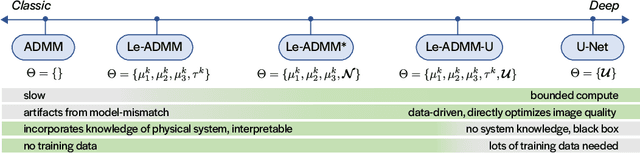
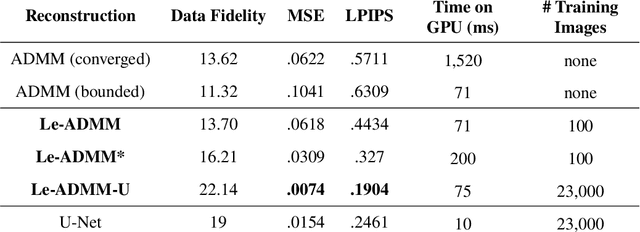
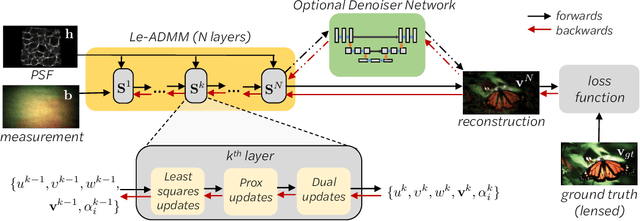
Abstract:Mask-based lensless imagers are smaller and lighter than traditional lensed cameras. In these imagers, the sensor does not directly record an image of the scene; rather, a computational algorithm reconstructs it. Typically, mask-based lensless imagers use a model-based reconstruction approach that suffers from long compute times and a heavy reliance on both system calibration and heuristically chosen denoisers. In this work, we address these limitations using a bounded-compute, trainable neural network to reconstruct the image. We leverage our knowledge of the physical system by unrolling a traditional model-based optimization algorithm, whose parameters we optimize using experimentally gathered ground-truth data. Optionally, images produced by the unrolled network are then fed into a jointly-trained denoiser. As compared to traditional methods, our architecture achieves better perceptual image quality and runs 20x faster, enabling interactive previewing of the scene. We explore a spectrum between model-based and deep learning methods, showing the benefits of using an intermediate approach. Finally, we test our network on images taken in the wild with a prototype mask-based camera, demonstrating that our network generalizes to natural images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge