"Image": models, code, and papers
Quantitative Error Prediction of Medical Image Registration using Regression Forests
May 18, 2019
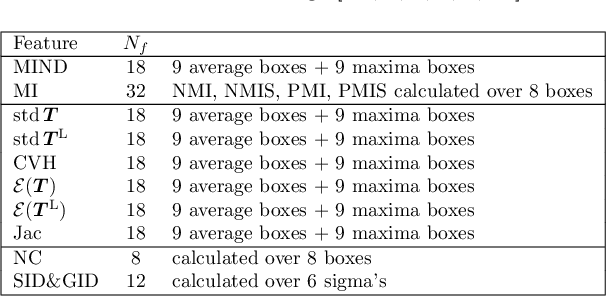
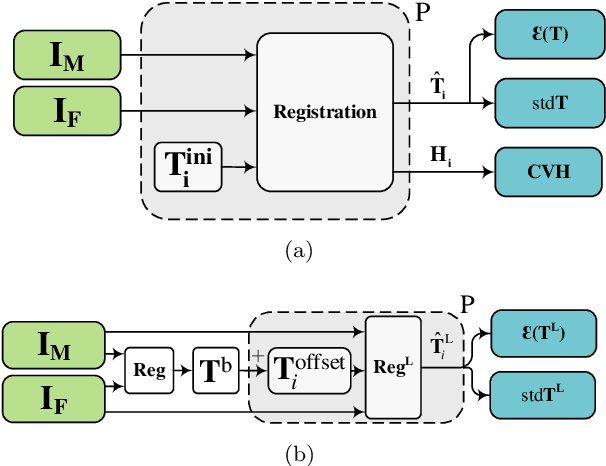
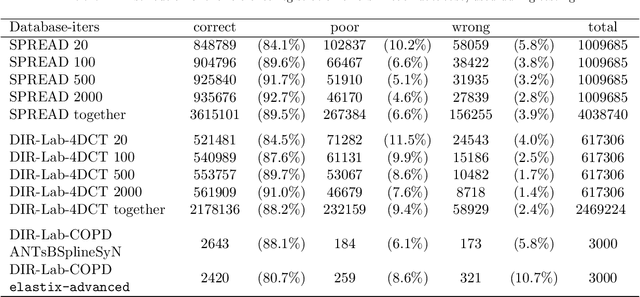
Predicting registration error can be useful for evaluation of registration procedures, which is important for the adoption of registration techniques in the clinic. In addition, quantitative error prediction can be helpful in improving the registration quality. The task of predicting registration error is demanding due to the lack of a ground truth in medical images. This paper proposes a new automatic method to predict the registration error in a quantitative manner, and is applied to chest CT scans. A random regression forest is utilized to predict the registration error locally. The forest is built with features related to the transformation model and features related to the dissimilarity after registration. The forest is trained and tested using manually annotated corresponding points between pairs of chest CT scans in two experiments: SPREAD (trained and tested on SPREAD) and inter-database (including three databases SPREAD, DIR-Lab-4DCT and DIR-Lab-COPDgene). The results show that the mean absolute errors of regression are 1.07 $\pm$ 1.86 and 1.76 $\pm$ 2.59 mm for the SPREAD and inter-database experiment, respectively. The overall accuracy of classification in three classes (correct, poor and wrong registration) is 90.7% and 75.4%, for SPREAD and inter-database respectively. The good performance of the proposed method enables important applications such as automatic quality control in large-scale image analysis.
Fusion of Real Time Thermal Image and 1D/2D/3D Depth Laser Readings for Remote Thermal Sensing in Industrial Plants by Means of UAVs and/or Robots
Jun 04, 2020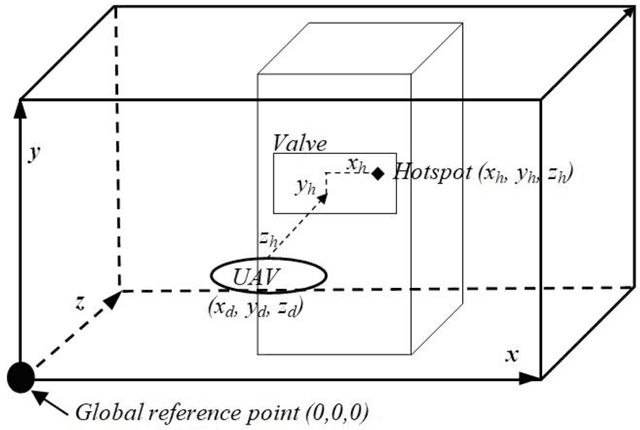
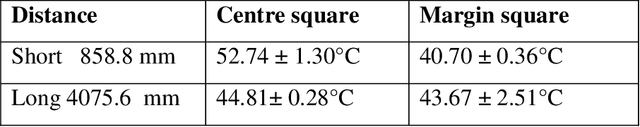
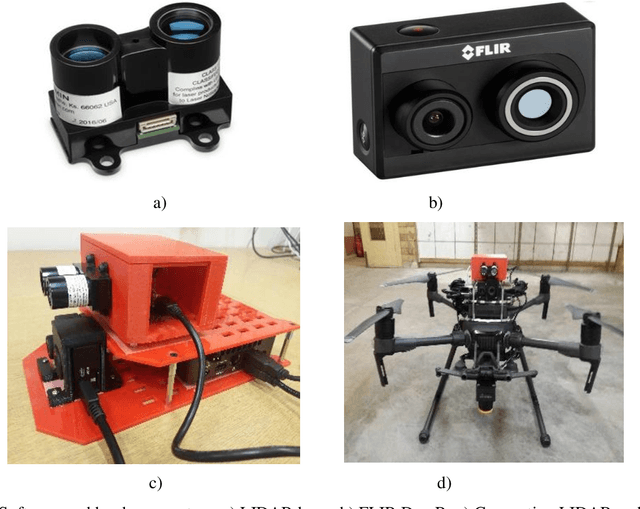


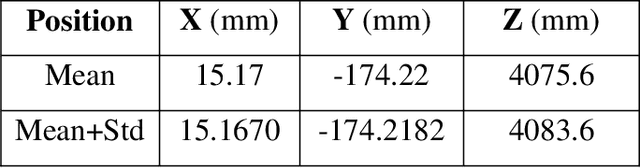
This paper presents fast procedures for thermal infrared remote sensing in dark, GPS-denied environments, such as those found in industrial plants such as in High-Voltage Direct Current (HVDC) converter stations. These procedures are based on the combination of the depth estimation obtained from either a 1-Dimensional LIDAR laser or a 2-Dimensional Hokuyo laser or a 3D MultiSense SLB laser sensor and the visible and thermal cameras from a FLIR Duo R dual-sensor thermal camera. The combination of these sensors/cameras is suitable to be mounted on Unmanned Aerial Vehicles (UAVs) and/or robots in order to provide reliable information about the potential malfunctions, which can be found within the hazardous environment. For example, the capabilities of the developed software and hardware system corresponding to the combination of the 1-D LIDAR sensor and the FLIR Duo R dual-sensor thermal camera is assessed from the point of the accuracy of results and the required computational times: the obtained computational times are under 10 ms, with a maximum localization error of 8 mm and an average standard deviation for the measured temperatures of 1.11 degree Celsius, which results are obtained for a number of test cases. The paper is structured as follows: the description of the system used for identification and localization of hotspots in industrial plants is presented in section II. In section III, the method for faults identification and localization in plants by using a 1-Dimensional LIDAR laser sensor and thermal images is described together with results. In section IV the real time thermal image processing is presented. Fusion of the 2-Dimensional depth laser Hokuyo and the thermal images is described in section V. In section VI the combination of the 3D MultiSense SLB laser and thermal images is described. In section VII a discussion and several conclusions are drawn.
Universal Joint Image Clustering and Registration using Partition Information
Nov 30, 2017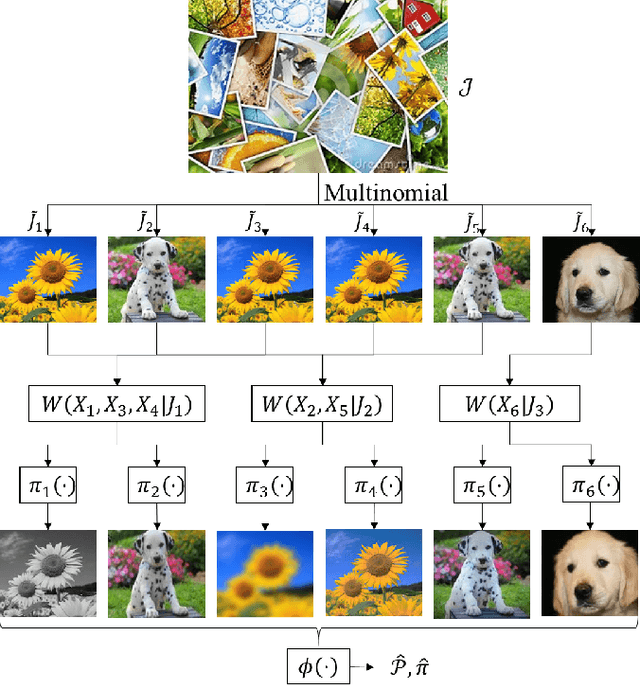
We consider the problem of universal joint clustering and registration of images and define algorithms using multivariate information functionals. We first study registering two images using maximum mutual information and prove its asymptotic optimality. We then show the shortcomings of pairwise registration in multi-image registration, and design an asymptotically optimal algorithm based on multiinformation. Further, we define a novel multivariate information functional to perform joint clustering and registration of images, and prove consistency of the algorithm. Finally, we consider registration and clustering of numerous limited-resolution images, defining algorithms that are order-optimal in scaling of number of pixels in each image with the number of images.
How Much Off-The-Shelf Knowledge Is Transferable From Natural Images To Pathology Images?
May 09, 2020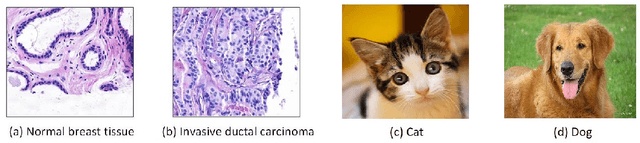
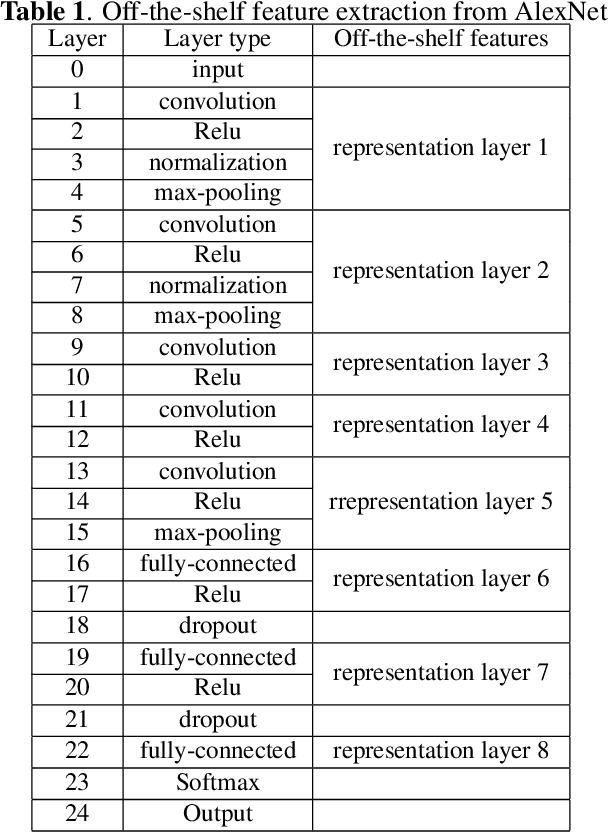
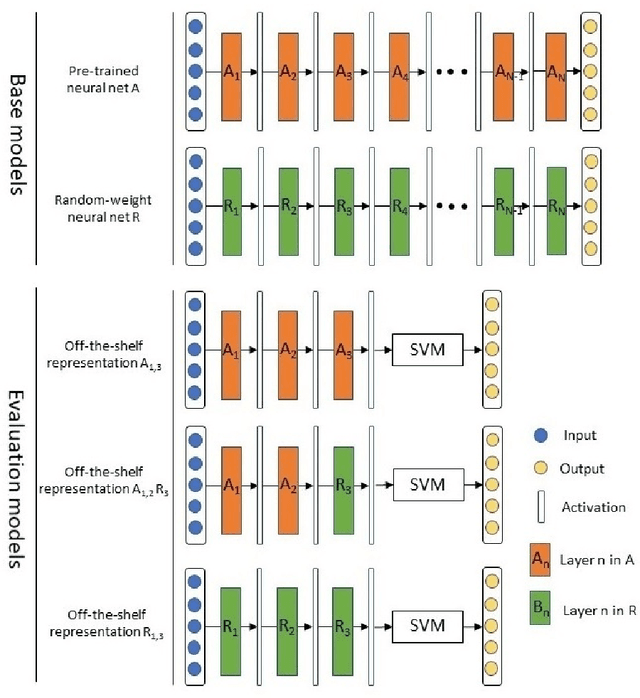
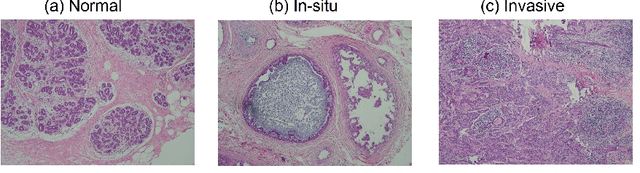
Deep learning has achieved a great success in natural image classification. To overcome data-scarcity in computational pathology, recent studies exploit transfer learning to reuse knowledge gained from natural images in pathology image analysis, aiming to build effective pathology image diagnosis models. Since transferability of knowledge heavily depends on the similarity of the original and target tasks, significant differences in image content and statistics between pathology images and natural images raise the questions: how much knowledge is transferable? Is the transferred information equally contributed by pre-trained layers? To answer these questions, this paper proposes a framework to quantify knowledge gain by a particular layer, conducts an empirical investigation in pathology image centered transfer learning, and reports some interesting observations. Particularly, compared to the performance baseline obtained by random-weight model, though transferability of off-the-shelf representations from deep layers heavily depend on specific pathology image sets, the general representation generated by early layers does convey transferred knowledge in various image classification applications. The observation in this study encourages further investigation of specific metric and tools to quantify effectiveness and feasibility of transfer learning in future.
Anonymization of labeled TOF-MRA images for brain vessel segmentation using generative adversarial networks
Sep 09, 2020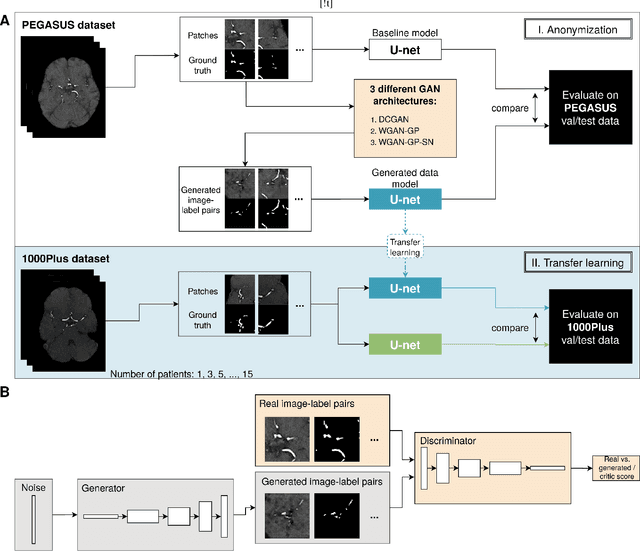
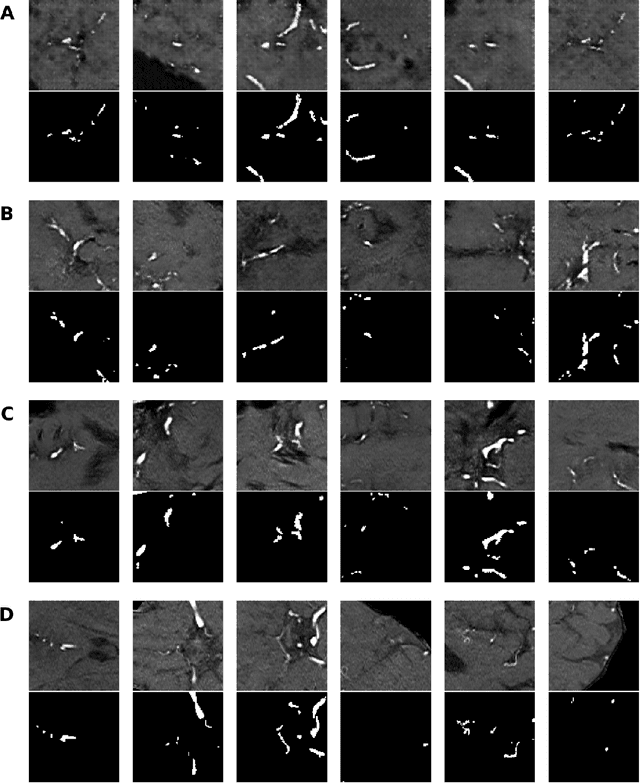
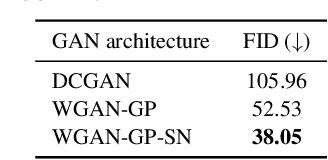

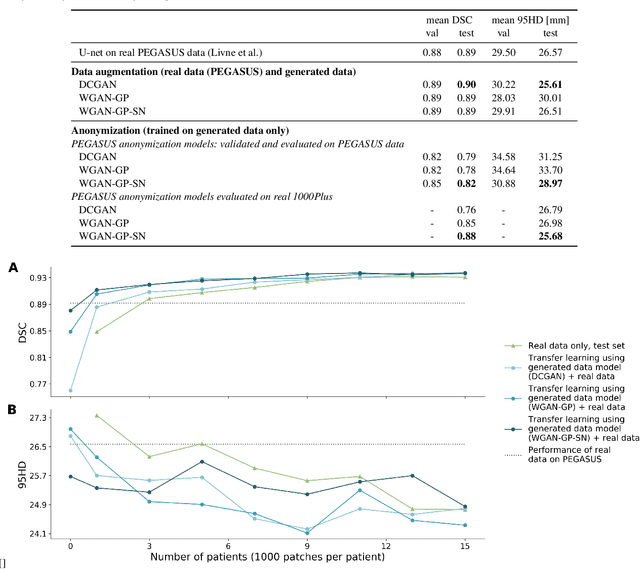
Anonymization and data sharing are crucial for privacy protection and acquisition of large datasets for medical image analysis. This is a big challenge, especially for neuroimaging. Here, the brain's unique structure allows for re-identification and thus requires non-conventional anonymization. Generative adversarial networks (GANs) have the potential to provide anonymous images while preserving predictive properties. Analyzing brain vessel segmentation as a use case, we trained 3 GANs on time-of-flight (TOF) magnetic resonance angiography (MRA) patches for image-label generation: 1) Deep convolutional GAN, 2) Wasserstein-GAN with gradient penalty (WGAN-GP) and 3) WGAN-GP with spectral normalization (WGAN-GP-SN). The generated image-labels from each GAN were used to train a U-net for segmentation and tested on real data. Moreover, we applied our synthetic patches using transfer learning on a second dataset. For an increasing number of up to 15 patients we evaluated the model performance on real data with and without pre-training. The performance for all models was assessed by the Dice Similarity Coefficient (DSC) and the 95th percentile of the Hausdorff Distance (95HD). Comparing the 3 GANs, the U-net trained on synthetic data generated by the WGAN-GP-SN showed the highest performance to predict vessels (DSC/95HD 0.82/28.97) benchmarked by the U-net trained on real data (0.89/26.61). The transfer learning approach showed superior performance for the same GAN compared to no pre-training, especially for one patient only (0.91/25.68 vs. 0.85/27.36). In this work, synthetic image-label pairs retained generalizable information and showed good performance for vessel segmentation. Besides, we showed that synthetic patches can be used in a transfer learning approach with independent data. This paves the way to overcome the challenges of scarce data and anonymization in medical imaging.
Learning Adaptive Sampling and Reconstruction for Volume Visualization
Jul 20, 2020
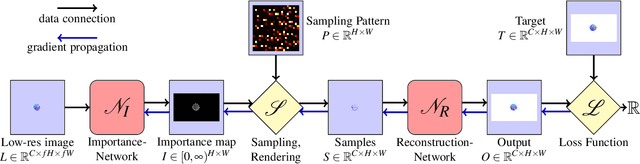
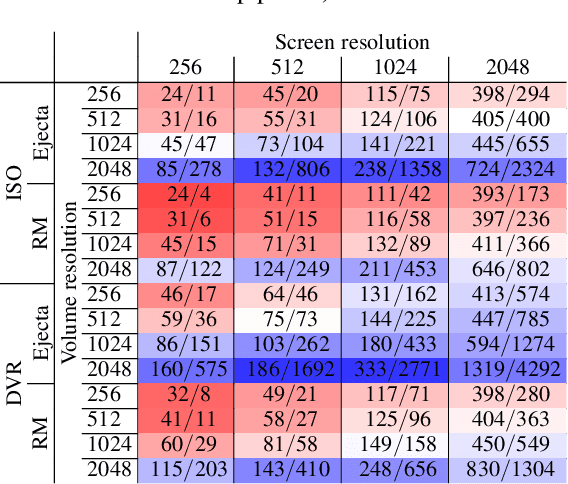
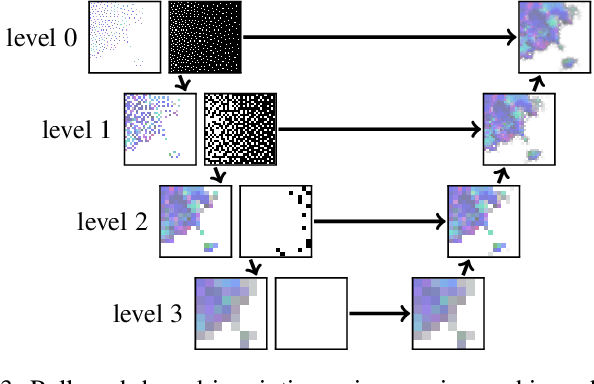
A central challenge in data visualization is to understand which data samples are required to generate an image of a data set in which the relevant information is encoded. In this work, we make a first step towards answering the question of whether an artificial neural network can predict where to sample the data with higher or lower density, by learning of correspondences between the data, the sampling patterns and the generated images. We introduce a novel neural rendering pipeline, which is trained end-to-end to generate a sparse adaptive sampling structure from a given low-resolution input image, and reconstructs a high-resolution image from the sparse set of samples. For the first time, to the best of our knowledge, we demonstrate that the selection of structures that are relevant for the final visual representation can be jointly learned together with the reconstruction of this representation from these structures. Therefore, we introduce differentiable sampling and reconstruction stages, which can leverage back-propagation based on supervised losses solely on the final image. We shed light on the adaptive sampling patterns generated by the network pipeline and analyze its use for volume visualization including isosurface and direct volume rendering.
Self-Supervised Learning of Graph Neural Networks: A Unified Review
Feb 23, 2021
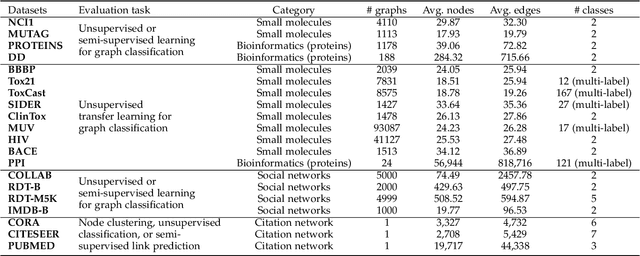
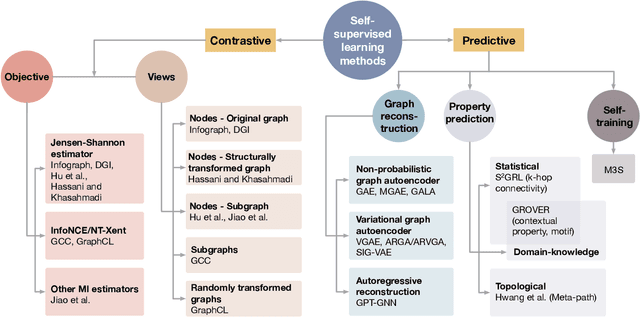
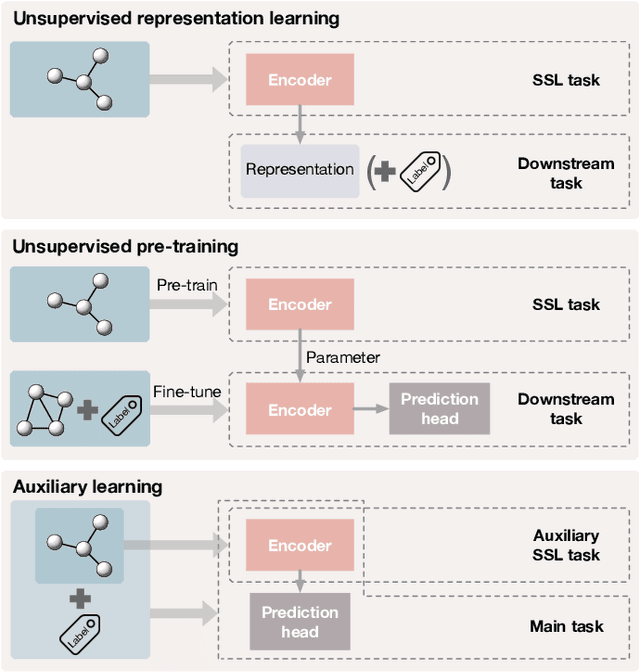
Deep models trained in supervised mode have achieved remarkable success on a variety of tasks. When labeled samples are limited, self-supervised learning (SSL) is emerging as a new paradigm for making use of large amounts of unlabeled samples. SSL has achieved promising performance on natural language and image learning tasks. Recently, there is a trend to extend such success to graph data using graph neural networks (GNNs). In this survey, we provide a unified review of different ways of training GNNs using SSL. Specifically, we categorize SSL methods into contrastive and predictive models. In either category, we provide a unified framework for methods as well as how these methods differ in each component under the framework. Our unified treatment of SSL methods for GNNs sheds light on the similarities and differences of various methods, setting the stage for developing new methods and algorithms. We also summarize different SSL settings and the corresponding datasets used in each setting. To facilitate methodological development and empirical comparison, we develop a standardized testbed for SSL in GNNs, including implementations of common baseline methods, datasets, and evaluation metrics.
Do Noises Bother Human and Neural Networks In the Same Way? A Medical Image Analysis Perspective
Nov 04, 2020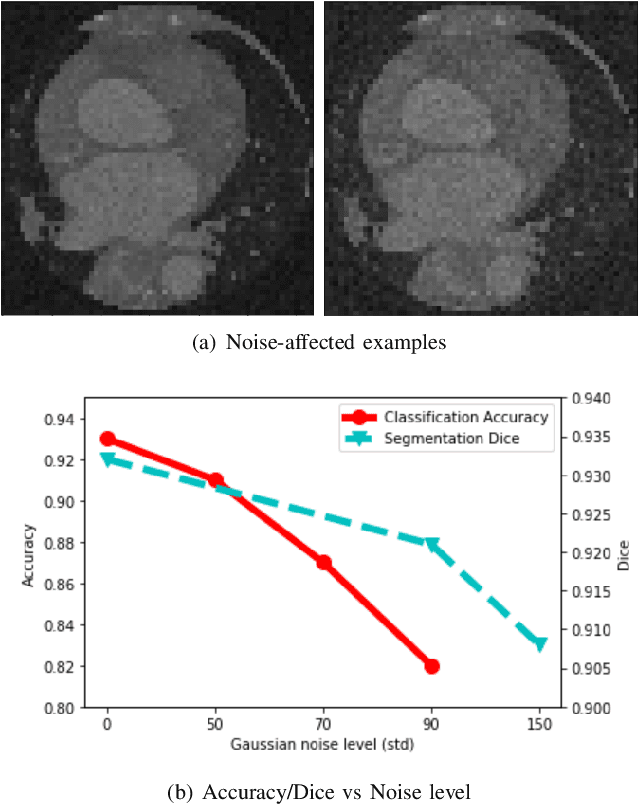
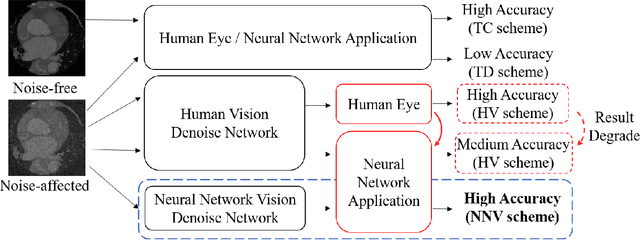
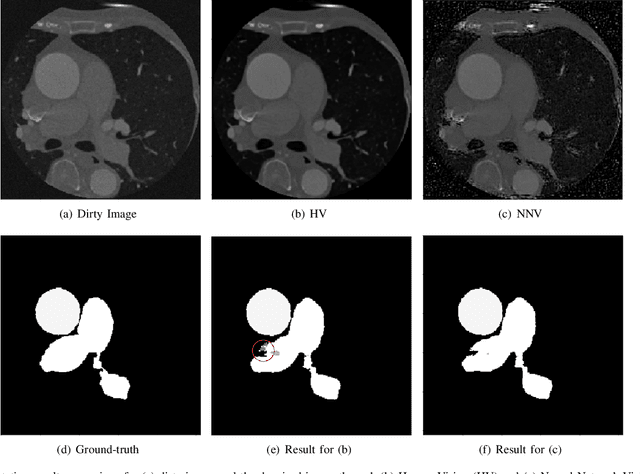
Deep learning had already demonstrated its power in medical images, including denoising, classification, segmentation, etc. All these applications are proposed to automatically analyze medical images beforehand, which brings more information to radiologists during clinical assessment for accuracy improvement. Recently, many medical denoising methods had shown their significant artifact reduction result and noise removal both quantitatively and qualitatively. However, those existing methods are developed around human-vision, i.e., they are designed to minimize the noise effect that can be perceived by human eyes. In this paper, we introduce an application-guided denoising framework, which focuses on denoising for the following neural networks. In our experiments, we apply the proposed framework to different datasets, models, and use cases. Experimental results show that our proposed framework can achieve a better result than human-vision denoising network.
A Haar Wavelet-Based Perceptual Similarity Index for Image Quality Assessment
Nov 06, 2017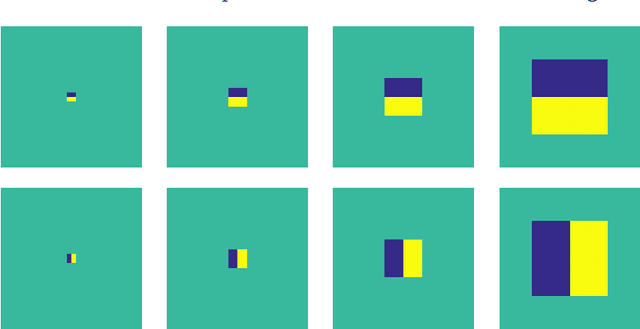
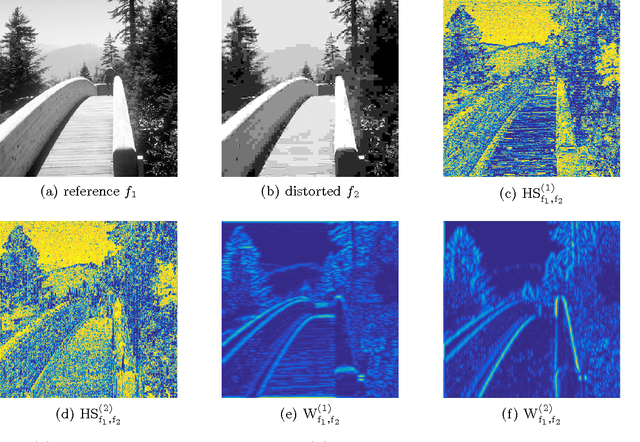
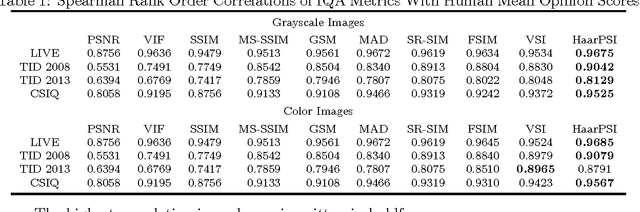

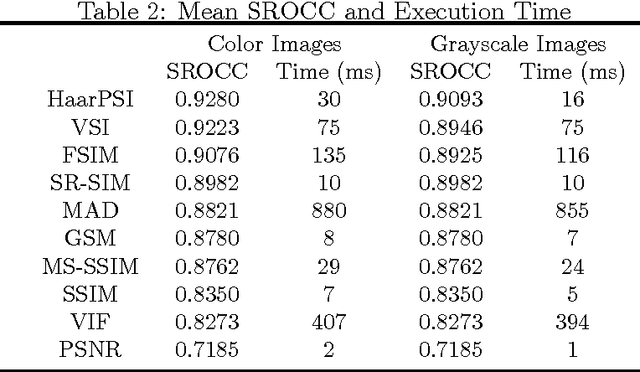
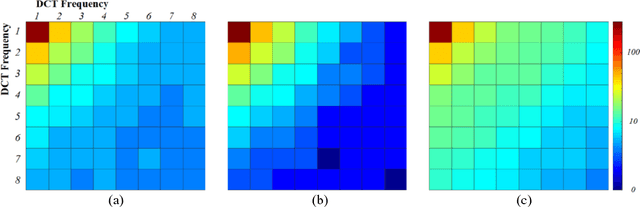
In most practical situations, the compression or transmission of images and videos creates distortions that will eventually be perceived by a human observer. Vice versa, image and video restoration techniques, such as inpainting or denoising, aim to enhance the quality of experience of human viewers. Correctly assessing the similarity between an image and an undistorted reference image as subjectively experienced by a human viewer can thus lead to significant improvements in any transmission, compression, or restoration system. This paper introduces the Haar wavelet-based perceptual similarity index (HaarPSI), a novel and computationally inexpensive similarity measure for full reference image quality assessment. The HaarPSI utilizes the coefficients obtained from a Haar wavelet decomposition to assess local similarities between two images, as well as the relative importance of image areas. The consistency of the HaarPSI with the human quality of experience was validated on four large benchmark databases containing thousands of differently distorted images. On these databases, the HaarPSI achieves higher correlations with human opinion scores than state-of-the-art full reference similarity measures like the structural similarity index (SSIM), the feature similarity index (FSIM), and the visual saliency-based index (VSI). Along with the simple computational structure and the short execution time, these experimental results suggest a high applicability of the HaarPSI in real world tasks.
Quality-aware semi-supervised learning for CMR segmentation
Sep 01, 2020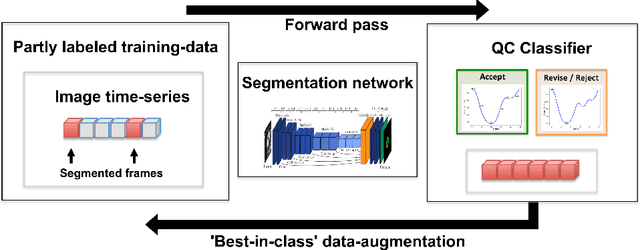
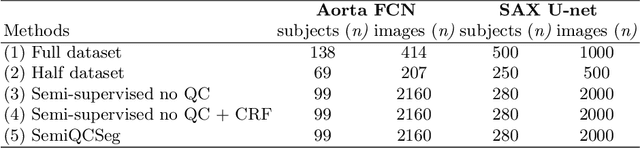
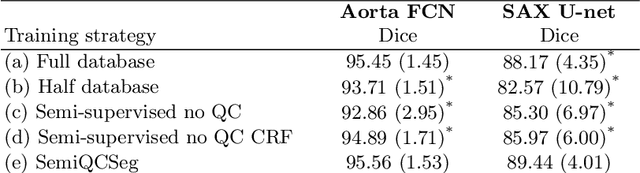
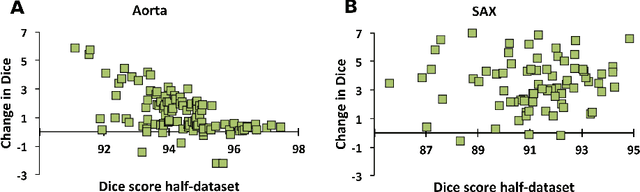
One of the challenges in developing deep learning algorithms for medical image segmentation is the scarcity of annotated training data. To overcome this limitation, data augmentation and semi-supervised learning (SSL) methods have been developed. However, these methods have limited effectiveness as they either exploit the existing data set only (data augmentation) or risk negative impact by adding poor training examples (SSL). Segmentations are rarely the final product of medical image analysis - they are typically used in downstream tasks to infer higher-order patterns to evaluate diseases. Clinicians take into account a wealth of prior knowledge on biophysics and physiology when evaluating image analysis results. We have used these clinical assessments in previous works to create robust quality-control (QC) classifiers for automated cardiac magnetic resonance (CMR) analysis. In this paper, we propose a novel scheme that uses QC of the downstream task to identify high quality outputs of CMR segmentation networks, that are subsequently utilised for further network training. In essence, this provides quality-aware augmentation of training data in a variant of SSL for segmentation networks (semiQCSeg). We evaluate our approach in two CMR segmentation tasks (aortic and short axis cardiac volume segmentation) using UK Biobank data and two commonly used network architectures (U-net and a Fully Convolutional Network) and compare against supervised and SSL strategies. We show that semiQCSeg improves training of the segmentation networks. It decreases the need for labelled data, while outperforming the other methods in terms of Dice and clinical metrics. SemiQCSeg can be an efficient approach for training segmentation networks for medical image data when labelled datasets are scarce.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge