Marius Staring
Clinical DVH metrics as a loss function for 3D dose prediction in head and neck radiotherapy
Mar 31, 2026Abstract:Purpose: Deep-learning-based three-dimensional (3D) dose prediction is widely used in automated radiotherapy workflows. However, most existing models are trained with voxel-wise regression losses, which are poorly aligned with clinical plan evaluation criteria based on dose-volume histogram (DVH) metrics. This study aims to develop a clinically guided loss formulation that directly optimizes clinically used DVH metrics while remaining computationally efficient for head and neck (H\&N) dose prediction. Methods: We propose a clinical DVH metric loss (CDM loss) that incorporates differentiable \textit{D-metrics} and surrogate \textit{V-metrics}, together with a lossless bit-mask region-of-interest (ROI) encoding to improve training efficiency. The method was evaluated on 174 H\&N patients using a temporal split (137 training, 37 testing). Results: Compared with MAE- and DVH-curve based losses, CDM loss substantially improved target coverage and satisfied all clinical constraints. Using a standard 3D U-Net, the PTV Score was reduced from 1.544 (MAE) to 0.491 (MAE + CDM), while OAR sparing remained comparable. Bit-mask encoding reduced training time by 83\% and lowered GPU memory usage. Conclusion: Directly optimizing clinically used DVH metrics enables 3D dose predictions that are better aligned with clinical treatment planning criteria than conventional voxel-wise or DVH-curve-based supervision. The proposed CDM loss, combined with efficient ROI bit-mask encoding, provides a practical and scalable framework for H\&N dose prediction.
CRUNet-MR-Univ: A Foundation Model for Diverse Cardiac MRI Reconstruction
Jan 07, 2026Abstract:In recent years, deep learning has attracted increasing at- tention in the field of Cardiac MRI (CMR) reconstruction due to its superior performance over traditional methods, particularly in handling higher acceleration factors, highlighting its potential for real-world clini- cal applications. However, current deep learning methods remain limited in generalizability. CMR scans exhibit wide variability in image contrast, sampling patterns, scanner vendors, anatomical structures, and disease types. Most existing models are designed to handle only a single or nar- row subset of these variations, leading to performance degradation when faced with distribution shifts. Therefore, it is beneficial to develop a unified model capable of generalizing across diverse CMR scenarios. To this end, we propose CRUNet-MR-Univ, a foundation model that lever- ages spatio-temporal correlations and prompt-based priors to effectively handle the full diversity of CMR scans. Our approach consistently out- performs baseline methods across a wide range of settings, highlighting its effectiveness and promise.
Efficient Large-Deformation Medical Image Registration via Recurrent Dynamic Correlation
Oct 25, 2025Abstract:Deformable image registration estimates voxel-wise correspondences between images through spatial transformations, and plays a key role in medical imaging. While deep learning methods have significantly reduced runtime, efficiently handling large deformations remains a challenging task. Convolutional networks aggregate local features but lack direct modeling of voxel correspondences, promoting recent works to explore explicit feature matching. Among them, voxel-to-region matching is more efficient for direct correspondence modeling by computing local correlation features whithin neighbourhoods, while region-to-region matching incurs higher redundancy due to excessive correlation pairs across large regions. However, the inherent locality of voxel-to-region matching hinders the capture of long-range correspondences required for large deformations. To address this, we propose a Recurrent Correlation-based framework that dynamically relocates the matching region toward more promising positions. At each step, local matching is performed with low cost, and the estimated offset guides the next search region, supporting efficient convergence toward large deformations. In addition, we uses a lightweight recurrent update module with memory capacity and decouples motion-related and texture features to suppress semantic redundancy. We conduct extensive experiments on brain MRI and abdominal CT datasets under two settings: with and without affine pre-registration. Results show that our method exibits a strong accuracy-computation trade-off, surpassing or matching the state-of-the-art performance. For example, it achieves comparable performance on the non-affine OASIS dataset, while using only 9.5% of the FLOPs and running 96% faster than RDP, a representative high-performing method.
CMRINet: Joint Groupwise Registration and Segmentation for Cardiac Function Quantification from Cine-MRI
May 22, 2025



Abstract:Accurate and efficient quantification of cardiac function is essential for the estimation of prognosis of cardiovascular diseases (CVDs). One of the most commonly used metrics for evaluating cardiac pumping performance is left ventricular ejection fraction (LVEF). However, LVEF can be affected by factors such as inter-observer variability and varying pre-load and after-load conditions, which can reduce its reproducibility. Additionally, cardiac dysfunction may not always manifest as alterations in LVEF, such as in heart failure and cardiotoxicity diseases. An alternative measure that can provide a relatively load-independent quantitative assessment of myocardial contractility is myocardial strain and strain rate. By using LVEF in combination with myocardial strain, it is possible to obtain a thorough description of cardiac function. Automated estimation of LVEF and other volumetric measures from cine-MRI sequences can be achieved through segmentation models, while strain calculation requires the estimation of tissue displacement between sequential frames, which can be accomplished using registration models. These tasks are often performed separately, potentially limiting the assessment of cardiac function. To address this issue, in this study we propose an end-to-end deep learning (DL) model that jointly estimates groupwise (GW) registration and segmentation for cardiac cine-MRI images. The proposed anatomically-guided Deep GW network was trained and validated on a large dataset of 4-chamber view cine-MRI image series of 374 subjects. A quantitative comparison with conventional GW registration using elastix and two DL-based methods showed that the proposed model improved performance and substantially reduced computation time.
UPCMR: A Universal Prompt-guided Model for Random Sampling Cardiac MRI Reconstruction
Feb 18, 2025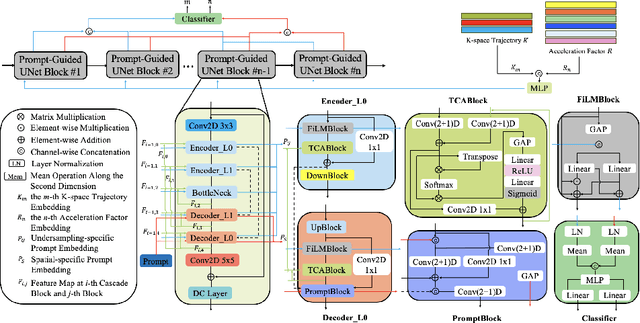
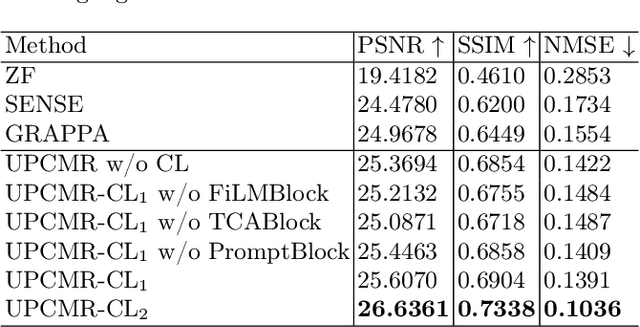
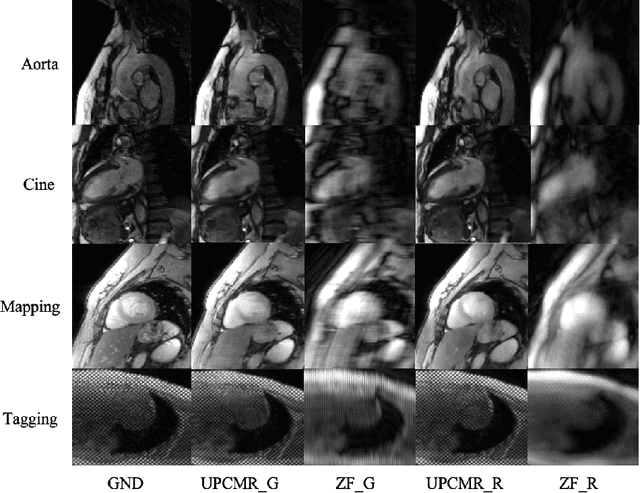
Abstract:Cardiac magnetic resonance imaging (CMR) is vital for diagnosing heart diseases, but long scan time remains a major drawback. To address this, accelerated imaging techniques have been introduced by undersampling k-space, which reduces the quality of the resulting images. Recent deep learning advancements aim to speed up scanning while preserving quality, but adapting to various sampling modes and undersampling factors remains challenging. Therefore, building a universal model is a promising direction. In this work, we introduce UPCMR, a universal unrolled model designed for CMR reconstruction. This model incorporates two kinds of learnable prompts, undersampling-specific prompt and spatial-specific prompt, and integrates them with a UNet structure in each block. Overall, by using the CMRxRecon2024 challenge dataset for training and validation, the UPCMR model highly enhances reconstructed image quality across all random sampling scenarios through an effective training strategy compared to some traditional methods, demonstrating strong adaptability potential for this task.
Efficient MedSAMs: Segment Anything in Medical Images on Laptop
Dec 20, 2024


Abstract:Promptable segmentation foundation models have emerged as a transformative approach to addressing the diverse needs in medical images, but most existing models require expensive computing, posing a big barrier to their adoption in clinical practice. In this work, we organized the first international competition dedicated to promptable medical image segmentation, featuring a large-scale dataset spanning nine common imaging modalities from over 20 different institutions. The top teams developed lightweight segmentation foundation models and implemented an efficient inference pipeline that substantially reduced computational requirements while maintaining state-of-the-art segmentation accuracy. Moreover, the post-challenge phase advanced the algorithms through the design of performance booster and reproducibility tasks, resulting in improved algorithms and validated reproducibility of the winning solution. Furthermore, the best-performing algorithms have been incorporated into the open-source software with a user-friendly interface to facilitate clinical adoption. The data and code are publicly available to foster the further development of medical image segmentation foundation models and pave the way for impactful real-world applications.
MCP-MedSAM: A Powerful Lightweight Medical Segment Anything Model Trained with a Single GPU in Just One Day
Dec 08, 2024



Abstract:Medical image segmentation involves partitioning medical images into meaningful regions, with a focus on identifying anatomical structures or abnormalities. It has broad applications in healthcare, and deep learning methods have enabled significant advancements in automating this process. Recently, the introduction of the Segmentation Anything Model (SAM), the first foundation model for segmentation task, has prompted researchers to adapt it for the medical domain to improve performance across various tasks. However, SAM's large model size and high GPU requirements hinder its scalability and development in the medical domain. To address these challenges, research has increasingly focused on lightweight adaptations of SAM to reduce its parameter count, enabling training with limited GPU resources while maintaining competitive segmentation performance. In this work, we propose MCP-MedSAM, a powerful and lightweight medical SAM model designed to be trainable on a single GPU within one day while delivering superior segmentation performance. Our method was trained and evaluated using a large-scale challenge dataset\footnote{\url{https://www.codabench.org/competitions/1847}\label{comp}}, compared to top-ranking methods on the challenge leaderboard, MCP-MedSAM achieved superior performance while requiring only one day of training on a single GPU. The code is publicly available at \url{https://github.com/dong845/MCP-MedSAM}.
Swin-LiteMedSAM: A Lightweight Box-Based Segment Anything Model for Large-Scale Medical Image Datasets
Sep 11, 2024Abstract:Medical imaging is essential for the diagnosis and treatment of diseases, with medical image segmentation as a subtask receiving high attention. However, automatic medical image segmentation models are typically task-specific and struggle to handle multiple scenarios, such as different imaging modalities and regions of interest. With the introduction of the Segment Anything Model (SAM), training a universal model for various clinical scenarios has become feasible. Recently, several Medical SAM (MedSAM) methods have been proposed, but these models often rely on heavy image encoders to achieve high performance, which may not be practical for real-world applications due to their high computational demands and slow inference speed. To address this issue, a lightweight version of the MedSAM (LiteMedSAM) can provide a viable solution, achieving high performance while requiring fewer resources and less time. In this work, we introduce Swin-LiteMedSAM, a new variant of LiteMedSAM. This model integrates the tiny Swin Transformer as the image encoder, incorporates multiple types of prompts, including box-based points and scribble generated from a given bounding box, and establishes skip connections between the image encoder and the mask decoder. In the \textit{Segment Anything in Medical Images on Laptop} challenge (CVPR 2024), our approach strikes a good balance between segmentation performance and speed, demonstrating significantly improved overall results across multiple modalities compared to the LiteMedSAM baseline provided by the challenge organizers. Our proposed model achieved a DSC score of \textbf{0.8678} and an NSD score of \textbf{0.8844} on the validation set. On the final test set, it attained a DSC score of \textbf{0.8193} and an NSD score of \textbf{0.8461}, securing fourth place in the challenge.
Improving Uncertainty-Error Correspondence in Deep Bayesian Medical Image Segmentation
Sep 05, 2024



Abstract:Increased usage of automated tools like deep learning in medical image segmentation has alleviated the bottleneck of manual contouring. This has shifted manual labour to quality assessment (QA) of automated contours which involves detecting errors and correcting them. A potential solution to semi-automated QA is to use deep Bayesian uncertainty to recommend potentially erroneous regions, thus reducing time spent on error detection. Previous work has investigated the correspondence between uncertainty and error, however, no work has been done on improving the "utility" of Bayesian uncertainty maps such that it is only present in inaccurate regions and not in the accurate ones. Our work trains the FlipOut model with the Accuracy-vs-Uncertainty (AvU) loss which promotes uncertainty to be present only in inaccurate regions. We apply this method on datasets of two radiotherapy body sites, c.f. head-and-neck CT and prostate MR scans. Uncertainty heatmaps (i.e. predictive entropy) are evaluated against voxel inaccuracies using Receiver Operating Characteristic (ROC) and Precision-Recall (PR) curves. Numerical results show that when compared to the Bayesian baseline the proposed method successfully suppresses uncertainty for accurate voxels, with similar presence of uncertainty for inaccurate voxels. Code to reproduce experiments is available at https://github.com/prerakmody/bayesuncertainty-error-correspondence
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2024:018
Vestibular schwannoma growth prediction from longitudinal MRI by time conditioned neural fields
Apr 04, 2024



Abstract:Vestibular schwannomas (VS) are benign tumors that are generally managed by active surveillance with MRI examination. To further assist clinical decision-making and avoid overtreatment, an accurate prediction of tumor growth based on longitudinal imaging is highly desirable. In this paper, we introduce DeepGrowth, a deep learning method that incorporates neural fields and recurrent neural networks for prospective tumor growth prediction. In the proposed method, each tumor is represented as a signed distance function (SDF) conditioned on a low-dimensional latent code. Unlike previous studies that perform tumor shape prediction directly in the image space, we predict the latent codes instead and then reconstruct future shapes from it. To deal with irregular time intervals, we introduce a time-conditioned recurrent module based on a ConvLSTM and a novel temporal encoding strategy, which enables the proposed model to output varying tumor shapes over time. The experiments on an in-house longitudinal VS dataset showed that the proposed model significantly improved the performance ($\ge 1.6\%$ Dice score and $\ge0.20$ mm 95\% Hausdorff distance), in particular for top 20\% tumors that grow or shrink the most ($\ge 4.6\%$ Dice score and $\ge 0.73$ mm 95\% Hausdorff distance). Our code is available at ~\burl{https://github.com/cyjdswx/DeepGrowth}
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge