"Image": models, code, and papers
Structure-Aware 3D Hourglass Network for Hand Pose Estimation from Single Depth Image
Dec 26, 2018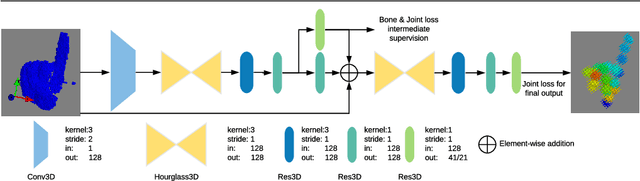

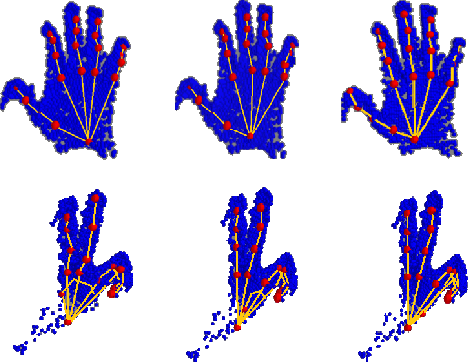
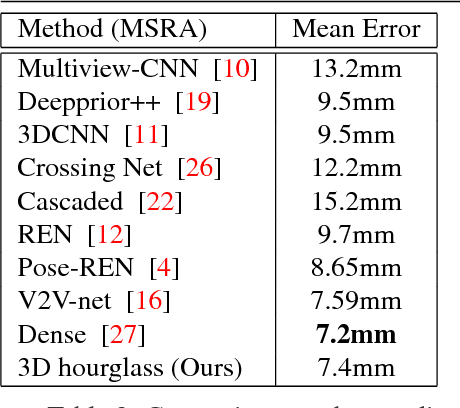
In this paper, we propose a novel structure-aware 3D hourglass network for hand pose estimation from a single depth image, which achieves state-of-the-art results on MSRA and NYU datasets. Compared to existing works that perform image-to-coordination regression, our network takes 3D voxel as input and directly regresses 3D heatmap for each joint. To be specific, we use hourglass network as our backbone network and modify it into 3D form. We explicitly model tree-like finger bone into the network as well as in the loss function in an end-to-end manner, in order to take the skeleton constraints into consideration. Final estimation can then be easily obtained from voxel density map with simple post-processing. Experimental results show that the proposed structure-aware 3D hourglass network is able to achieve a mean joint error of 7.4 mm in MSRA and 8.9 mm in NYU datasets, respectively.
Automated Seizure Detection and Seizure Type Classification From Electroencephalography With a Graph Neural Network and Self-Supervised Pre-Training
Apr 16, 2021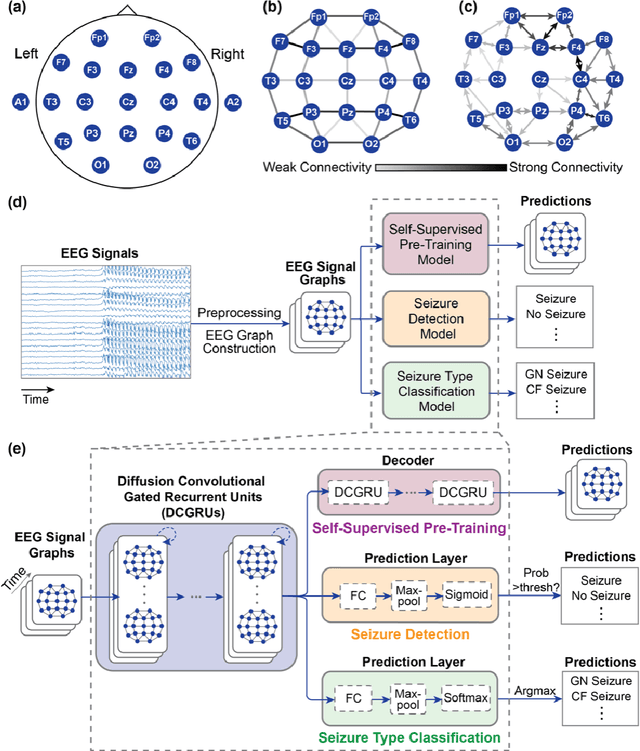
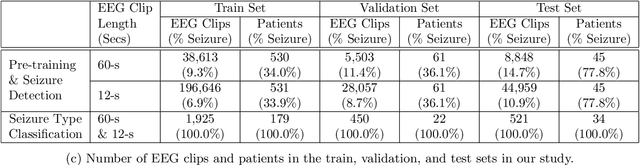
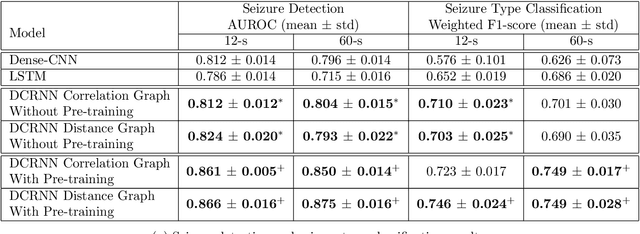
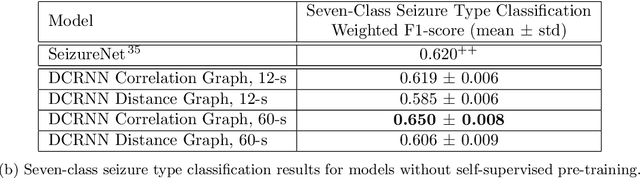
Automated seizure detection and classification from electroencephalography (EEG) can greatly improve the diagnosis and treatment of seizures. While prior studies mainly used convolutional neural networks (CNNs) that assume image-like structure in EEG signals or spectrograms, this modeling choice does not reflect the natural geometry of or connectivity between EEG electrodes. In this study, we propose modeling EEGs as graphs and present a graph neural network for automated seizure detection and classification. In addition, we leverage unlabeled EEG data using a self-supervised pre-training strategy. Our graph model with self-supervised pre-training significantly outperforms previous state-of-the-art CNN and Long Short-Term Memory (LSTM) models by 6.3 points (7.8%) in Area Under the Receiver Operating Characteristic curve (AUROC) for seizure detection and 6.3 points (9.2%) in weighted F1-score for seizure type classification. Ablation studies show that our graph-based modeling approach significantly outperforms existing CNN or LSTM models, and that self-supervision helps further improve the model performance. Moreover, we find that self-supervised pre-training substantially improves model performance on combined tonic seizures, a low-prevalence seizure type. Furthermore, our model interpretability analysis suggests that our model is better at identifying seizure regions compared to an existing CNN. In summary, our graph-based modeling approach integrates domain knowledge about EEG, sets a new state-of-the-art for seizure detection and classification on a large public dataset (5,499 EEG files), and provides better ability to identify seizure regions.
UnrealROX+: An Improved Tool for Acquiring Synthetic Data from Virtual 3D Environments
Apr 23, 2021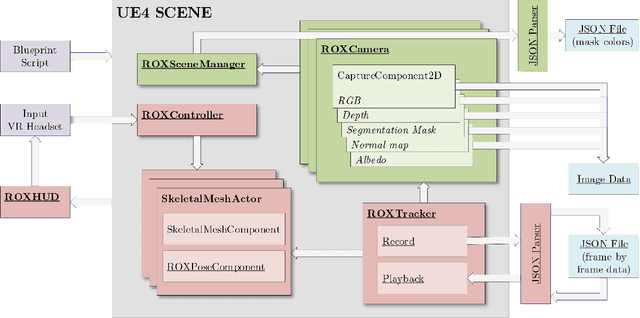
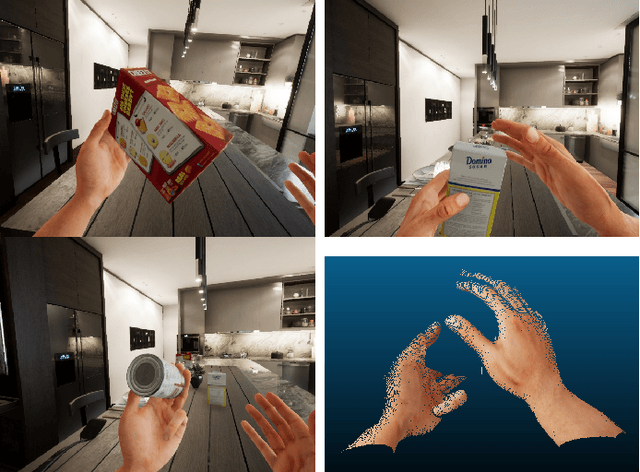



Synthetic data generation has become essential in last years for feeding data-driven algorithms, which surpassed traditional techniques performance in almost every computer vision problem. Gathering and labelling the amount of data needed for these data-hungry models in the real world may become unfeasible and error-prone, while synthetic data give us the possibility of generating huge amounts of data with pixel-perfect annotations. However, most synthetic datasets lack from enough realism in their rendered images. In that context UnrealROX generation tool was presented in 2019, allowing to generate highly realistic data, at high resolutions and framerates, with an efficient pipeline based on Unreal Engine, a cutting-edge videogame engine. UnrealROX enabled robotic vision researchers to generate realistic and visually plausible data with full ground truth for a wide variety of problems such as class and instance semantic segmentation, object detection, depth estimation, visual grasping, and navigation. Nevertheless, its workflow was very tied to generate image sequences from a robotic on-board camera, making hard to generate data for other purposes. In this work, we present UnrealROX+, an improved version of UnrealROX where its decoupled and easy-to-use data acquisition system allows to quickly design and generate data in a much more flexible and customizable way. Moreover, it is packaged as an Unreal plug-in, which makes it more comfortable to use with already existing Unreal projects, and it also includes new features such as generating albedo or a Python API for interacting with the virtual environment from Deep Learning frameworks.
Deep Learning with Inaccurate Training Data for Image Restoration
Nov 18, 2018
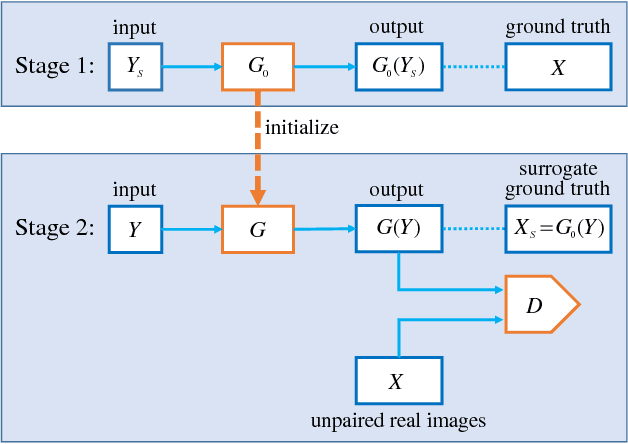


In many applications of deep learning, particularly those in image restoration, it is either very difficult, prohibitively expensive, or outright impossible to obtain paired training data precisely as in the real world. In such cases, one is forced to use synthesized paired data to train the deep convolutional neural network (DCNN). However, due to the unavoidable generalization error in statistical learning, the synthetically trained DCNN often performs poorly on real world data. To overcome this problem, we propose a new general training method that can compensate for, to a large extent, the generalization errors of synthetically trained DCNNs.
Embedded Self-Distillation in Compact Multi-Branch Ensemble Network for Remote Sensing Scene Classification
Apr 01, 2021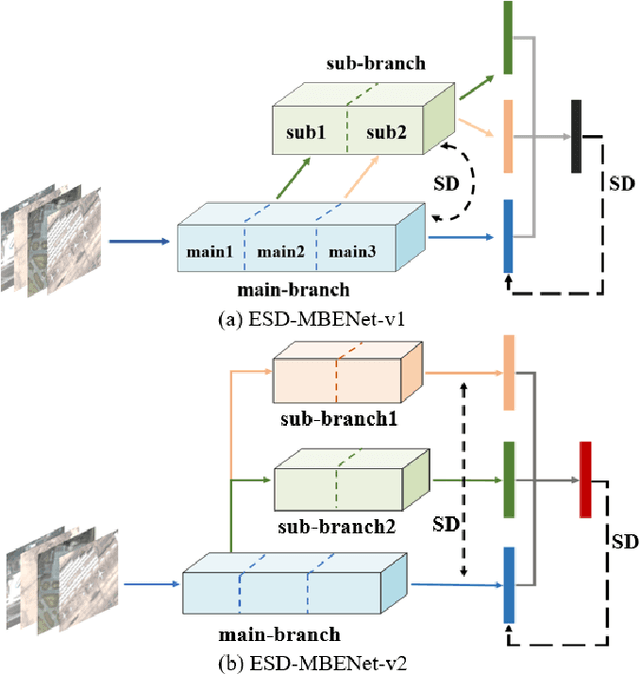
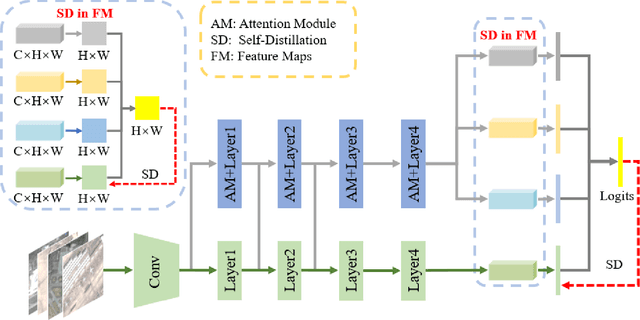
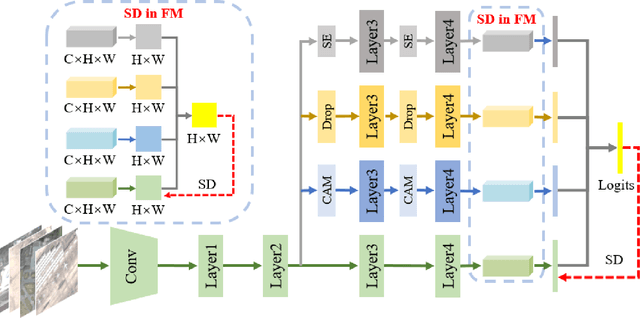
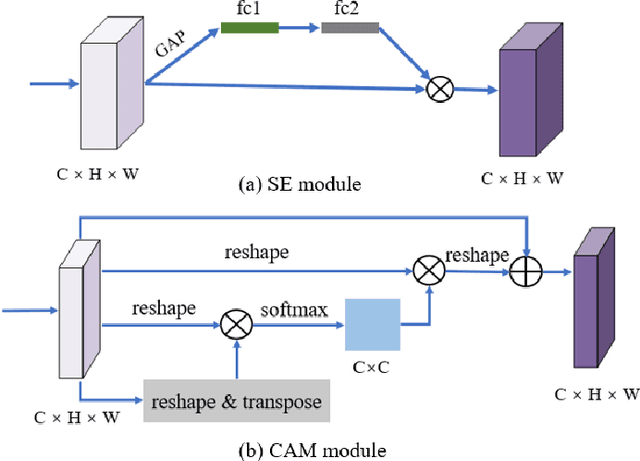
Remote sensing (RS) image scene classification task faces many challenges due to the interference from different characteristics of different geographical elements. To solve this problem, we propose a multi-branch ensemble network to enhance the feature representation ability by fusing features in final output logits and intermediate feature maps. However, simply adding branches will increase the complexity of models and decline the inference efficiency. On this issue, we embed self-distillation (SD) method to transfer knowledge from ensemble network to main-branch in it. Through optimizing with SD, main-branch will have close performance as ensemble network. During inference, we can cut other branches to simplify the whole model. In this paper, we first design compact multi-branch ensemble network, which can be trained in an end-to-end manner. Then, we insert SD method on output logits and feature maps. Compared to previous methods, our proposed architecture (ESD-MBENet) performs strongly on classification accuracy with compact design. Extensive experiments are applied on three benchmark RS datasets AID, NWPU-RESISC45 and UC-Merced with three classic baseline models, VGG16, ResNet50 and DenseNet121. Results prove that our proposed ESD-MBENet can achieve better accuracy than previous state-of-the-art (SOTA) complex models. Moreover, abundant visualization analysis make our method more convincing and interpretable.
Comparison of various image fusion methods for impervious surface classification from VNREDSat-1
May 04, 2018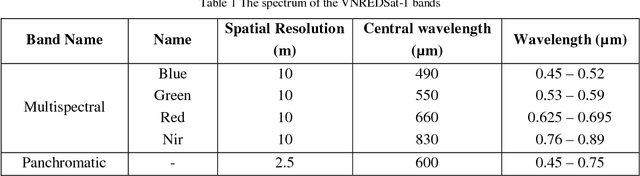
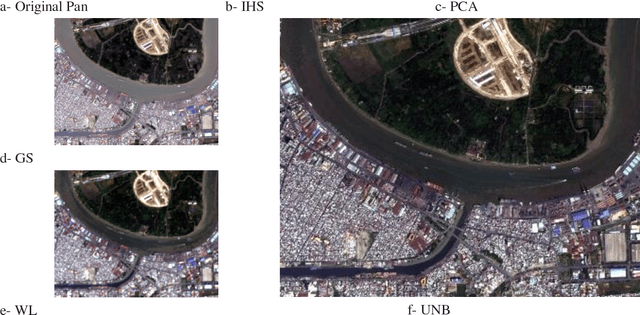

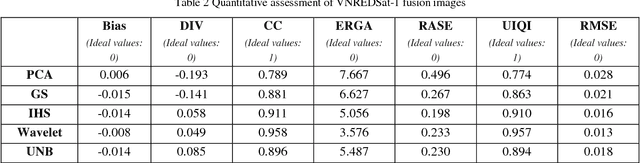
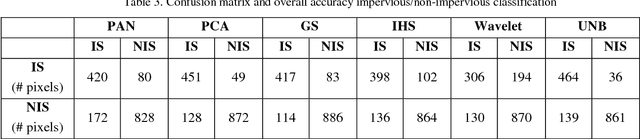
Impervious surface is an important indicator for urban development monitoring. Accurate urban impervious surfaces mapping with VNREDSat-1 remains challenging due to their spectral diversity not captured by individual PAN image. In this artical, five multi-resolution image fusion techniques were compared for classification task of urban impervious surface. The result shows that for VNREDSat-1 dataset, UNB and Wavelet tranform methods are the best techniques reserving spatial and spectral information of original MS image, respectively. However, the UNB technique gives best results when it comes to impervious surface classification especially in the case of shadow area included in non-impervious surface group.
Learning from Subjective Ratings Using Auto-Decoded Deep Latent Embeddings
Apr 16, 2021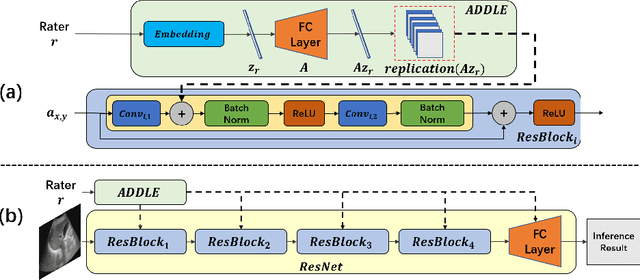
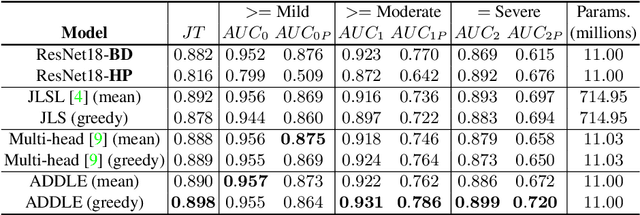
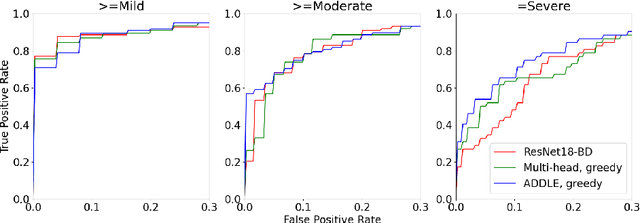
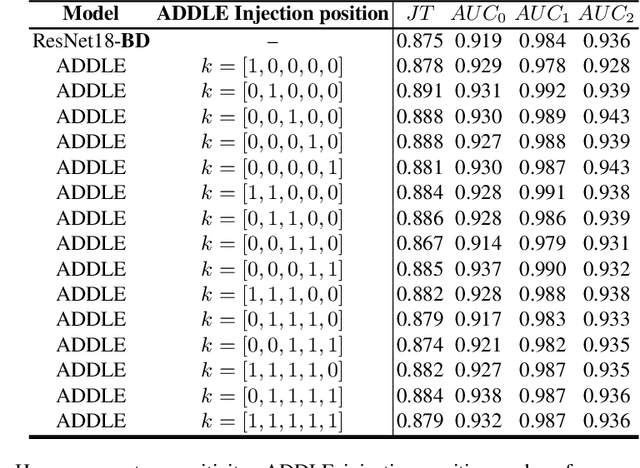
Depending on the application, radiological diagnoses can be associated with high inter- and intra-rater variabilities. Most computer-aided diagnosis (CAD) solutions treat such data as incontrovertible, exposing learning algorithms to considerable and possibly contradictory label noise and biases. Thus, managing subjectivity in labels is a fundamental problem in medical imaging analysis. To address this challenge, we introduce auto-decoded deep latent embeddings (ADDLE), which explicitly models the tendencies of each rater using an auto-decoder framework. After a simple linear transformation, the latent variables can be injected into any backbone at any and multiple points, allowing the model to account for rater-specific effects on the diagnosis. Importantly, ADDLE does not expect multiple raters per image in training, meaning it can readily learn from data mined from hospital archives. Moreover, the complexity of training ADDLE does not increase as more raters are added. During inference each rater can be simulated and a 'mean' or 'greedy' virtual rating can be produced. We test ADDLE on the problem of liver steatosis diagnosis from 2D ultrasound (US) by collecting 46 084 studies along with clinical US diagnoses originating from 65 different raters. We evaluated diagnostic performance using a separate dataset with gold-standard biopsy diagnoses. ADDLE can improve the partial areas under the curve (AUCs) for diagnosing severe steatosis by 10.5% over standard classifiers while outperforming other annotator-noise approaches, including those requiring 65 times the parameters.
SuctionNet-1Billion: A Large-Scale Benchmark for Suction Grasping
Mar 23, 2021



Suction is an important solution for the longstanding robotic grasping problem. Compared with other kinds of grasping, suction grasping is easier to represent and often more reliable in practice. Though preferred in many scenarios, it is not fully investigated and lacks sufficient training data and evaluation benchmarks. To address that, firstly, we propose a new physical model to analytically evaluate seal formation and wrench resistance of a suction grasping, which are two key aspects of grasp success. Secondly, a two-step methodology is adopted to generate annotations on a large-scale dataset collected in real-world cluttered scenarios. Thirdly, a standard online evaluation system is proposed to evaluate suction poses in continuous operation space, which can benchmark different algorithms fairly without the need of exhaustive labeling. Real-robot experiments are conducted to show that our annotations align well with real world. Meanwhile, we propose a method to predict numerous suction poses from an RGB-D image of a cluttered scene and demonstrate our superiority against several previous methods. Result analyses are further provided to help readers better understand the challenges in this area. Data and source code are publicly available at www.graspnet.net.
MR image reconstruction using deep density priors
Jan 17, 2018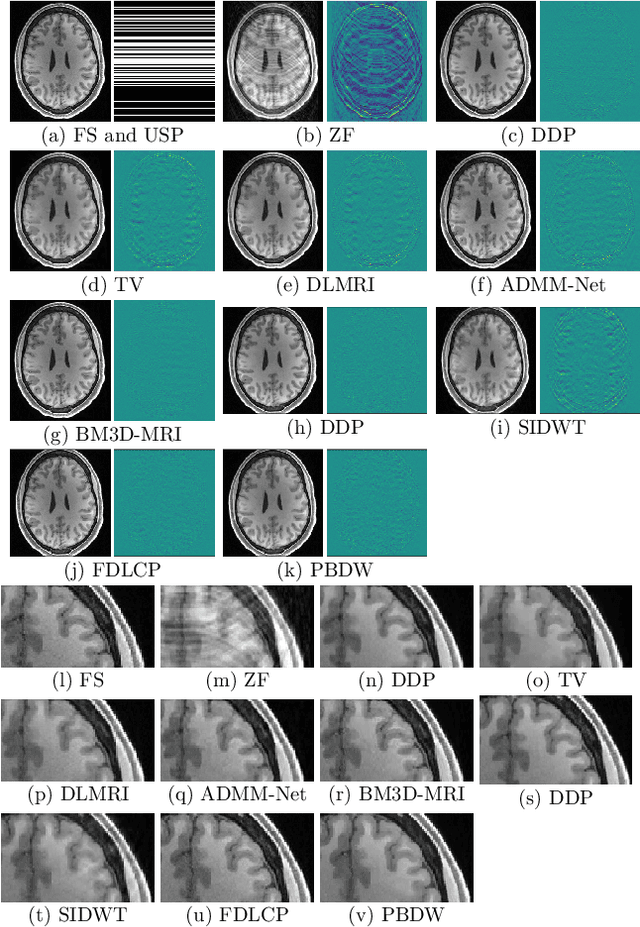
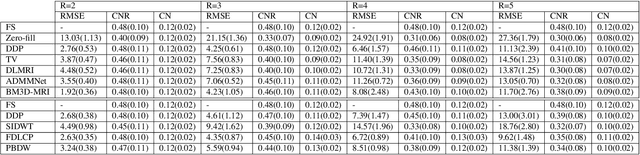
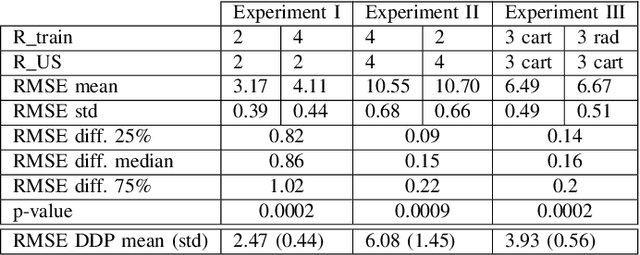
Purpose: MR image reconstruction exploits regularization to compensate for missing k-space data. In this work, we propose to learn the probability distribution of MR image patches with neural networks and use this distribution as prior information constraining images during reconstruction, effectively employing it as regularization. Methods: We use variational autoencoders (VAE) to learn the distribution of MR image patches, which models the high-dimensional distribution by a latent parameter model of lower dimensions in a non-linear fashion. The proposed algorithm uses the learned prior in a Maximum-A-Posteriori estimation formulation. We evaluate the proposed reconstruction method with T1 weighted images and also apply our method on images with white matter lesions. Results: Visual evaluation of the samples showed that the VAE algorithm can approximate the distribution of MR patches well. The proposed reconstruction algorithm using the VAE prior produced high quality reconstructions. The algorithm achieved normalized RMSE, CNR and CN values of 2.77\%, 0.43, 0.11; 4.29\%, 0.43, 0.11, 6.36\%, 0.47, 0.11 and 10.00\%, 0.42, 0.10 for undersampling ratios of 2, 3, 4 and 5, respectively, where it outperformed most of the alternative methods. In the experiments on images with white matter lesions, the method faithfully reconstructed the lesions. Conclusion: We introduced a novel method for MR reconstruction, which takes a new perspective on regularization by using priors learned by neural networks. Results suggest the method compares favorably against the other evaluated methods and can reconstruct lesions as well. Keywords: Reconstruction, MRI, prior probability, MAP estimation, machine learning, variational inference, deep learning
Algorithm-driven Advances for Scientific CT Instruments: From Model-based to Deep Learning-based Approaches
Apr 16, 2021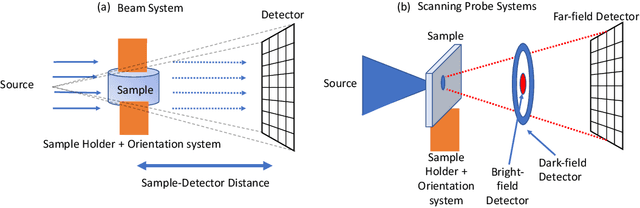
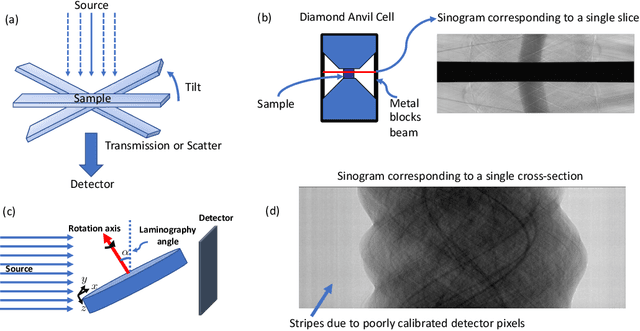
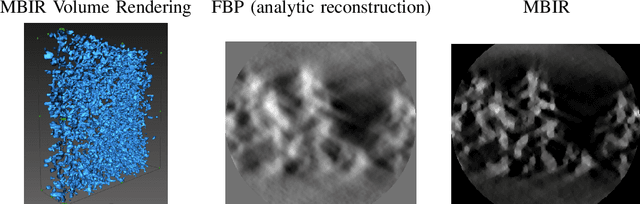
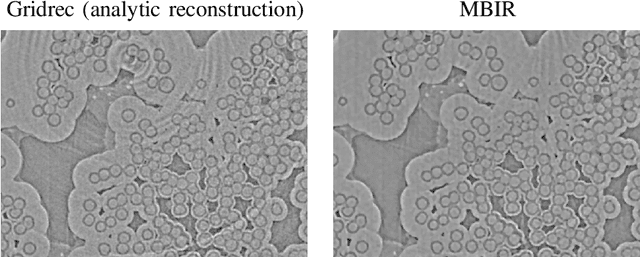
Multi-scale 3D characterization is widely used by materials scientists to further their understanding of the relationships between microscopic structure and macroscopic function. Scientific computed tomography (CT) instruments are one of the most popular choices for 3D non-destructive characterization of materials at length scales ranging from the angstrom-scale to the micron-scale. These instruments typically have a source of radiation that interacts with the sample to be studied and a detector assembly to capture the result of this interaction. A collection of such high-resolution measurements are made by re-orienting the sample which is mounted on a specially designed stage/holder after which reconstruction algorithms are used to produce the final 3D volume of interest. The end goal of scientific CT scans include determining the morphology,chemical composition or dynamic behavior of materials when subjected to external stimuli. In this article, we will present an overview of recent advances in reconstruction algorithms that have enabled significant improvements in the performance of scientific CT instruments - enabling faster, more accurate and novel imaging capabilities. In the first part, we will focus on model-based image reconstruction algorithms that formulate the inversion as solving a high-dimensional optimization problem involving a data-fidelity term and a regularization term. In the last part of the article, we will present an overview of recent approaches using deep-learning based algorithms for improving scientific CT instruments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge