"Image": models, code, and papers
Shortcut Detection with Variational Autoencoders
Feb 08, 2023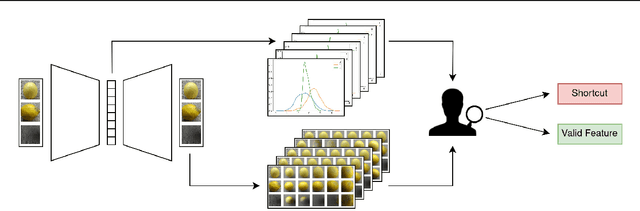
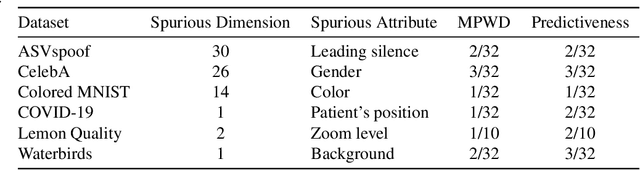
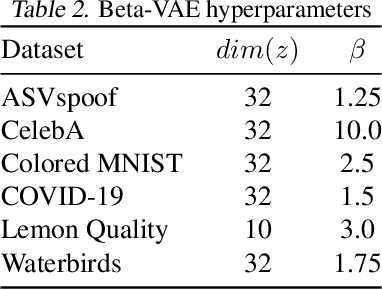

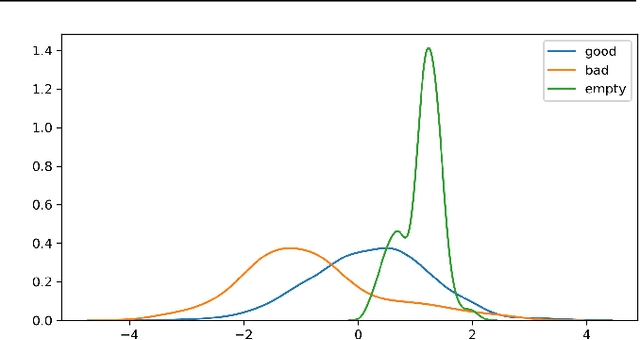

For real-world applications of machine learning (ML), it is essential that models make predictions based on well-generalizing features rather than spurious correlations in the data. The identification of such spurious correlations, also known as shortcuts, is a challenging problem and has so far been scarcely addressed. In this work, we present a novel approach to detect shortcuts in image and audio datasets by leveraging variational autoencoders (VAEs). The disentanglement of features in the latent space of VAEs allows us to discover correlations in datasets and semi-automatically evaluate them for ML shortcuts. We demonstrate the applicability of our method on several real-world datasets and identify shortcuts that have not been discovered before. Based on these findings, we also investigate the construction of shortcut adversarial examples.
GradMix for nuclei segmentation and classification in imbalanced pathology image datasets
Oct 24, 2022An automated segmentation and classification of nuclei is an essential task in digital pathology. The current deep learning-based approaches require a vast amount of annotated datasets by pathologists. However, the existing datasets are imbalanced among different types of nuclei in general, leading to a substantial performance degradation. In this paper, we propose a simple but effective data augmentation technique, termed GradMix, that is specifically designed for nuclei segmentation and classification. GradMix takes a pair of a major-class nucleus and a rare-class nucleus, creates a customized mixing mask, and combines them using the mask to generate a new rare-class nucleus. As it combines two nuclei, GradMix considers both nuclei and the neighboring environment by using the customized mixing mask. This allows us to generate realistic rare-class nuclei with varying environments. We employed two datasets to evaluate the effectiveness of GradMix. The experimental results suggest that GradMix is able to improve the performance of nuclei segmentation and classification in imbalanced pathology image datasets.
MILAN: Masked Image Pretraining on Language Assisted Representation
Aug 15, 2022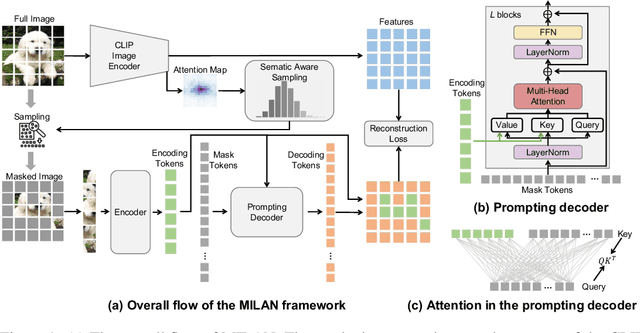
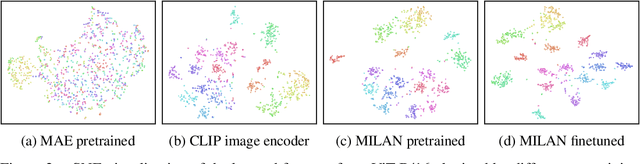
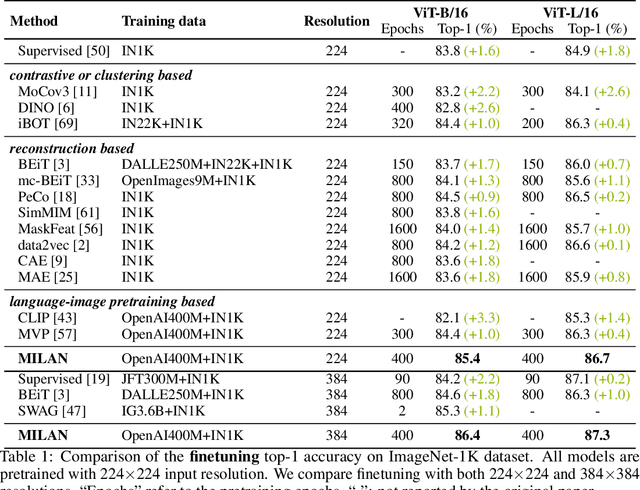

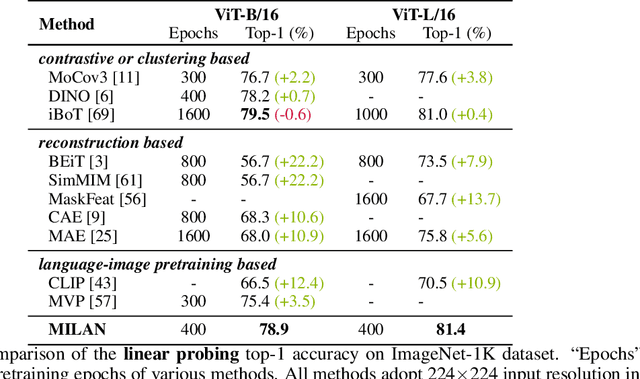
Self-attention based transformer models have been dominating many computer vision tasks in the past few years. Their superb model qualities heavily depend on the excessively large labeled image datasets. In order to reduce the reliance on large labeled datasets, reconstruction based masked autoencoders are gaining popularity, which learn high quality transferable representations from unlabeled images. For the same purpose, recent weakly supervised image pretraining methods explore language supervision from text captions accompanying the images. In this work, we propose masked image pretraining on language assisted representation, dubbed as MILAN. Instead of predicting raw pixels or low level features, our pretraining objective is to reconstruct the image features with substantial semantic signals that are obtained using caption supervision. Moreover, to accommodate our reconstruction target, we propose a more efficient prompting decoder architecture and a semantic aware mask sampling mechanism, which further advance the transfer performance of the pretrained model. Experimental results demonstrate that MILAN delivers higher accuracy than the previous works. When the masked autoencoder is pretrained and finetuned on ImageNet-1K dataset with an input resolution of 224x224, MILAN achieves a top-1 accuracy of 85.4% on ViTB/16, surpassing previous state-of-the-arts by 1%. In the downstream semantic segmentation task, MILAN achieves 52.7 mIoU using ViT-B/16 backbone on ADE20K dataset, outperforming previous masked pretraining results by 4 points.
Curvilinear object segmentation in medical images based on ODoS filter and deep learning network
Jan 18, 2023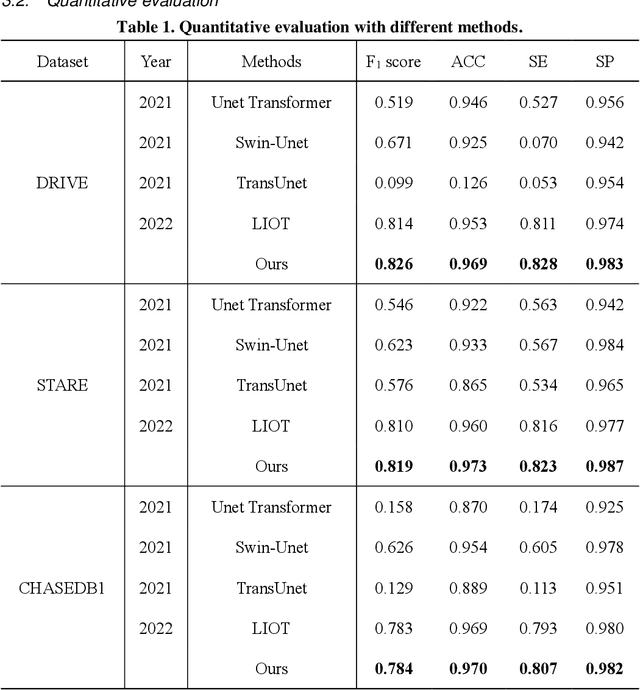
Automatic segmentation of curvilinear objects in medical images plays an important role in the diagnosis and evaluation of human diseases, yet it is a challenging uncertainty for the complex segmentation task due to different issues like various image appearance, low contrast between curvilinear objects and their surrounding backgrounds, thin and uneven curvilinear structures, and improper background illumination. To overcome these challenges, we present a unique curvilinear structure segmentation framework based on oriented derivative of stick (ODoS) filter and deep learning network for curvilinear object segmentation in medical images. Currently, a large number of deep learning models emphasis on developing deep architectures and ignore capturing the structural features of curvature objects, which may lead to unsatisfactory results. In consequence, a new approach that incorporates the ODoS filter as part of a deep learning network is presented to improve the spatial attention of curvilinear objects. In which, the original image is considered as principal part to describe various image appearance and complex background illumination, the multi-step strategy is used to enhance contrast between curvilinear objects and their surrounding backgrounds, and the vector field is applied to discriminate thin and uneven curvilinear structures. Subsequently, a deep learning framework is employed to extract varvious structural features for curvilinear object segmentation in medical images. The performance of the computational model was validated in experiments with publicly available DRIVE, STARE and CHASEDB1 datasets. Experimental results indicate that the presented model has yielded surprising results compared with some state-of-the-art methods.
Deep Learning for Medical Image Segmentation: Tricks, Challenges and Future Directions
Sep 21, 2022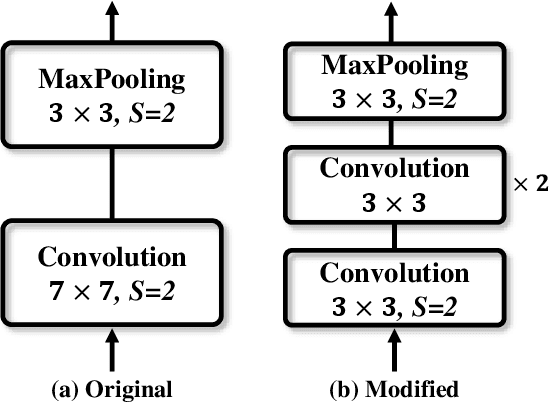
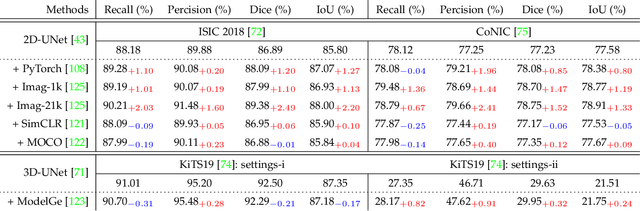
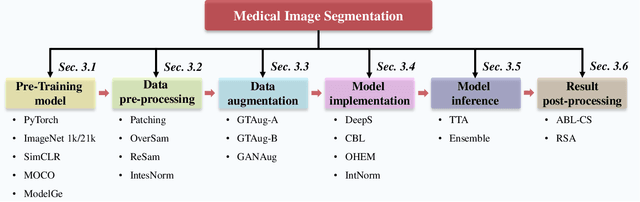
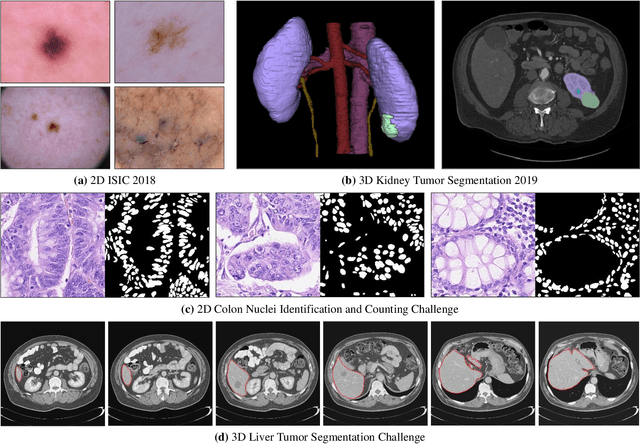
Over the past few years, the rapid development of deep learning technologies for computer vision has greatly promoted the performance of medical image segmentation (MedISeg). However, the recent MedISeg publications usually focus on presentations of the major contributions (e.g., network architectures, training strategies, and loss functions) while unwittingly ignoring some marginal implementation details (also known as "tricks"), leading to a potential problem of the unfair experimental result comparisons. In this paper, we collect a series of MedISeg tricks for different model implementation phases (i.e., pre-training model, data pre-processing, data augmentation, model implementation, model inference, and result post-processing), and experimentally explore the effectiveness of these tricks on the consistent baseline models. Compared to paper-driven surveys that only blandly focus on the advantages and limitation analyses of segmentation models, our work provides a large number of solid experiments and is more technically operable. With the extensive experimental results on both the representative 2D and 3D medical image datasets, we explicitly clarify the effect of these tricks. Moreover, based on the surveyed tricks, we also open-sourced a strong MedISeg repository, where each of its components has the advantage of plug-and-play. We believe that this milestone work not only completes a comprehensive and complementary survey of the state-of-the-art MedISeg approaches, but also offers a practical guide for addressing the future medical image processing challenges including but not limited to small dataset learning, class imbalance learning, multi-modality learning, and domain adaptation. The code has been released at: https://github.com/hust-linyi/MedISeg
Ultra-NeRF: Neural Radiance Fields for Ultrasound Imaging
Jan 25, 2023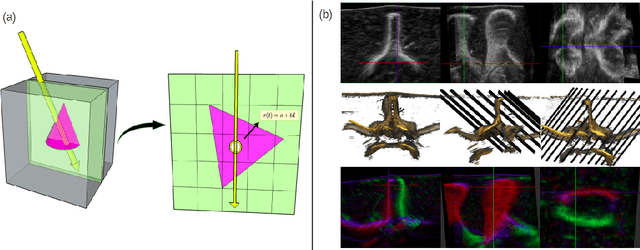
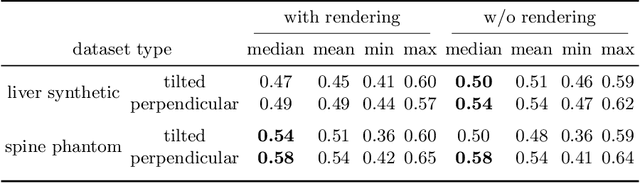
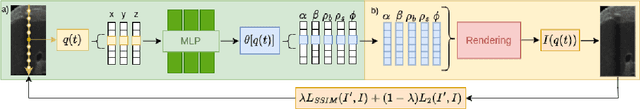
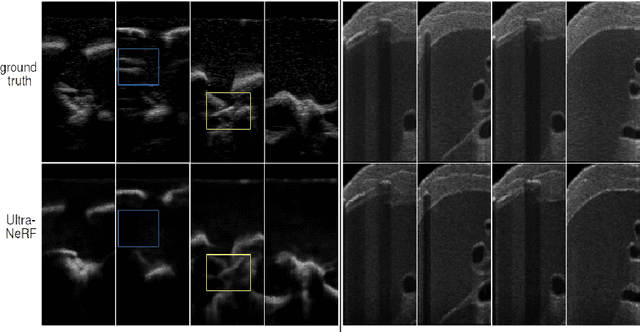
We present a physics-enhanced implicit neural representation (INR) for ultrasound (US) imaging that learns tissue properties from overlapping US sweeps. Our proposed method leverages a ray-tracing-based neural rendering for novel view US synthesis. Recent publications demonstrated that INR models could encode a representation of a three-dimensional scene from a set of two-dimensional US frames. However, these models fail to consider the view-dependent changes in appearance and geometry intrinsic to US imaging. In our work, we discuss direction-dependent changes in the scene and show that a physics-inspired rendering improves the fidelity of US image synthesis. In particular, we demonstrate experimentally that our proposed method generates geometrically accurate B-mode images for regions with ambiguous representation owing to view-dependent differences of the US images. We conduct our experiments using simulated B-mode US sweeps of the liver and acquired US sweeps of a spine phantom tracked with a robotic arm. The experiments corroborate that our method generates US frames that enable consistent volume compounding from previously unseen views. To the best of our knowledge, the presented work is the first to address view-dependent US image synthesis using INR.
Variational Deep Image Restoration
Jul 03, 2022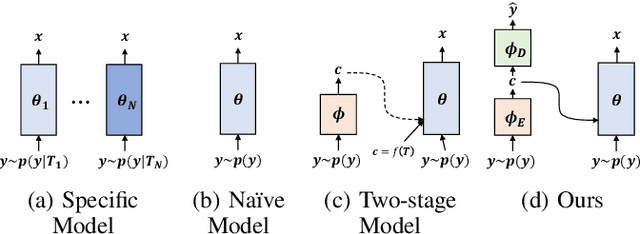
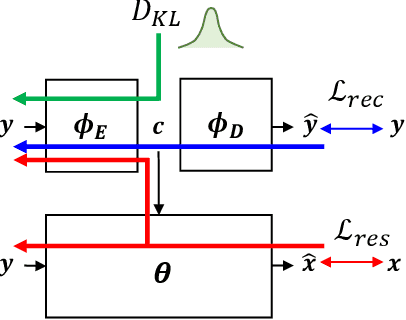
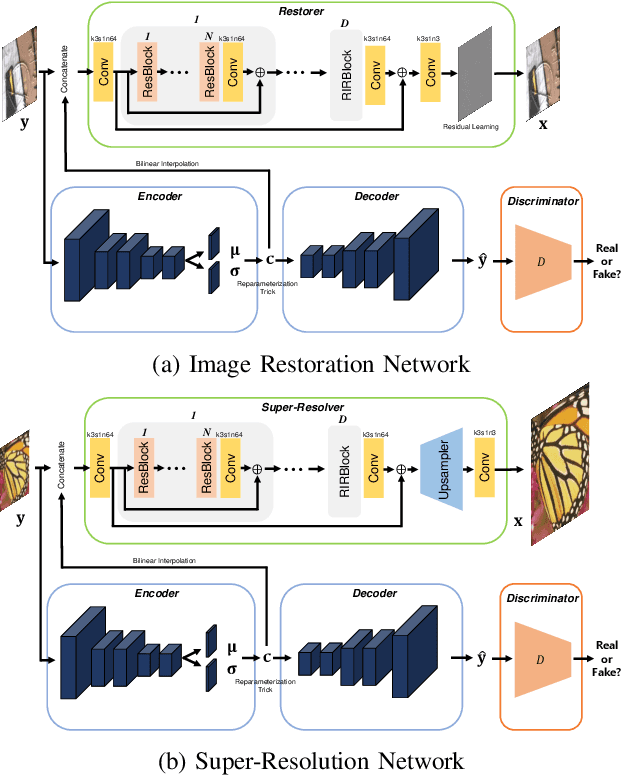
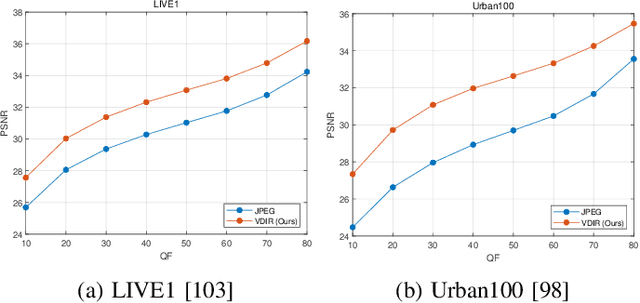
This paper presents a new variational inference framework for image restoration and a convolutional neural network (CNN) structure that can solve the restoration problems described by the proposed framework. Earlier CNN-based image restoration methods primarily focused on network architecture design or training strategy with non-blind scenarios where the degradation models are known or assumed. For a step closer to real-world applications, CNNs are also blindly trained with the whole dataset, including diverse degradations. However, the conditional distribution of a high-quality image given a diversely degraded one is too complicated to be learned by a single CNN. Therefore, there have also been some methods that provide additional prior information to train a CNN. Unlike previous approaches, we focus more on the objective of restoration based on the Bayesian perspective and how to reformulate the objective. Specifically, our method relaxes the original posterior inference problem to better manageable sub-problems and thus behaves like a divide-and-conquer scheme. As a result, the proposed framework boosts the performance of several restoration problems compared to the previous ones. Specifically, our method delivers state-of-the-art performance on Gaussian denoising, real-world noise reduction, blind image super-resolution, and JPEG compression artifacts reduction.
Learning to Collocate Visual-Linguistic Neural Modules for Image Captioning
Oct 04, 2022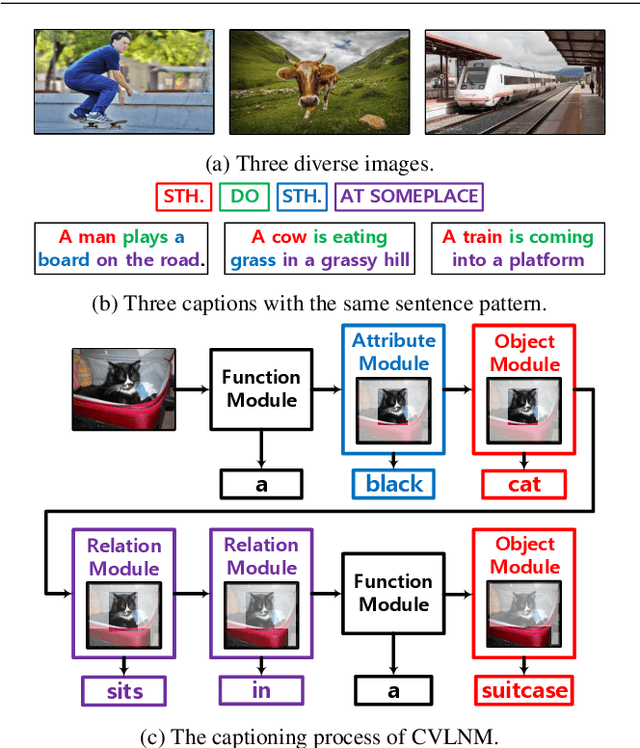
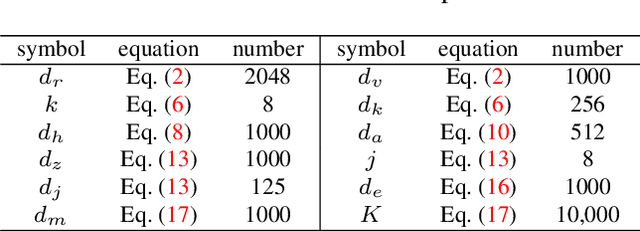


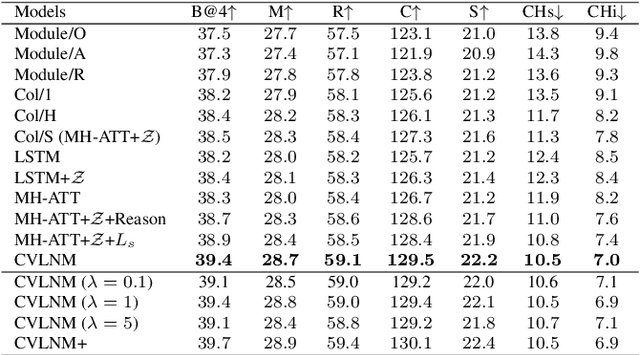
Humans tend to decompose a sentence into different parts like \textsc{sth do sth at someplace} and then fill each part with certain content. Inspired by this, we follow the \textit{principle of modular design} to propose a novel image captioner: learning to Collocate Visual-Linguistic Neural Modules (CVLNM). Unlike the \re{widely used} neural module networks in VQA, where the language (\ie, question) is fully observable, \re{the task of collocating visual-linguistic modules is more challenging.} This is because the language is only partially observable, for which we need to dynamically collocate the modules during the process of image captioning. To sum up, we make the following technical contributions to design and train our CVLNM: 1) \textit{distinguishable module design} -- \re{four modules in the encoder} including one linguistic module for function words and three visual modules for different content words (\ie, noun, adjective, and verb) and another linguistic one in the decoder for commonsense reasoning, 2) a self-attention based \textit{module controller} for robustifying the visual reasoning, 3) a part-of-speech based \textit{syntax loss} imposed on the module controller for further regularizing the training of our CVLNM. Extensive experiments on the MS-COCO dataset show that our CVLNM is more effective, \eg, achieving a new state-of-the-art 129.5 CIDEr-D, and more robust, \eg, being less likely to overfit to dataset bias and suffering less when fewer training samples are available. Codes are available at \url{https://github.com/GCYZSL/CVLMN}
Retinal Image Restoration and Vessel Segmentation using Modified Cycle-CBAM and CBAM-UNet
Sep 09, 2022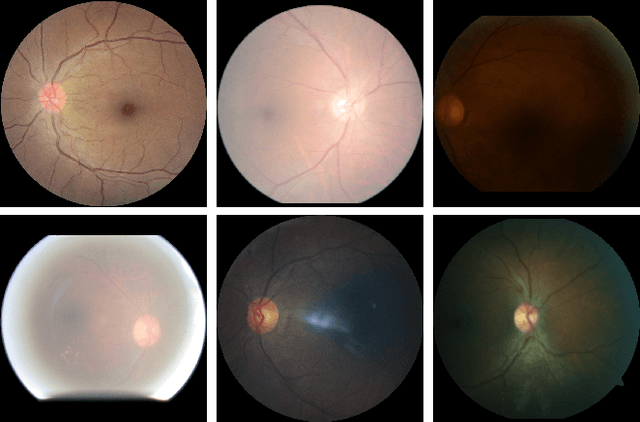
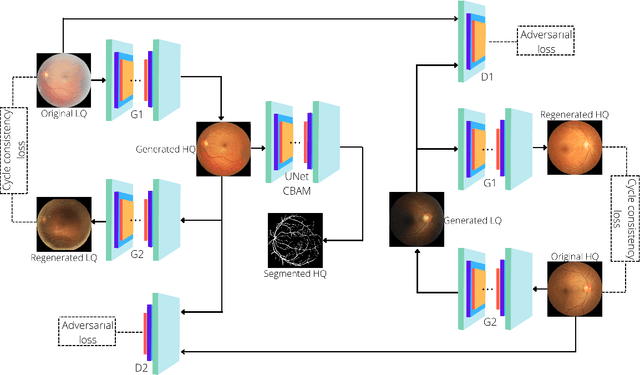
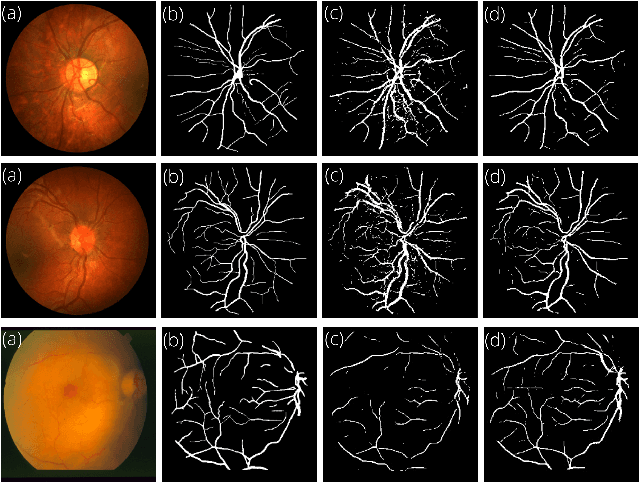
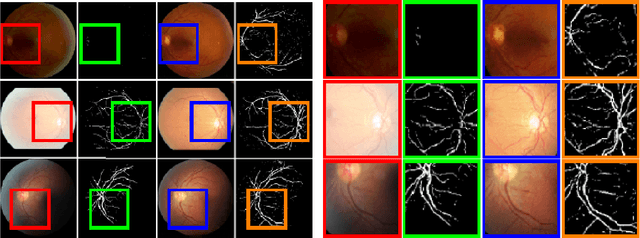
Clinical screening with low-quality fundus images is challenging and significantly leads to misdiagnosis. This paper addresses the issue of improving the retinal image quality and vessel segmentation through retinal image restoration. More specifically, a cycle-consistent generative adversarial network (CycleGAN) with a convolution block attention module (CBAM) is used for retinal image restoration. A modified UNet is used for retinal vessel segmentation for the restored retinal images (CBAM-UNet). The proposed model consists of two generators and two discriminators. Generators translate images from one domain to another, i.e., from low to high quality and vice versa. Discriminators classify generated and original images. The retinal vessel segmentation model uses downsampling, bottlenecking, and upsampling layers to generate segmented images. The CBAM has been used to enhance the feature extraction of these models. The proposed method does not require paired image datasets, which are challenging to produce. Instead, it uses unpaired data that consists of low- and high-quality fundus images retrieved from publicly available datasets. The restoration performance of the proposed method was evaluated using full-reference evaluation metrics, e.g., peak signal-to-noise ratio (PSNR) and structural similarity index measure (SSIM). The retinal vessel segmentation performance was compared with the ground-truth fundus images. The proposed method can significantly reduce the degradation effects caused by out-of-focus blurring, color distortion, low, high, and uneven illumination. Experimental results show the effectiveness of the proposed method for retinal image restoration and vessel segmentation.
Scale-MAE: A Scale-Aware Masked Autoencoder for Multiscale Geospatial Representation Learning
Jan 02, 2023
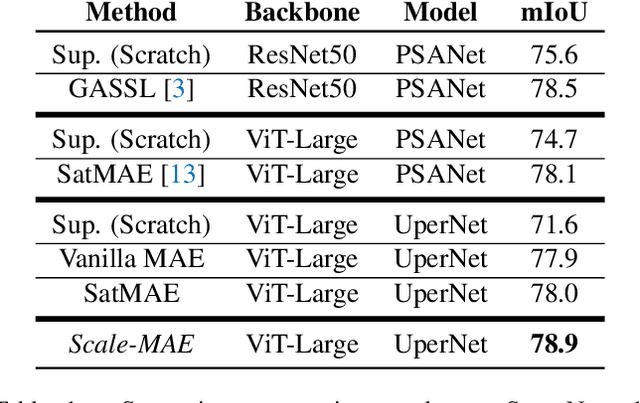
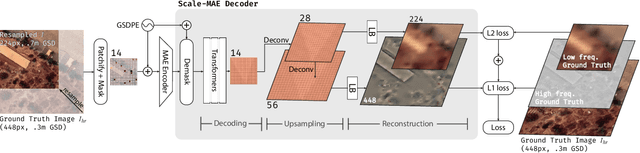
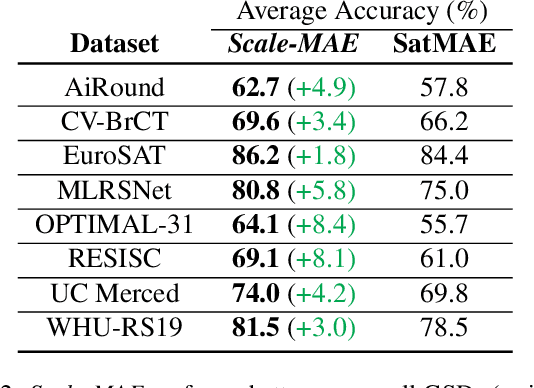
Remote sensing imagery provides comprehensive views of the Earth, where different sensors collect complementary data at different spatial scales. Large, pretrained models are commonly finetuned with imagery that is heavily augmented to mimic different conditions and scales, with the resulting models used for various tasks with imagery from a range of spatial scales. Such models overlook scale-specific information in the data. In this paper, we present Scale-MAE, a pretraining method that explicitly learns relationships between data at different, known scales throughout the pretraining process. Scale-MAE pretrains a network by masking an input image at a known input scale, where the area of the Earth covered by the image determines the scale of the ViT positional encoding, not the image resolution. Scale-MAE encodes the masked image with a standard ViT backbone, and then decodes the masked image through a bandpass filter to reconstruct low/high frequency images at lower/higher scales. We find that tasking the network with reconstructing both low/high frequency images leads to robust multiscale representations for remote sensing imagery. Scale-MAE achieves an average of a $5.0\%$ non-parametric kNN classification improvement across eight remote sensing datasets compared to current state-of-the-art and obtains a $0.9$ mIoU to $3.8$ mIoU improvement on the SpaceNet building segmentation transfer task for a range of evaluation scales.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge