Chexnet
Papers and Code
Reproducing and Improving CheXNet: Deep Learning for Chest X-ray Disease Classification
May 10, 2025Deep learning for radiologic image analysis is a rapidly growing field in biomedical research and is likely to become a standard practice in modern medicine. On the publicly available NIH ChestX-ray14 dataset, containing X-ray images that are classified by the presence or absence of 14 different diseases, we reproduced an algorithm known as CheXNet, as well as explored other algorithms that outperform CheXNet's baseline metrics. Model performance was primarily evaluated using the F1 score and AUC-ROC, both of which are critical metrics for imbalanced, multi-label classification tasks in medical imaging. The best model achieved an average AUC-ROC score of 0.85 and an average F1 score of 0.39 across all 14 disease classifications present in the dataset.
Lung Disease Detection with Vision Transformers: A Comparative Study of Machine Learning Methods
Nov 18, 2024



Recent advancements in medical image analysis have predominantly relied on Convolutional Neural Networks (CNNs), achieving impressive performance in chest X-ray classification tasks, such as the 92% AUC reported by AutoThorax-Net and the 88% AUC achieved by ChexNet in classifcation tasks. However, in the medical field, even small improvements in accuracy can have significant clinical implications. This study explores the application of Vision Transformers (ViT), a state-of-the-art architecture in machine learning, to chest X-ray analysis, aiming to push the boundaries of diagnostic accuracy. I present a comparative analysis of two ViT-based approaches: one utilizing full chest X-ray images and another focusing on segmented lung regions. Experiments demonstrate that both methods surpass the performance of traditional CNN-based models, with the full-image ViT achieving up to 97.83% accuracy and the lung-segmented ViT reaching 96.58% accuracy in classifcation of diseases on three label and AUC of 94.54% when label numbers are increased to eight. Notably, the full-image approach showed superior performance across all metrics, including precision, recall, F1 score, and AUC-ROC. These findings suggest that Vision Transformers can effectively capture relevant features from chest X-rays without the need for explicit lung segmentation, potentially simplifying the preprocessing pipeline while maintaining high accuracy. This research contributes to the growing body of evidence supporting the efficacy of transformer-based architectures in medical image analysis and highlights their potential to enhance diagnostic precision in clinical settings.
Calibrating Deep Neural Network using Euclidean Distance
Oct 23, 2024



Uncertainty is a fundamental aspect of real-world scenarios, where perfect information is rarely available. Humans naturally develop complex internal models to navigate incomplete data and effectively respond to unforeseen or partially observed events. In machine learning, Focal Loss is commonly used to reduce misclassification rates by emphasizing hard-to-classify samples. However, it does not guarantee well-calibrated predicted probabilities and may result in models that are overconfident or underconfident. High calibration error indicates a misalignment between predicted probabilities and actual outcomes, affecting model reliability. This research introduces a novel loss function called Focal Calibration Loss (FCL), designed to improve probability calibration while retaining the advantages of Focal Loss in handling difficult samples. By minimizing the Euclidean norm through a strictly proper loss, FCL penalizes the instance-wise calibration error and constrains bounds. We provide theoretical validation for proposed method and apply it to calibrate CheXNet for potential deployment in web-based health-care systems. Extensive evaluations on various models and datasets demonstrate that our method achieves SOTA performance in both calibration and accuracy metrics.
Shadow and Light: Digitally Reconstructed Radiographs for Disease Classification
Jun 06, 2024



In this paper, we introduce DRR-RATE, a large-scale synthetic chest X-ray dataset derived from the recently released CT-RATE dataset. DRR-RATE comprises of 50,188 frontal Digitally Reconstructed Radiographs (DRRs) from 21,304 unique patients. Each image is paired with a corresponding radiology text report and binary labels for 18 pathology classes. Given the controllable nature of DRR generation, it facilitates the inclusion of lateral view images and images from any desired viewing position. This opens up avenues for research into new and novel multimodal applications involving paired CT, X-ray images from various views, text, and binary labels. We demonstrate the applicability of DRR-RATE alongside existing large-scale chest X-ray resources, notably the CheXpert dataset and CheXnet model. Experiments demonstrate that CheXnet, when trained and tested on the DRR-RATE dataset, achieves sufficient to high AUC scores for the six common pathologies cited in common literature: Atelectasis, Cardiomegaly, Consolidation, Lung Lesion, Lung Opacity, and Pleural Effusion. Additionally, CheXnet trained on the CheXpert dataset can accurately identify several pathologies, even when operating out of distribution. This confirms that the generated DRR images effectively capture the essential pathology features from CT images. The dataset and labels are publicly accessible at https://huggingface.co/datasets/farrell236/DRR-RATE.
COVID-19 Detection Based on Self-Supervised Transfer Learning Using Chest X-Ray Images
Dec 19, 2022Purpose: Considering several patients screened due to COVID-19 pandemic, computer-aided detection has strong potential in assisting clinical workflow efficiency and reducing the incidence of infections among radiologists and healthcare providers. Since many confirmed COVID-19 cases present radiological findings of pneumonia, radiologic examinations can be useful for fast detection. Therefore, chest radiography can be used to fast screen COVID-19 during the patient triage, thereby determining the priority of patient's care to help saturated medical facilities in a pandemic situation. Methods: In this paper, we propose a new learning scheme called self-supervised transfer learning for detecting COVID-19 from chest X-ray (CXR) images. We compared six self-supervised learning (SSL) methods (Cross, BYOL, SimSiam, SimCLR, PIRL-jigsaw, and PIRL-rotation) with the proposed method. Additionally, we compared six pretrained DCNNs (ResNet18, ResNet50, ResNet101, CheXNet, DenseNet201, and InceptionV3) with the proposed method. We provide quantitative evaluation on the largest open COVID-19 CXR dataset and qualitative results for visual inspection. Results: Our method achieved a harmonic mean (HM) score of 0.985, AUC of 0.999, and four-class accuracy of 0.953. We also used the visualization technique Grad-CAM++ to generate visual explanations of different classes of CXR images with the proposed method to increase the interpretability. Conclusions: Our method shows that the knowledge learned from natural images using transfer learning is beneficial for SSL of the CXR images and boosts the performance of representation learning for COVID-19 detection. Our method promises to reduce the incidence of infections among radiologists and healthcare providers.
A knee cannot have lung disease: out-of-distribution detection with in-distribution voting using the medical example of chest X-ray classification
Aug 01, 2022



Deep learning models are being applied to more and more use cases with astonishing success stories, but how do they perform in the real world? To test a model, a specific cleaned data set is assembled. However, when deployed in the real world, the model will face unexpected, out-of-distribution (OOD) data. In this work, we show that the so-called "radiologist-level" CheXnet model fails to recognize all OOD images and classifies them as having lung disease. To address this issue, we propose in-distribution voting, a novel method to classify out-of-distribution images for multi-label classification. Using independent class-wise in-distribution (ID) predictors trained on ID and OOD data we achieve, on average, 99 % ID classification specificity and 98 % sensitivity, improving the end-to-end performance significantly compared to previous works on the chest X-ray 14 data set. Our method surpasses other output-based OOD detectors even when trained solely with ImageNet as OOD data and tested with X-ray OOD images.
Transfer learning approach to Classify the X-ray image that corresponds to corona disease Using ResNet50 pretrained by ChexNet
May 18, 2021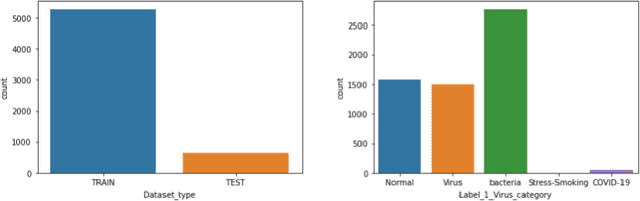
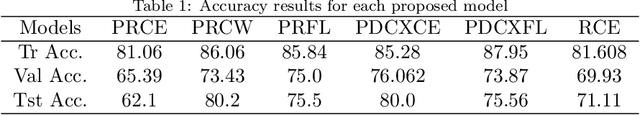
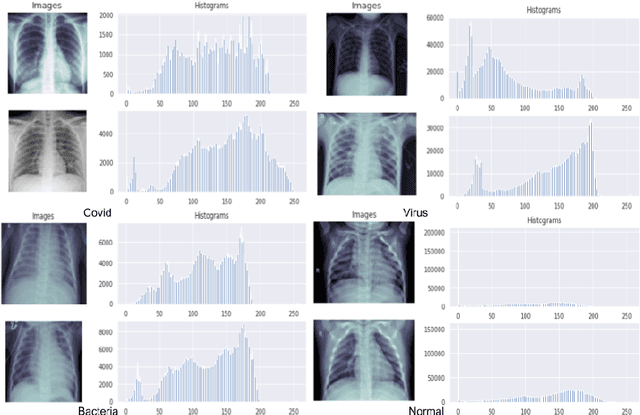
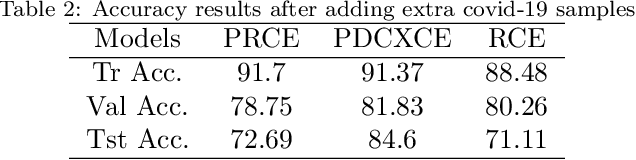
Coronavirus adversely has affected people worldwide. There are common symptoms between the Covid19 virus disease and other respiratory diseases like pneumonia or Influenza. Therefore, diagnosing it fast is crucial not only to save patients but also to prevent it from spreading. One of the most reliant methods of diagnosis is through X-ray images of a lung. With the help of deep learning approaches, we can teach the deep model to learn the condition of an affected lung. Therefore, it can classify the new sample as if it is a Covid19 infected patient or not. In this project, we train a deep model based on ResNet50 pretrained by ImageNet dataset and CheXNet dataset. Based on the imbalanced CoronaHack Chest X-Ray dataset introducing by Kaggle we applied both binary and multi-class classification. Also, we compare the results when using Focal loss and Cross entropy loss.
Rethinking annotation granularity for overcoming deep shortcut learning: A retrospective study on chest radiographs
Apr 21, 2021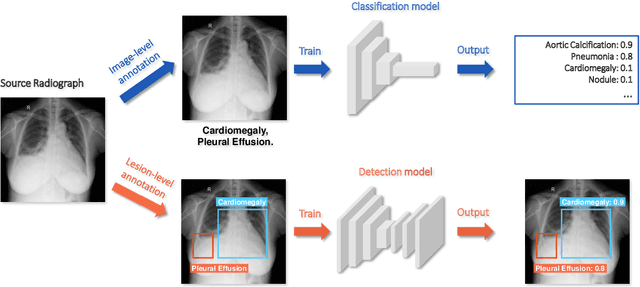
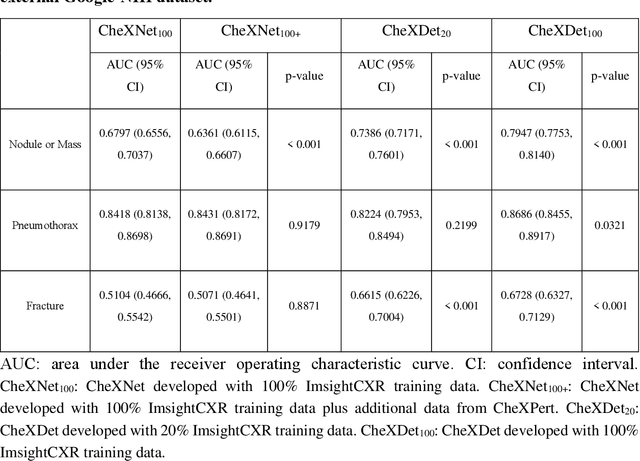
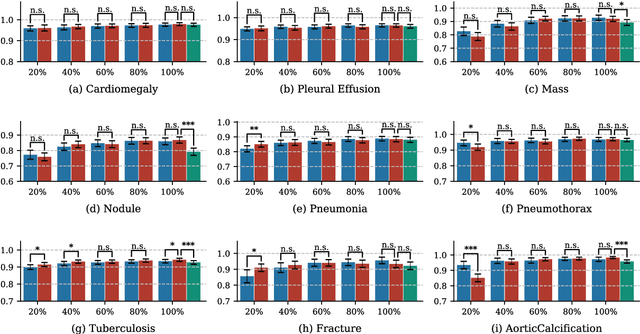
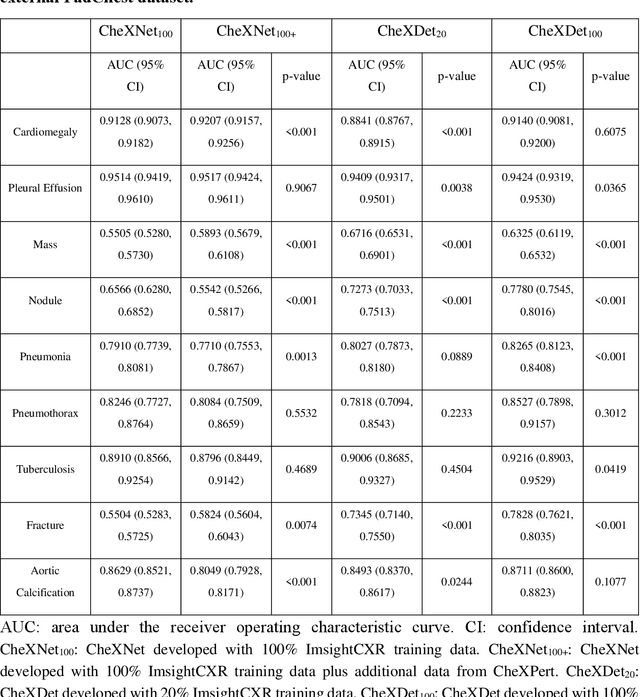
Deep learning has demonstrated radiograph screening performances that are comparable or superior to radiologists. However, recent studies show that deep models for thoracic disease classification usually show degraded performance when applied to external data. Such phenomena can be categorized into shortcut learning, where the deep models learn unintended decision rules that can fit the identically distributed training and test set but fail to generalize to other distributions. A natural way to alleviate this defect is explicitly indicating the lesions and focusing the model on learning the intended features. In this paper, we conduct extensive retrospective experiments to compare a popular thoracic disease classification model, CheXNet, and a thoracic lesion detection model, CheXDet. We first showed that the two models achieved similar image-level classification performance on the internal test set with no significant differences under many scenarios. Meanwhile, we found incorporating external training data even led to performance degradation for CheXNet. Then, we compared the models' internal performance on the lesion localization task and showed that CheXDet achieved significantly better performance than CheXNet even when given 80% less training data. By further visualizing the models' decision-making regions, we revealed that CheXNet learned patterns other than the target lesions, demonstrating its shortcut learning defect. Moreover, CheXDet achieved significantly better external performance than CheXNet on both the image-level classification task and the lesion localization task. Our findings suggest improving annotation granularity for training deep learning systems as a promising way to elevate future deep learning-based diagnosis systems for clinical usage.
PneumoXttention: A CNN compensating for Human Fallibility when Detecting Pneumonia through CXR images with Attention
Aug 11, 2020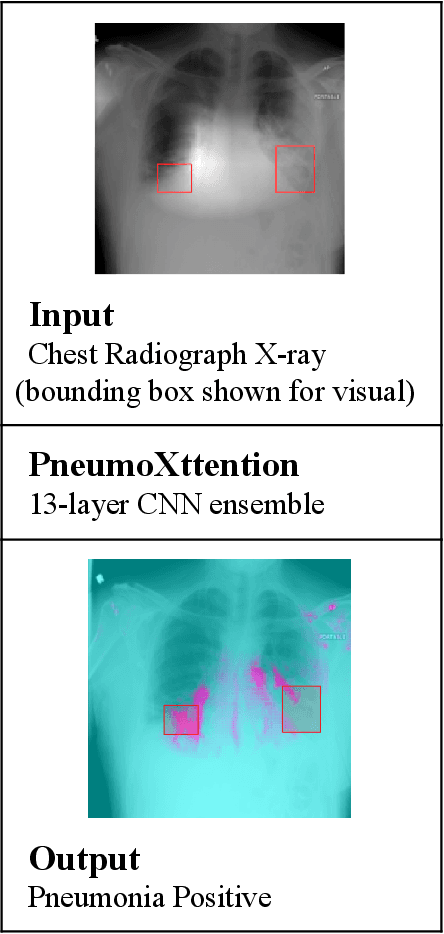

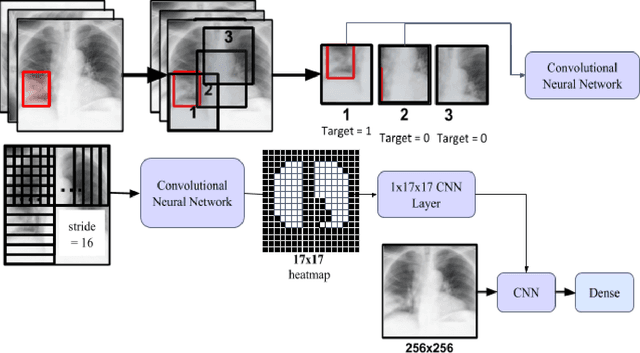
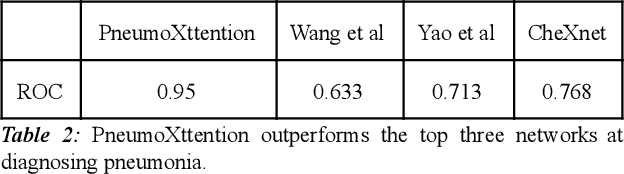
Automatic Chest Radiograph X-ray (CXR) interpretation by machines is an important research topic of Artificial Intelligence. As part of my journey through the California Science Fair, I have developed an algorithm that can detect pneumonia from a CXR image to compensate for human fallibility. My algorithm, PneumoXttention, is an ensemble of two 13 layer convolutional neural network trained on the RSNA dataset, a dataset provided by the Radiological Society of North America, containing 26,684 frontal X-ray images split into the categories of pneumonia and no pneumonia. The dataset was annotated by many professional radiologists in North America. It achieved an impressive F1 score, 0.82, on the test set (20% random split of RSNA dataset) and completely compensated Human Radiologists on a random set of 25 test images drawn from RSNA and NIH. I don't have a direct comparison but Stanford's Chexnet has a F1 score of 0.435 on the NIH dataset for category Pneumonia.
Reliable Tuberculosis Detection using Chest X-ray with Deep Learning, Segmentation and Visualization
Jul 29, 2020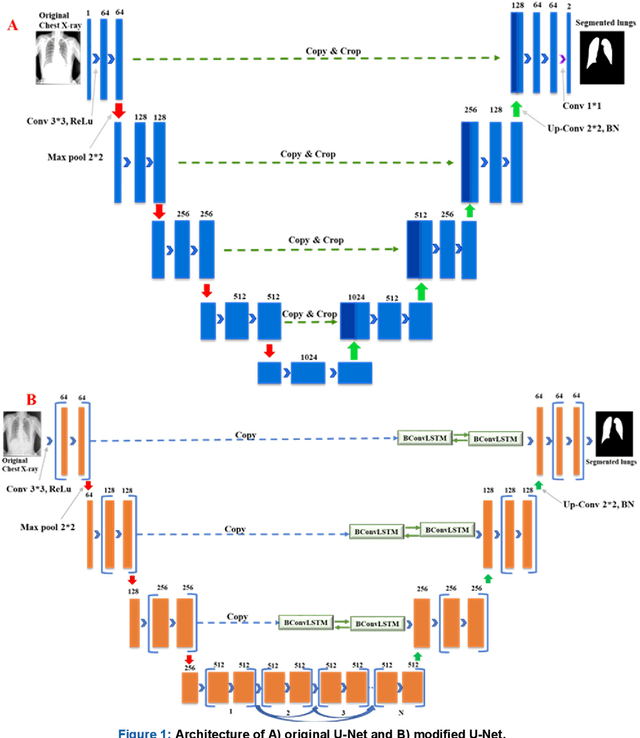
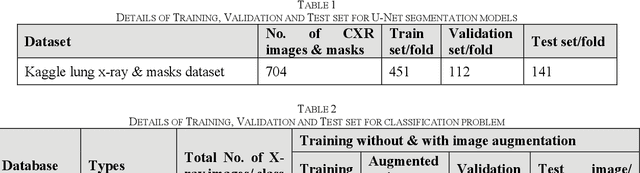
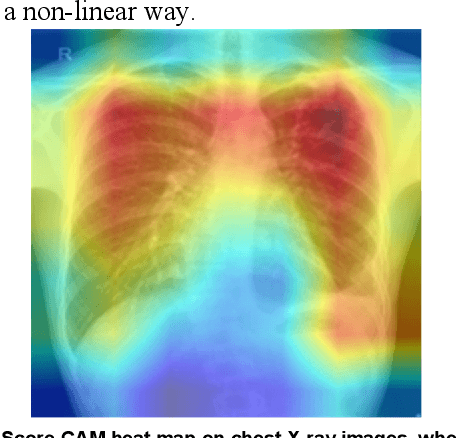
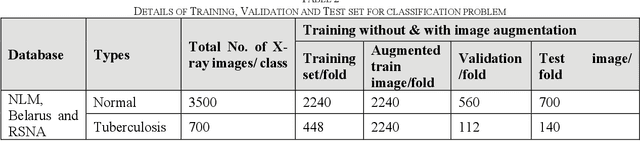
Tuberculosis (TB) is a chronic lung disease that occurs due to bacterial infection and is one of the top 10 leading causes of death. Accurate and early detection of TB is very important, otherwise, it could be life-threatening. In this work, we have detected TB reliably from the chest X-ray images using image pre-processing, data augmentation, image segmentation, and deep-learning classification techniques. Several public databases were used to create a database of 700 TB infected and 3500 normal chest X-ray images for this study. Nine different deep CNNs (ResNet18, ResNet50, ResNet101, ChexNet, InceptionV3, Vgg19, DenseNet201, SqueezeNet, and MobileNet), which were used for transfer learning from their pre-trained initial weights and trained, validated and tested for classifying TB and non-TB normal cases. Three different experiments were carried out in this work: segmentation of X-ray images using two different U-net models, classification using X-ray images, and segmented lung images. The accuracy, precision, sensitivity, F1-score, specificity in the detection of tuberculosis using X-ray images were 97.07 %, 97.34 %, 97.07 %, 97.14 % and 97.36 % respectively. However, segmented lungs for the classification outperformed than whole X-ray image-based classification and accuracy, precision, sensitivity, F1-score, specificity were 99.9 %, 99.91 %, 99.9 %, 99.9 %, and 99.52 % respectively. The paper also used a visualization technique to confirm that CNN learns dominantly from the segmented lung regions results in higher detection accuracy. The proposed method with state-of-the-art performance can be useful in the computer-aided faster diagnosis of tuberculosis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge