Sanskriti Singh
CheX-Nomaly: Segmenting Lung Abnormalities from Chest Radiographs using Machine Learning
Nov 03, 2023Abstract:The global challenge in chest radiograph X-ray (CXR) abnormalities often being misdiagnosed is primarily associated with perceptual errors, where healthcare providers struggle to accurately identify the location of abnormalities, rather than misclassification errors. We currently address this problem through disease-specific segmentation models. Unfortunately, these models cannot be released in the field due to their lack of generalizability across all thoracic diseases. A binary model tends to perform poorly when it encounters a disease that isn't represented in the dataset. We present CheX-nomaly: a binary localization U-net model that leverages transfer learning techniques with the incorporation of an innovative contrastive learning approach. Trained on the VinDr-CXR dataset, which encompasses 14 distinct diseases in addition to 'no finding' cases, my model achieves generalizability across these 14 diseases and others it has not seen before. We show that we can significantly improve the generalizability of an abnormality localization model by incorporating a contrastive learning method and dissociating the bounding boxes with its disease class. We also introduce a new loss technique to apply to enhance the U-nets performance on bounding box segmentation. By introducing CheX-nomaly, we offer a promising solution to enhance the precision of chest disease diagnosis, with a specific focus on reducing the significant number of perceptual errors in healthcare.
Automated Coronary Calcium Scoring using U-Net Models through Semi-supervised Learning on Non-Gated CT Scans
Jun 13, 2022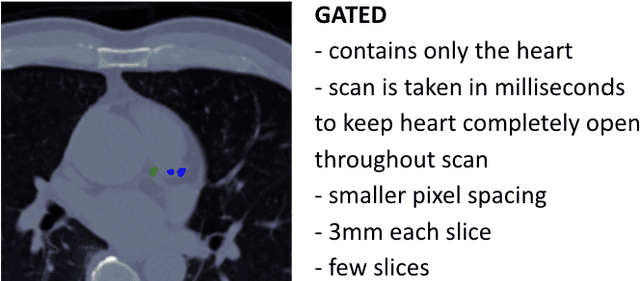
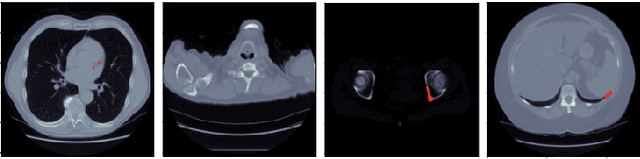

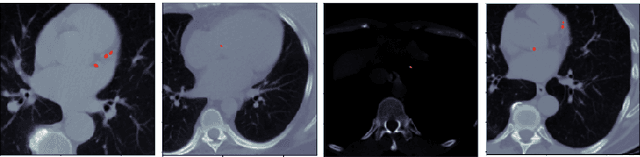
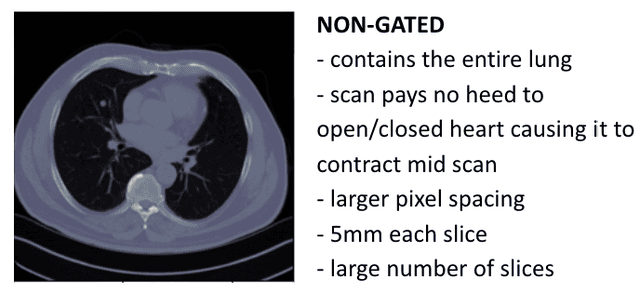
Abstract:Every year, thousands of innocent people die due to heart attacks. Often undiagnosed heart attacks can hit people by surprise since many current medical plans don't cover the costs to require the searching of calcification on these scans. Only if someone is suspected to have a heart problem, a gated CT scan is taken, otherwise, there's no way for the patient to be aware of a possible heart attack/disease. While nongated CT scans are more periodically taken, it is harder to detect calcification and is usually taken for a purpose other than locating calcification in arteries. In fact, in real time coronary artery calcification scores are only calculated on gated CT scans, not nongated CT scans. After training a unet model on the Coronary Calcium and chest CT's gated scans, it received a DICE coefficient of 0.95 on its untouched test set. This model was used to predict on nongated CT scans, performing with a mean absolute error (MAE) of 674.19 and bucket classification accuracy of 41% (5 classes). Through the analysis of the images and the information stored in the images, mathematical equations were derived and used to automatically crop the images around the location of the heart. By performing semi-supervised learning the new cropped nongated scans were able to closely resemble gated CT scans, improving the performance by 91% in MAE (62.38) and 23% in accuracy.
Emphasis on the Minimization of False Negatives or False Positives in Binary Classification
Apr 06, 2022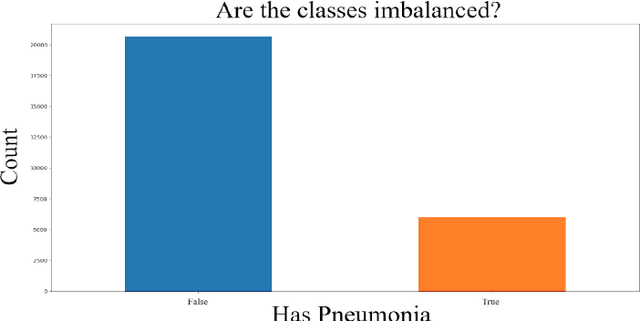
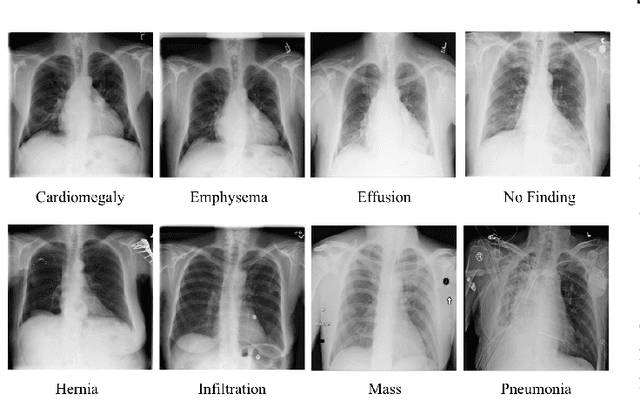
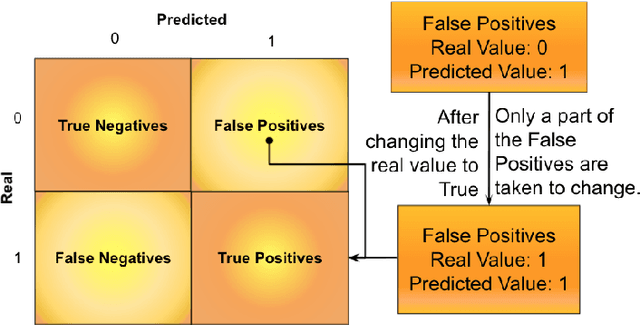
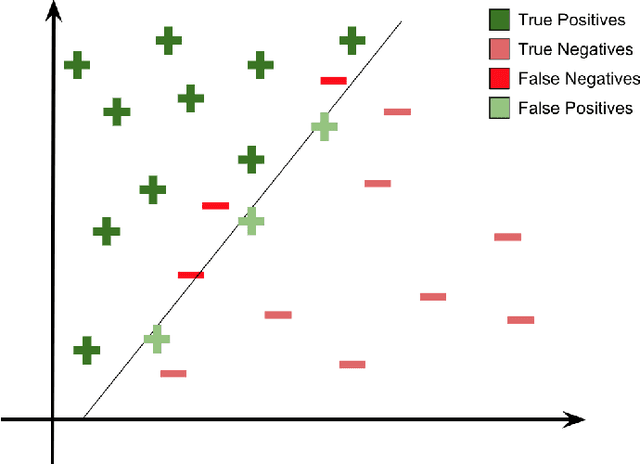
Abstract:The minimization of specific cases in binary classification, such as false negatives or false positives, grows increasingly important as humans begin to implement more machine learning into current products. While there are a few methods to put a bias towards the reduction of specific cases, these methods aren't very effective, hence their minimal use in models. To this end, a new method is introduced to reduce the False Negatives or False positives without drastically changing the overall performance or F1 score of the model. This method involving the careful change to the real value of the input after pre-training the model. Presenting the results of this method being applied on various datasets, some being more complex than others. Through experimentation on multiple model architectures on these datasets, the best model was found. In all the models, an increase in the recall or precision, minimization of False Negatives or False Positives respectively, was shown without a large drop in F1 score.
A Novel Mask R-CNN Model to Segment Heterogeneous Brain Tumors through Image Subtraction
Apr 04, 2022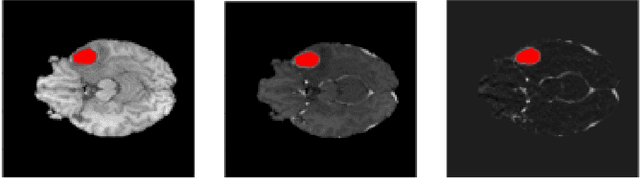
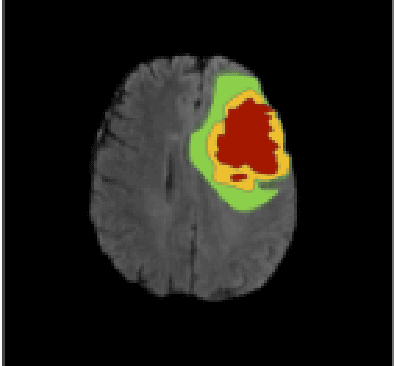
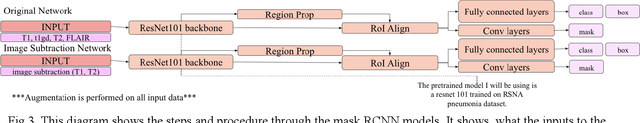
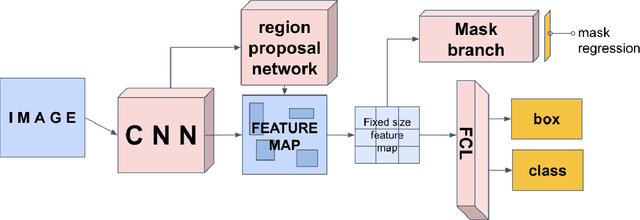
Abstract:The segmentation of diseases is a popular topic explored by researchers in the field of machine learning. Brain tumors are extremely dangerous and require the utmost precision to segment for a successful surgery. Patients with tumors usually take 4 MRI scans, T1, T1gd, T2, and FLAIR, which are then sent to radiologists to segment and analyze for possible future surgery. To create a second segmentation, it would be beneficial to both radiologists and patients in being more confident in their conclusions. We propose using a method performed by radiologists called image segmentation and applying it to machine learning models to prove a better segmentation. Using Mask R-CNN, its ResNet backbone being pre-trained on the RSNA pneumonia detection challenge dataset, we can train a model on the Brats2020 Brain Tumor dataset. Center for Biomedical Image Computing & Analytics provides MRI data on patients with and without brain tumors and the corresponding segmentations. We can see how well the method of image subtraction works by comparing it to models without image subtraction through DICE coefficient (F1 score), recall, and precision on the untouched test set. Our model performed with a DICE coefficient of 0.75 in comparison to 0.69 without image subtraction. To further emphasize the usefulness of image subtraction, we compare our final model to current state-of-the-art models to segment tumors from MRI scans.
PneumoXttention: A CNN compensating for Human Fallibility when Detecting Pneumonia through CXR images with Attention
Aug 11, 2020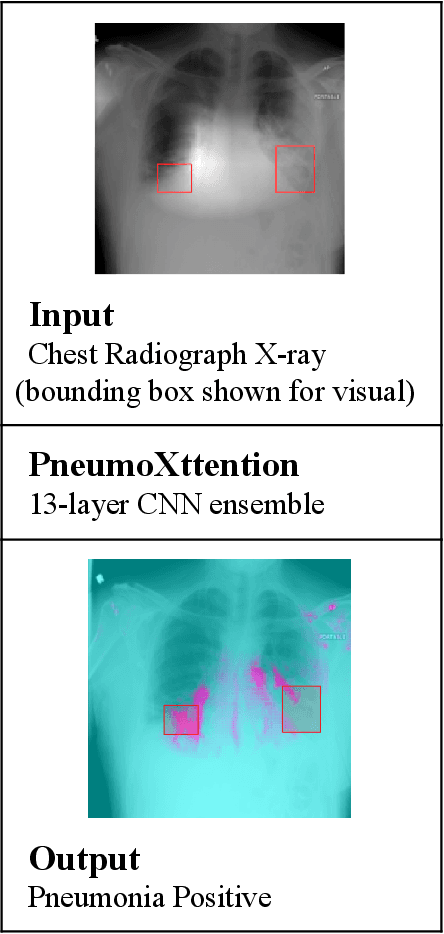

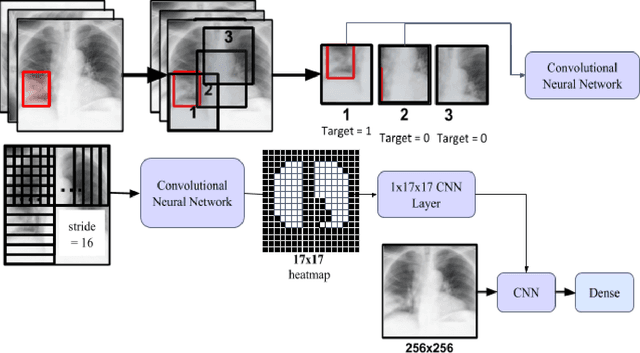
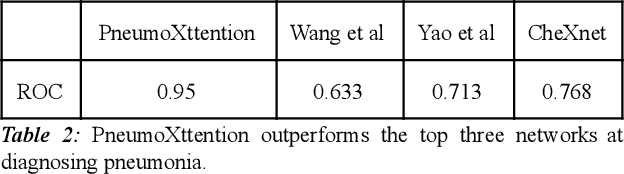
Abstract:Automatic Chest Radiograph X-ray (CXR) interpretation by machines is an important research topic of Artificial Intelligence. As part of my journey through the California Science Fair, I have developed an algorithm that can detect pneumonia from a CXR image to compensate for human fallibility. My algorithm, PneumoXttention, is an ensemble of two 13 layer convolutional neural network trained on the RSNA dataset, a dataset provided by the Radiological Society of North America, containing 26,684 frontal X-ray images split into the categories of pneumonia and no pneumonia. The dataset was annotated by many professional radiologists in North America. It achieved an impressive F1 score, 0.82, on the test set (20% random split of RSNA dataset) and completely compensated Human Radiologists on a random set of 25 test images drawn from RSNA and NIH. I don't have a direct comparison but Stanford's Chexnet has a F1 score of 0.435 on the NIH dataset for category Pneumonia.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge