Zhiqiang Hu
LLM-Adapters: An Adapter Family for Parameter-Efficient Fine-Tuning of Large Language Models
Apr 04, 2023Abstract:The success of large language models (LLMs), like GPT-3 and ChatGPT, has led to the development of numerous cost-effective and accessible alternatives that are created by fine-tuning open-access LLMs with task-specific data (e.g., ChatDoctor) or instruction data (e.g., Alpaca). Among the various fine-tuning methods, adapter-based parameter-efficient fine-tuning (PEFT) is undoubtedly one of the most attractive topics, as it only requires fine-tuning a few external parameters instead of the entire LLMs while achieving comparable or even better performance. To enable further research on PEFT methods of LLMs, this paper presents LLM-Adapters, an easy-to-use framework that integrates various adapters into LLMs and can execute these adapter-based PEFT methods of LLMs for different tasks. The framework includes state-of-the-art open-access LLMs such as LLaMA, BLOOM, OPT, and GPT-J, as well as widely used adapters such as Series adapter, Parallel adapter, and LoRA. The framework is designed to be research-friendly, efficient, modular, and extendable, allowing the integration of new adapters and the evaluation of them with new and larger-scale LLMs. Furthermore, to evaluate the effectiveness of adapters in LLMs-Adapters, we conduct experiments on six math reasoning datasets. The results demonstrate that using adapter-based PEFT in smaller-scale LLMs (7B) with few extra trainable parameters yields comparable, and in some cases superior, performance to that of powerful LLMs (175B) in zero-shot inference on simple math reasoning datasets. Overall, we provide a promising framework for fine-tuning large LLMs on downstream tasks. We believe the proposed LLMs-Adapters will advance adapter-based PEFT research, facilitate the deployment of research pipelines, and enable practical applications to real-world systems.
Adaptive Supervised PatchNCE Loss for Learning H&E-to-IHC Stain Translation with Inconsistent Groundtruth Image Pairs
Mar 10, 2023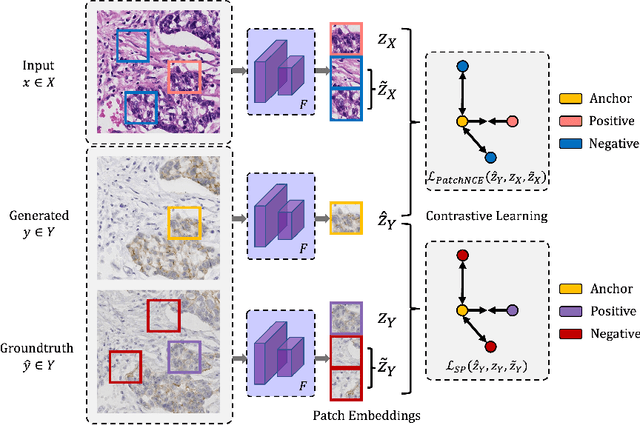



Abstract:Immunohistochemical (IHC) staining highlights the molecular information critical to diagnostics in tissue samples. However, compared to H&E staining, IHC staining can be much more expensive in terms of both labor and the laboratory equipment required. This motivates recent research that demonstrates that the correlations between the morphological information present in the H&E-stained slides and the molecular information in the IHC-stained slides can be used for H&E-to-IHC stain translation. However, due to a lack of pixel-perfect H&E-IHC groundtruth pairs, most existing methods have resorted to relying on expert annotations. To remedy this situation, we present a new loss function, Adaptive Supervised PatchNCE (ASP), to directly deal with the input to target inconsistencies in a proposed H&E-to-IHC image-to-image translation framework. The ASP loss is built upon a patch-based contrastive learning criterion, named Supervised PatchNCE (SP), and augments it further with weight scheduling to mitigate the negative impact of noisy supervision. Lastly, we introduce the Multi-IHC Stain Translation (MIST) dataset, which contains aligned H&E-IHC patches for 4 different IHC stains critical to breast cancer diagnosis. In our experiment, we demonstrate that our proposed method outperforms existing image-to-image translation methods for stain translation to multiple IHC stains. All of our code and datasets are available at https://github.com/lifangda01/AdaptiveSupervisedPatchNCE.
Are Current Task-oriented Dialogue Systems Able to Satisfy Impolite Users?
Oct 24, 2022



Abstract:Task-oriented dialogue (TOD) systems have assisted users on many tasks, including ticket booking and service inquiries. While existing TOD systems have shown promising performance in serving customer needs, these systems mostly assume that users would interact with the dialogue agent politely. This assumption is unrealistic as impatient or frustrated customers may also interact with TOD systems impolitely. This paper aims to address this research gap by investigating impolite users' effects on TOD systems. Specifically, we constructed an impolite dialogue corpus and conducted extensive experiments to evaluate the state-of-the-art TOD systems on our impolite dialogue corpus. Our experimental results show that existing TOD systems are unable to handle impolite user utterances. We also present a data augmentation method to improve TOD performance in impolite dialogues. Nevertheless, handling impolite dialogues remains a very challenging research task. We hope by releasing the impolite dialogue corpus and establishing the benchmark evaluations, more researchers are encouraged to investigate this new challenging research task.
Contrastive and Selective Hidden Embeddings for Medical Image Segmentation
Jan 21, 2022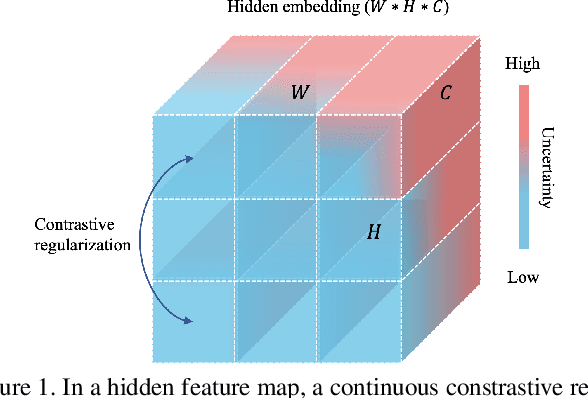
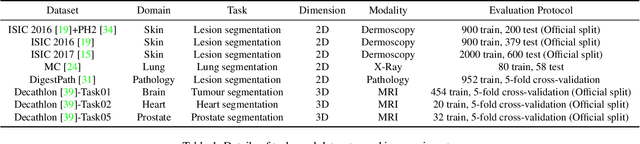
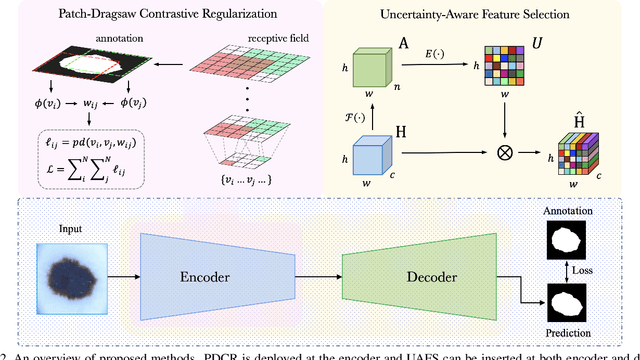
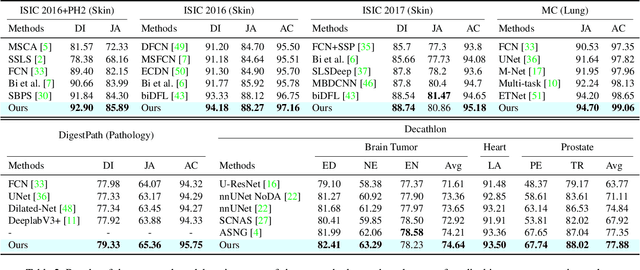
Abstract:Medical image segmentation has been widely recognized as a pivot procedure for clinical diagnosis, analysis, and treatment planning. However, the laborious and expensive annotation process lags down the speed of further advances. Contrastive learning-based weight pre-training provides an alternative by leveraging unlabeled data to learn a good representation. In this paper, we investigate how contrastive learning benefits the general supervised medical segmentation tasks. To this end, patch-dragsaw contrastive regularization (PDCR) is proposed to perform patch-level tugging and repulsing with the extent controlled by a continuous affinity score. And a new structure dubbed uncertainty-aware feature selection block (UAFS) is designed to perform the feature selection process, which can handle the learning target shift caused by minority features with high uncertainty. By plugging the proposed 2 modules into the existing segmentation architecture, we achieve state-of-the-art results across 8 public datasets from 6 domains. Newly designed modules further decrease the amount of training data to a quarter while achieving comparable, if not better, performances. From this perspective, we take the opposite direction of the original self/un-supervised contrastive learning by further excavating information contained within the label.
Improving Text Auto-Completion with Next Phrase Prediction
Sep 15, 2021
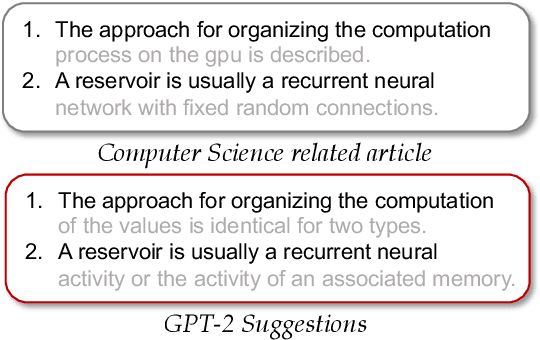

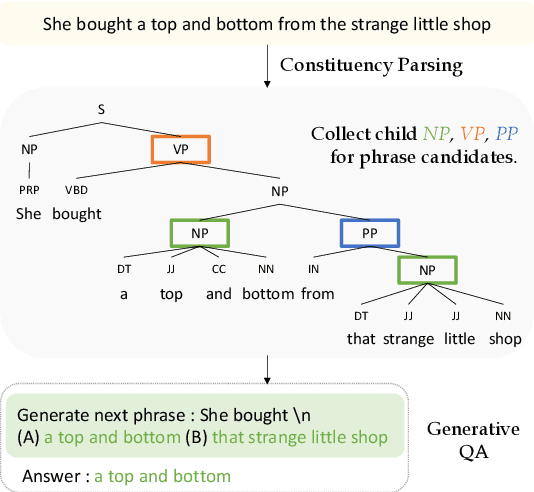
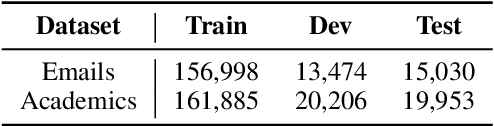
Abstract:Language models such as GPT-2 have performed well on constructing syntactically sound sentences for text auto-completion task. However, such models often require considerable training effort to adapt to specific writing domains (e.g., medical). In this paper, we propose an intermediate training strategy to enhance pre-trained language models' performance in the text auto-completion task and fastly adapt them to specific domains. Our strategy includes a novel self-supervised training objective called Next Phrase Prediction (NPP), which encourages a language model to complete the partial query with enriched phrases and eventually improve the model's text auto-completion performance. Preliminary experiments have shown that our approach is able to outperform the baselines in auto-completion for email and academic writing domains.
Syntax Matters! Syntax-Controlled in Text Style Transfer
Aug 12, 2021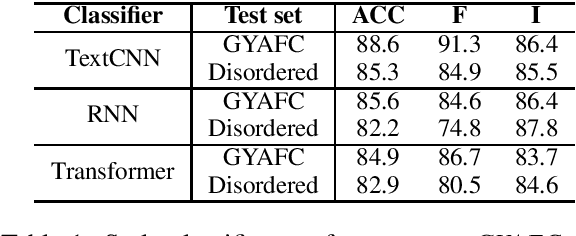
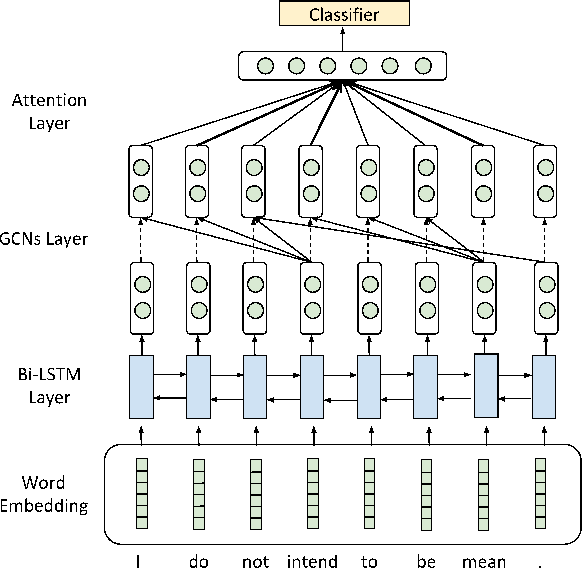
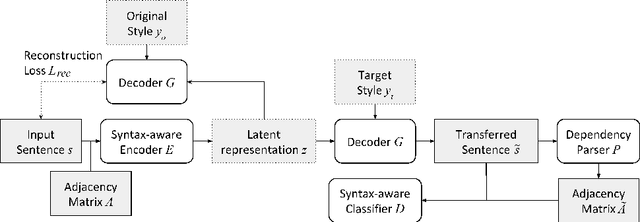
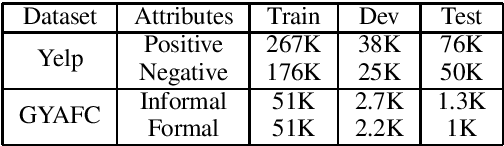
Abstract:Existing text style transfer (TST) methods rely on style classifiers to disentangle the text's content and style attributes for text style transfer. While the style classifier plays a critical role in existing TST methods, there is no known investigation on its effect on the TST methods. In this paper, we conduct an empirical study on the limitations of the style classifiers used in existing TST methods. We demonstrate that the existing style classifiers cannot learn sentence syntax effectively and ultimately worsen existing TST models' performance. To address this issue, we propose a novel Syntax-Aware Controllable Generation (SACG) model, which includes a syntax-aware style classifier that ensures learned style latent representations effectively capture the syntax information for TST. Through extensive experiments on two popular TST tasks, we show that our proposed method significantly outperforms the state-of-the-art methods. Our case studies have also demonstrated SACG's ability to generate fluent target-style sentences that preserved the original content.
Multi-frame Collaboration for Effective Endoscopic Video Polyp Detection via Spatial-Temporal Feature Transformation
Jul 08, 2021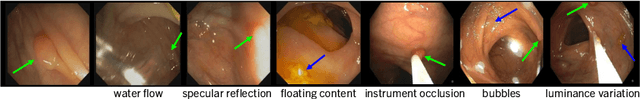
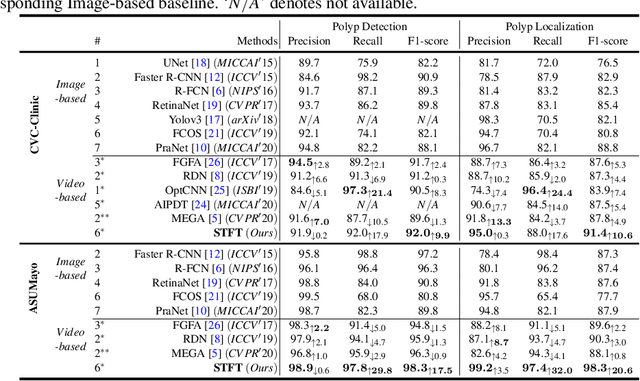
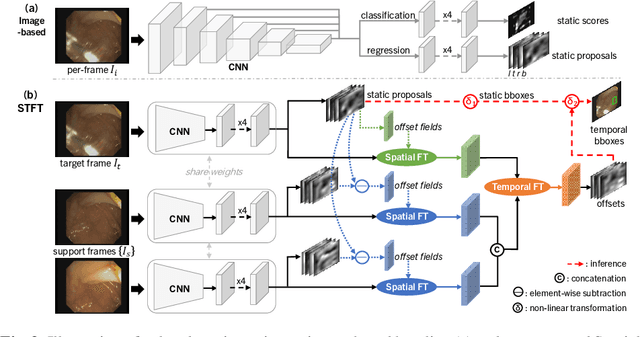

Abstract:Precise localization of polyp is crucial for early cancer screening in gastrointestinal endoscopy. Videos given by endoscopy bring both richer contextual information as well as more challenges than still images. The camera-moving situation, instead of the common camera-fixed-object-moving one, leads to significant background variation between frames. Severe internal artifacts (e.g. water flow in the human body, specular reflection by tissues) can make the quality of adjacent frames vary considerately. These factors hinder a video-based model to effectively aggregate features from neighborhood frames and give better predictions. In this paper, we present Spatial-Temporal Feature Transformation (STFT), a multi-frame collaborative framework to address these issues. Spatially, STFT mitigates inter-frame variations in the camera-moving situation with feature alignment by proposal-guided deformable convolutions. Temporally, STFT proposes a channel-aware attention module to simultaneously estimate the quality and correlation of adjacent frames for adaptive feature aggregation. Empirical studies and superior results demonstrate the effectiveness and stability of our method. For example, STFT improves the still image baseline FCOS by 10.6% and 20.6% on the comprehensive F1-score of the polyp localization task in CVC-Clinic and ASUMayo datasets, respectively, and outperforms the state-of-the-art video-based method by 3.6% and 8.0%, respectively. Code is available at \url{https://github.com/lingyunwu14/STFT}.
Mixed Supervision Learning for Whole Slide Image Classification
Jul 05, 2021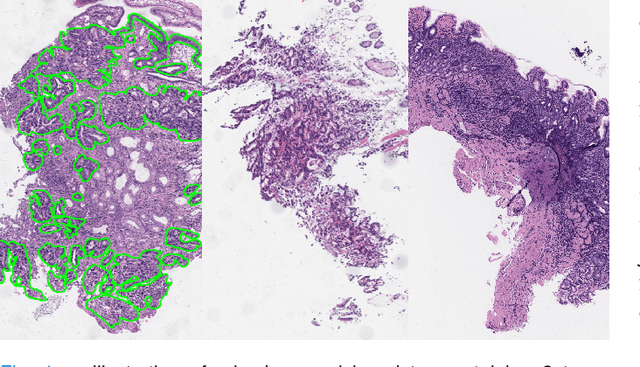
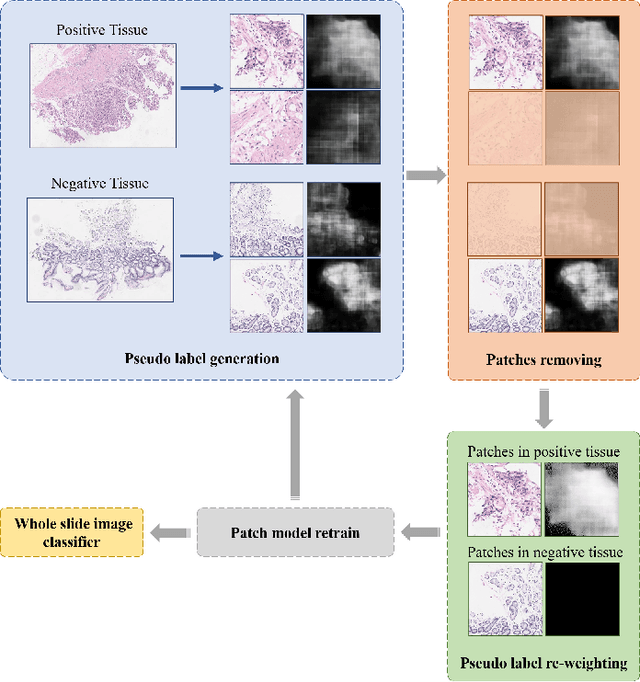
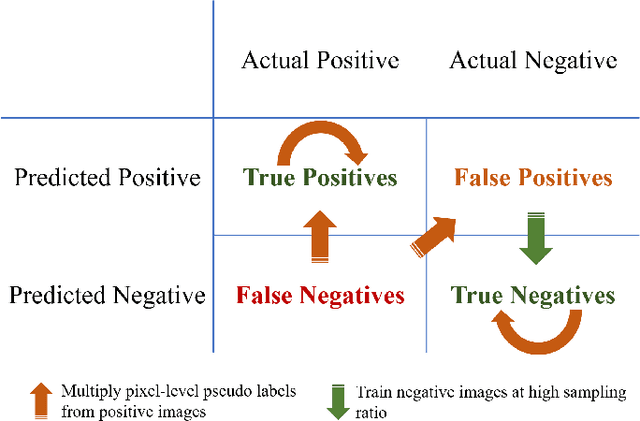
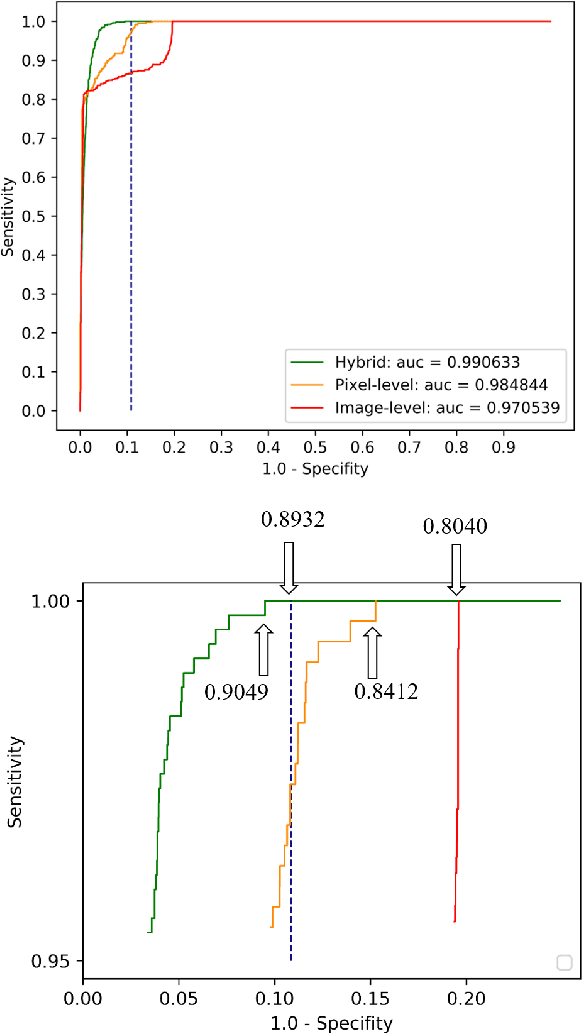
Abstract:Weak supervision learning on classification labels has demonstrated high performance in various tasks. When a few pixel-level fine annotations are also affordable, it is natural to leverage both of the pixel-level (e.g., segmentation) and image level (e.g., classification) annotation to further improve the performance. In computational pathology, however, such weak or mixed supervision learning is still a challenging task, since the high resolution of whole slide images makes it unattainable to perform end-to-end training of classification models. An alternative approach is to analyze such data by patch-base model training, i.e., using self-supervised learning to generate pixel-level pseudo labels for patches. However, such methods usually have model drifting issues, i.e., hard to converge, because the noise accumulates during the self-training process. To handle those problems, we propose a mixed supervision learning framework for super high-resolution images to effectively utilize their various labels (e.g., sufficient image-level coarse annotations and a few pixel-level fine labels). During the patch training stage, this framework can make use of coarse image-level labels to refine self-supervised learning and generate high-quality pixel-level pseudo labels. A comprehensive strategy is proposed to suppress pixel-level false positives and false negatives. Three real-world datasets with very large number of images (i.e., more than 10,000 whole slide images) and various types of labels are used to evaluate the effectiveness of mixed supervision learning. We reduced the false positive rate by around one third compared to state of the art while retaining 100% sensitivity, in the task of image-level classification.
Learning Unknown from Correlations: Graph Neural Network for Inter-novel-protein Interaction Prediction
Jun 01, 2021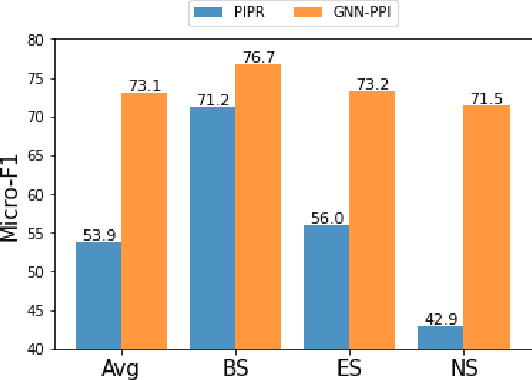
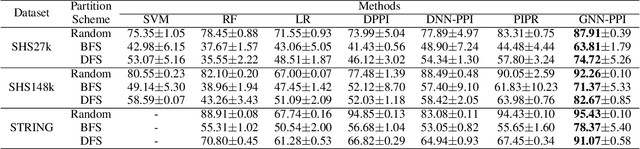
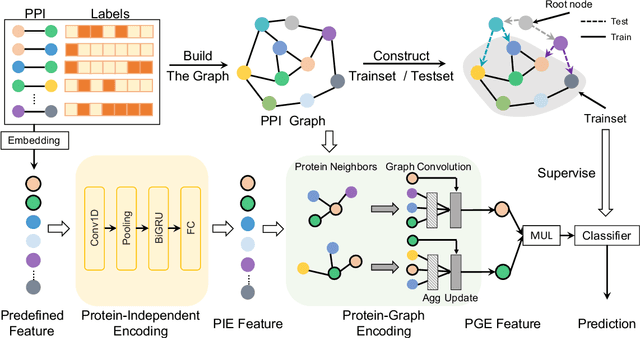
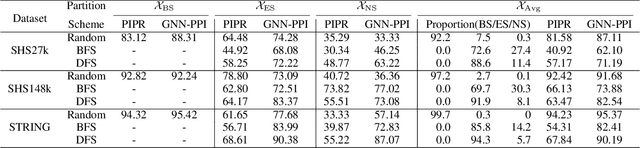
Abstract:The study of multi-type Protein-Protein Interaction (PPI) is fundamental for understanding biological processes from a systematic perspective and revealing disease mechanisms. Existing methods suffer from significant performance degradation when tested in unseen dataset. In this paper, we investigate the problem and find that it is mainly attributed to the poor performance for inter-novel-protein interaction prediction. However, current evaluations overlook the inter-novel-protein interactions, and thus fail to give an instructive assessment. As a result, we propose to address the problem from both the evaluation and the methodology. Firstly, we design a new evaluation framework that fully respects the inter-novel-protein interactions and gives consistent assessment across datasets. Secondly, we argue that correlations between proteins must provide useful information for analysis of novel proteins, and based on this, we propose a graph neural network based method (GNN-PPI) for better inter-novel-protein interaction prediction. Experimental results on real-world datasets of different scales demonstrate that GNN-PPI significantly outperforms state-of-the-art PPI prediction methods, especially for the inter-novel-protein interaction prediction.
DeepStyle: User Style Embedding for Authorship Attribution of Short Texts
Mar 14, 2021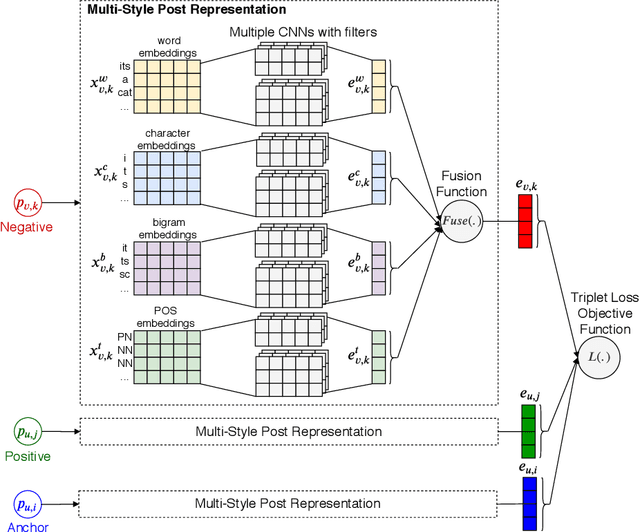
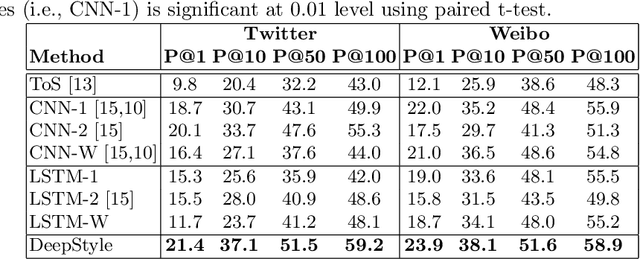
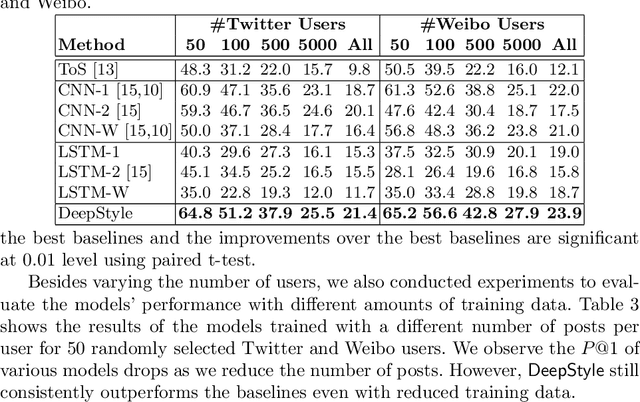
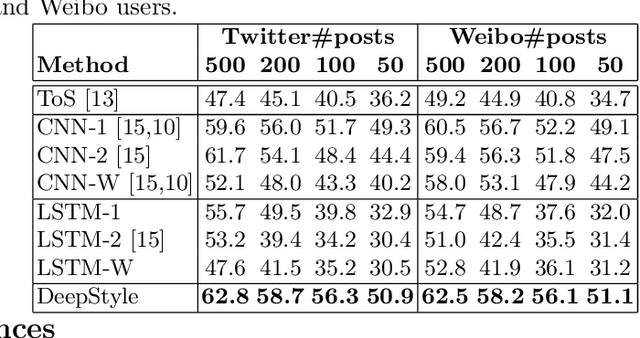
Abstract:Authorship attribution (AA), which is the task of finding the owner of a given text, is an important and widely studied research topic with many applications. Recent works have shown that deep learning methods could achieve significant accuracy improvement for the AA task. Nevertheless, most of these proposed methods represent user posts using a single type of feature (e.g., word bi-grams) and adopt a text classification approach to address the task. Furthermore, these methods offer very limited explainability of the AA results. In this paper, we address these limitations by proposing DeepStyle, a novel embedding-based framework that learns the representations of users' salient writing styles. We conduct extensive experiments on two real-world datasets from Twitter and Weibo. Our experiment results show that DeepStyle outperforms the state-of-the-art baselines on the AA task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge