Yujia Li
Compressed domain vibration detection and classification for distributed acoustic sensing
Dec 27, 2022



Abstract:Distributed acoustic sensing (DAS) is a novel enabling technology that can turn existing fibre optic networks to distributed acoustic sensors. However, it faces the challenges of transmitting, storing, and processing massive streams of data which are orders of magnitude larger than that collected from point sensors. The gap between intensive data generated by DAS and modern computing system with limited reading/writing speed and storage capacity imposes restrictions on many applications. Compressive sensing (CS) is a revolutionary signal acquisition method that allows a signal to be acquired and reconstructed with significantly fewer samples than that required by Nyquist-Shannon theorem. Though the data size is greatly reduced in the sampling stage, the reconstruction of the compressed data is however time and computation consuming. To address this challenge, we propose to map the feature extractor from Nyquist-domain to compressed-domain and therefore vibration detection and classification can be directly implemented in compressed-domain. The measured results show that our framework can be used to reduce the transmitted data size by 70% while achieves 99.4% true positive rate (TPR) and 0.04% false positive rate (TPR) along 5 km sensing fibre and 95.05% classification accuracy on a 5-class classification task.
Scaling Language Models: Methods, Analysis & Insights from Training Gopher
Dec 08, 2021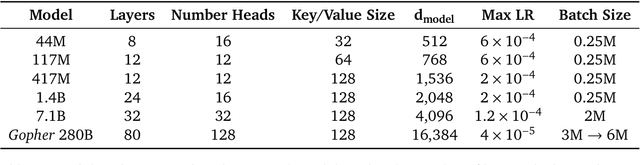
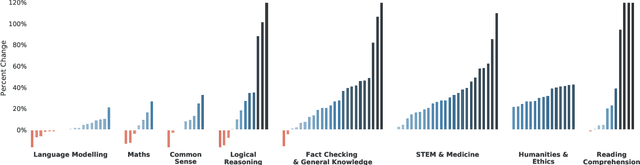
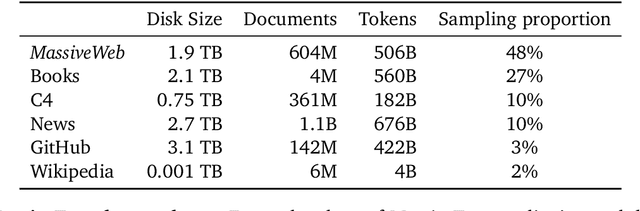
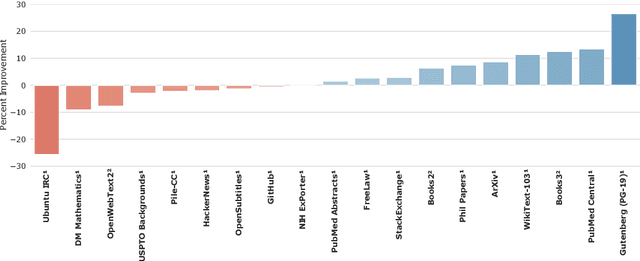
Abstract:Language modelling provides a step towards intelligent communication systems by harnessing large repositories of written human knowledge to better predict and understand the world. In this paper, we present an analysis of Transformer-based language model performance across a wide range of model scales -- from models with tens of millions of parameters up to a 280 billion parameter model called Gopher. These models are evaluated on 152 diverse tasks, achieving state-of-the-art performance across the majority. Gains from scale are largest in areas such as reading comprehension, fact-checking, and the identification of toxic language, but logical and mathematical reasoning see less benefit. We provide a holistic analysis of the training dataset and model's behaviour, covering the intersection of model scale with bias and toxicity. Finally we discuss the application of language models to AI safety and the mitigation of downstream harms.
ETA Prediction with Graph Neural Networks in Google Maps
Aug 25, 2021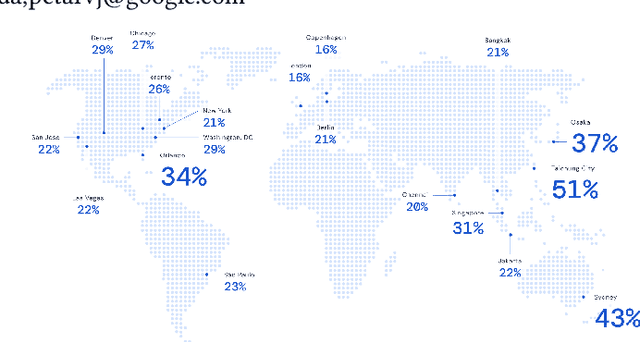
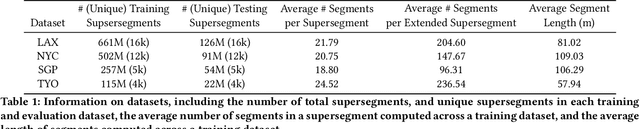
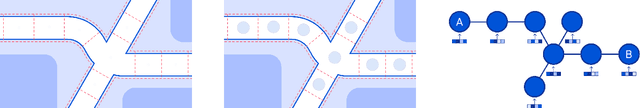
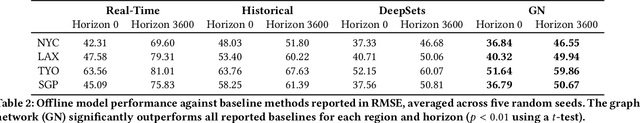
Abstract:Travel-time prediction constitutes a task of high importance in transportation networks, with web mapping services like Google Maps regularly serving vast quantities of travel time queries from users and enterprises alike. Further, such a task requires accounting for complex spatiotemporal interactions (modelling both the topological properties of the road network and anticipating events -- such as rush hours -- that may occur in the future). Hence, it is an ideal target for graph representation learning at scale. Here we present a graph neural network estimator for estimated time of arrival (ETA) which we have deployed in production at Google Maps. While our main architecture consists of standard GNN building blocks, we further detail the usage of training schedule methods such as MetaGradients in order to make our model robust and production-ready. We also provide prescriptive studies: ablating on various architectural decisions and training regimes, and qualitative analyses on real-world situations where our model provides a competitive edge. Our GNN proved powerful when deployed, significantly reducing negative ETA outcomes in several regions compared to the previous production baseline (40+% in cities like Sydney).
WikiGraphs: A Wikipedia Text - Knowledge Graph Paired Dataset
Jul 20, 2021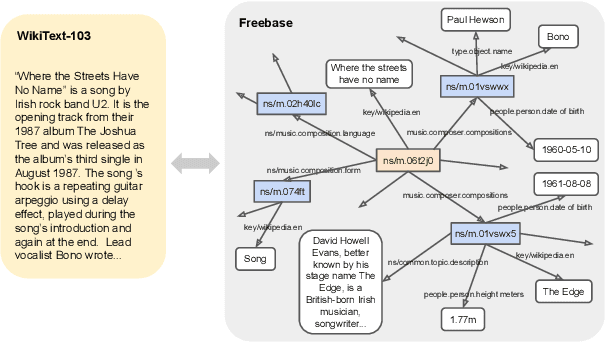
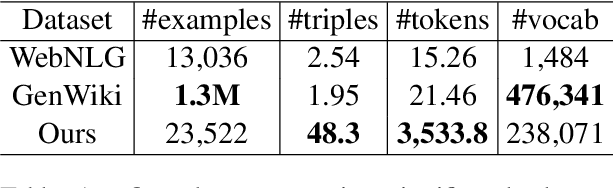
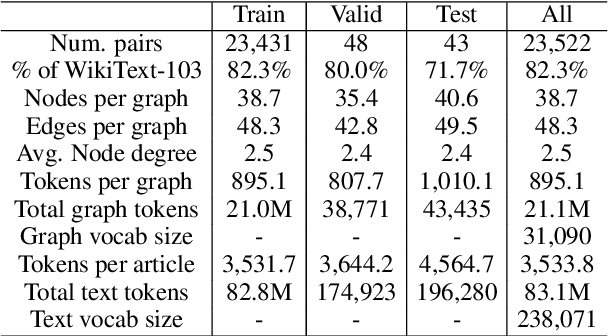
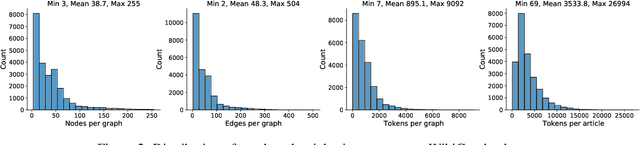
Abstract:We present a new dataset of Wikipedia articles each paired with a knowledge graph, to facilitate the research in conditional text generation, graph generation and graph representation learning. Existing graph-text paired datasets typically contain small graphs and short text (1 or few sentences), thus limiting the capabilities of the models that can be learned on the data. Our new dataset WikiGraphs is collected by pairing each Wikipedia article from the established WikiText-103 benchmark (Merity et al., 2016) with a subgraph from the Freebase knowledge graph (Bollacker et al., 2008). This makes it easy to benchmark against other state-of-the-art text generative models that are capable of generating long paragraphs of coherent text. Both the graphs and the text data are of significantly larger scale compared to prior graph-text paired datasets. We present baseline graph neural network and transformer model results on our dataset for 3 tasks: graph -> text generation, graph -> text retrieval and text -> graph retrieval. We show that better conditioning on the graph provides gains in generation and retrieval quality but there is still large room for improvement.
Computer-Aided Design as Language
May 06, 2021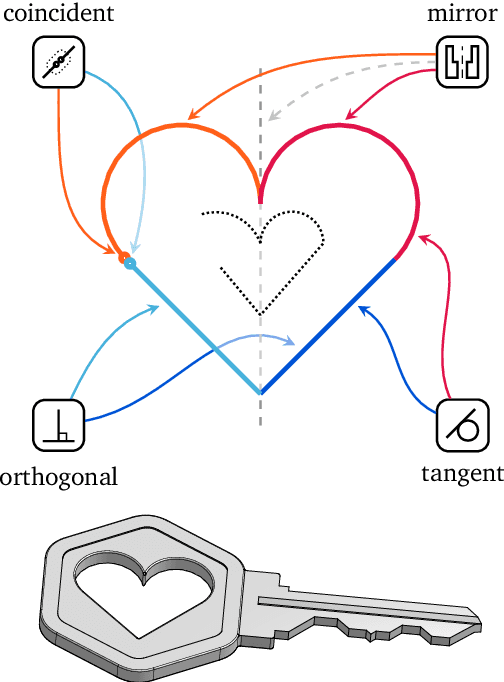
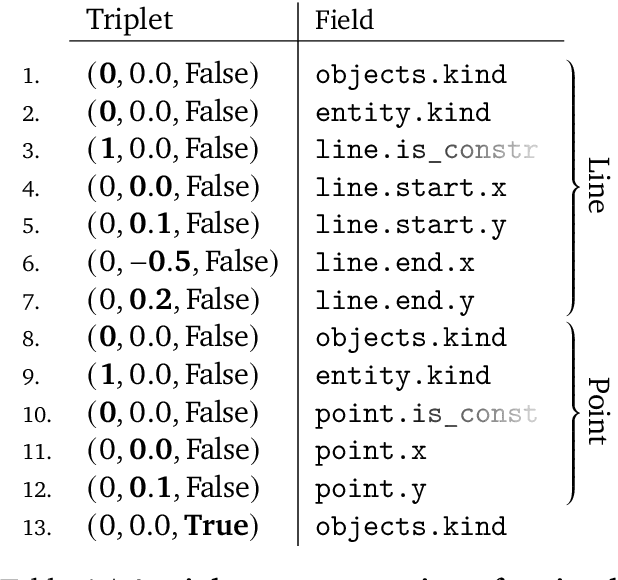
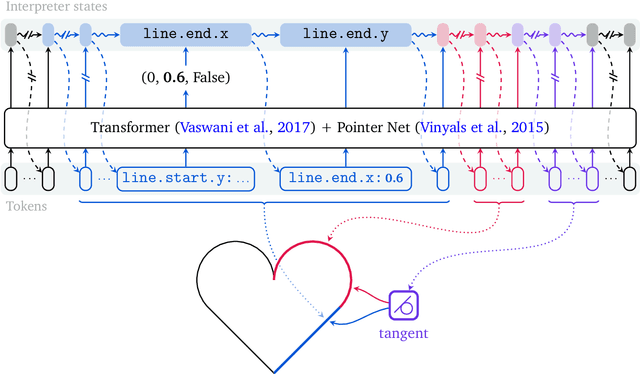
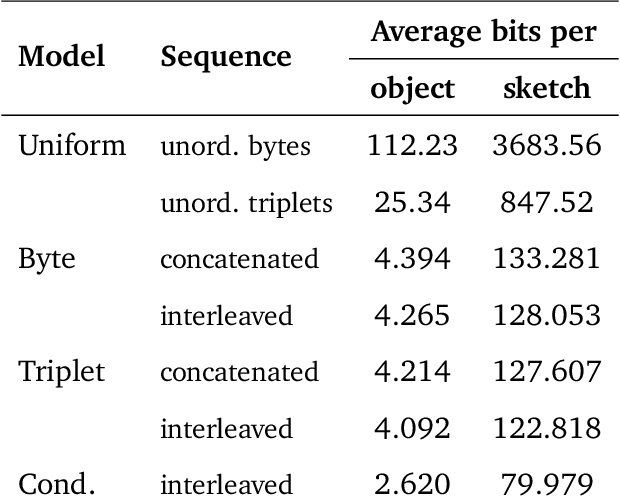
Abstract:Computer-Aided Design (CAD) applications are used in manufacturing to model everything from coffee mugs to sports cars. These programs are complex and require years of training and experience to master. A component of all CAD models particularly difficult to make are the highly structured 2D sketches that lie at the heart of every 3D construction. In this work, we propose a machine learning model capable of automatically generating such sketches. Through this, we pave the way for developing intelligent tools that would help engineers create better designs with less effort. Our method is a combination of a general-purpose language modeling technique alongside an off-the-shelf data serialization protocol. We show that our approach has enough flexibility to accommodate the complexity of the domain and performs well for both unconditional synthesis and image-to-sketch translation.
Solving Mixed Integer Programs Using Neural Networks
Dec 23, 2020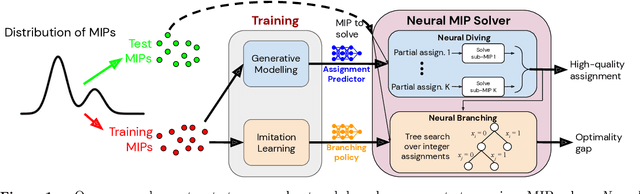
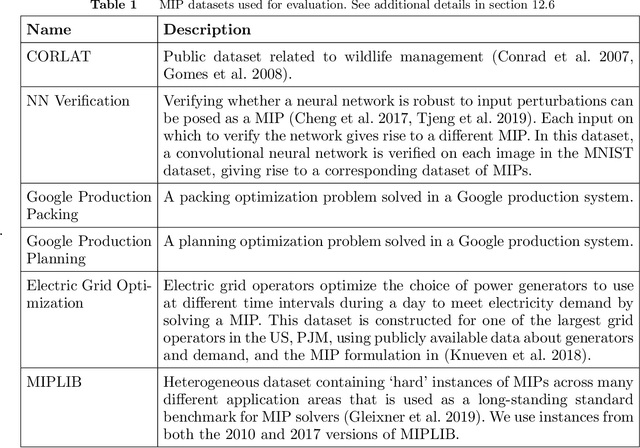
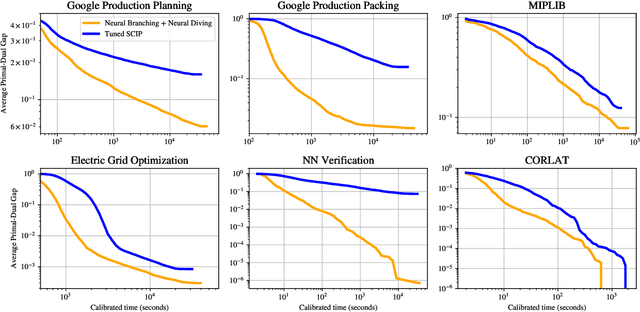
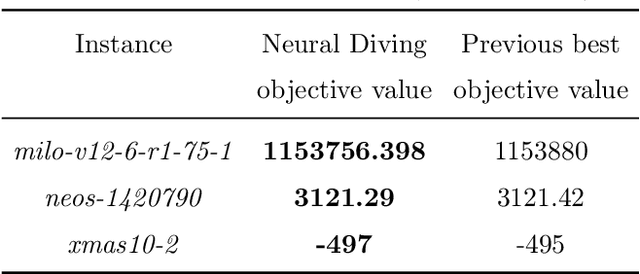
Abstract:Mixed Integer Programming (MIP) solvers rely on an array of sophisticated heuristics developed with decades of research to solve large-scale MIP instances encountered in practice. Machine learning offers to automatically construct better heuristics from data by exploiting shared structure among instances in the data. This paper applies learning to the two key sub-tasks of a MIP solver, generating a high-quality joint variable assignment, and bounding the gap in objective value between that assignment and an optimal one. Our approach constructs two corresponding neural network-based components, Neural Diving and Neural Branching, to use in a base MIP solver such as SCIP. Neural Diving learns a deep neural network to generate multiple partial assignments for its integer variables, and the resulting smaller MIPs for un-assigned variables are solved with SCIP to construct high quality joint assignments. Neural Branching learns a deep neural network to make variable selection decisions in branch-and-bound to bound the objective value gap with a small tree. This is done by imitating a new variant of Full Strong Branching we propose that scales to large instances using GPUs. We evaluate our approach on six diverse real-world datasets, including two Google production datasets and MIPLIB, by training separate neural networks on each. Most instances in all the datasets combined have $10^3-10^6$ variables and constraints after presolve, which is significantly larger than previous learning approaches. Comparing solvers with respect to primal-dual gap averaged over a held-out set of instances, the learning-augmented SCIP is 2x to 10x better on all datasets except one on which it is $10^5$x better, at large time limits. To the best of our knowledge, ours is the first learning approach to demonstrate such large improvements over SCIP on both large-scale real-world application datasets and MIPLIB.
Strong Generalization and Efficiency in Neural Programs
Jul 08, 2020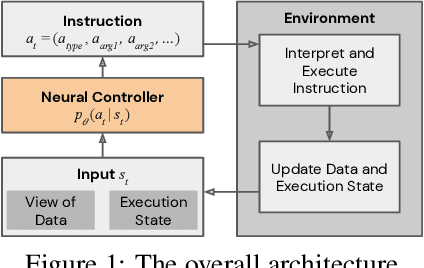
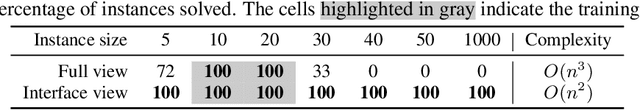
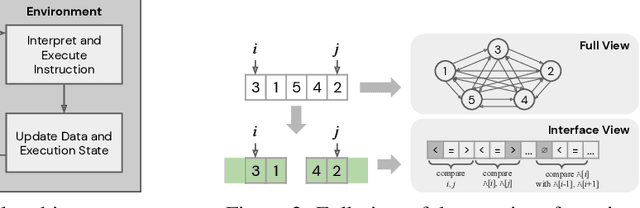
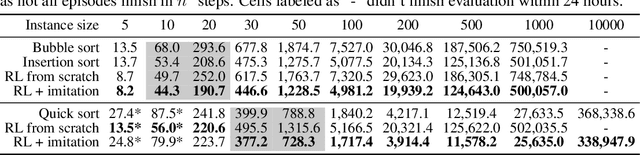
Abstract:We study the problem of learning efficient algorithms that strongly generalize in the framework of neural program induction. By carefully designing the input / output interfaces of the neural model and through imitation, we are able to learn models that produce correct results for arbitrary input sizes, achieving strong generalization. Moreover, by using reinforcement learning, we optimize for program efficiency metrics, and discover new algorithms that surpass the teacher used in imitation. With this, our approach can learn to outperform custom-written solutions for a variety of problems, as we tested it on sorting, searching in ordered lists and the NP-complete 0/1 knapsack problem, which sets a notable milestone in the field of Neural Program Induction. As highlights, our learned model can perform sorting perfectly on any input data size we tested on, with $O(n log n)$ complexity, whilst outperforming hand-coded algorithms, including quick sort, in number of operations even for list sizes far beyond those seen during training.
Scalable Deep Generative Modeling for Sparse Graphs
Jun 28, 2020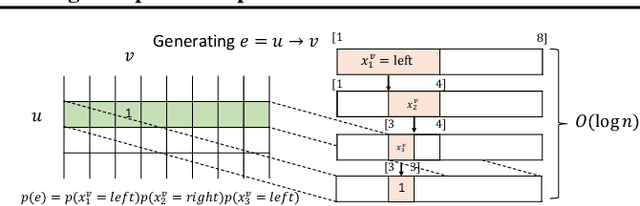
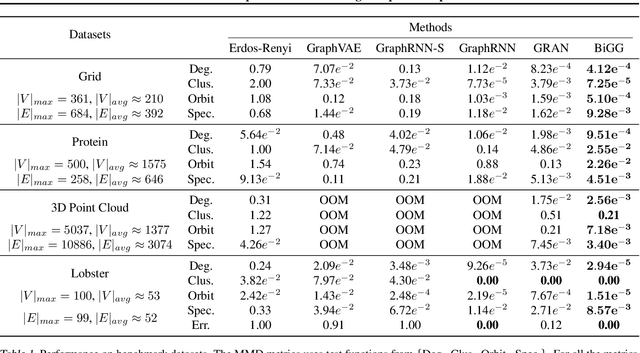
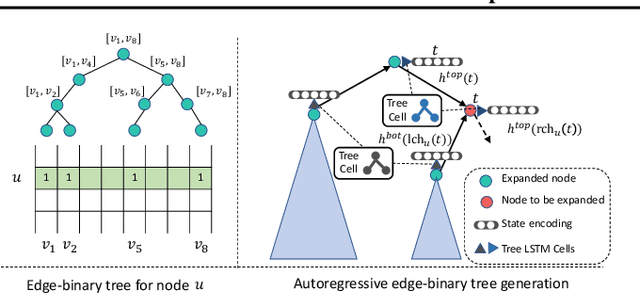
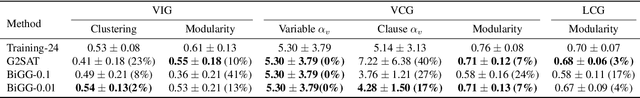
Abstract:Learning graph generative models is a challenging task for deep learning and has wide applicability to a range of domains like chemistry, biology and social science. However current deep neural methods suffer from limited scalability: for a graph with $n$ nodes and $m$ edges, existing deep neural methods require $\Omega(n^2)$ complexity by building up the adjacency matrix. On the other hand, many real world graphs are actually sparse in the sense that $m\ll n^2$. Based on this, we develop a novel autoregressive model, named BiGG, that utilizes this sparsity to avoid generating the full adjacency matrix, and importantly reduces the graph generation time complexity to $O((n + m)\log n)$. Furthermore, during training this autoregressive model can be parallelized with $O(\log n)$ synchronization stages, which makes it much more efficient than other autoregressive models that require $\Omega(n)$. Experiments on several benchmarks show that the proposed approach not only scales to orders of magnitude larger graphs than previously possible with deep autoregressive graph generative models, but also yields better graph generation quality.
A sparse negative binomial mixture model for clustering RNA-seq count data
Dec 05, 2019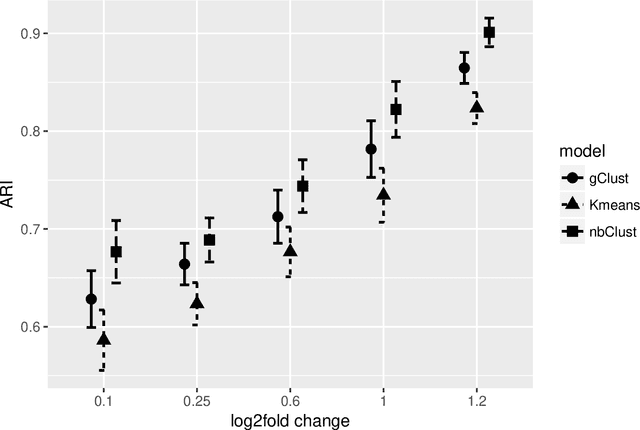
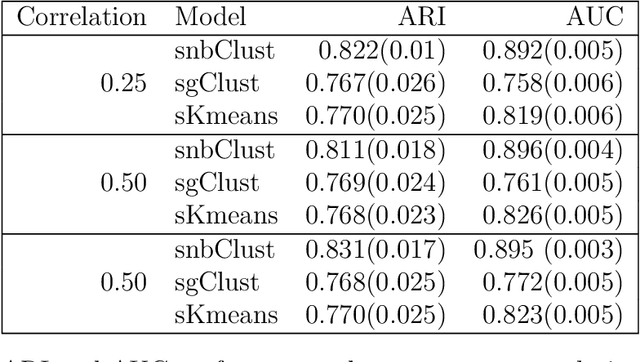
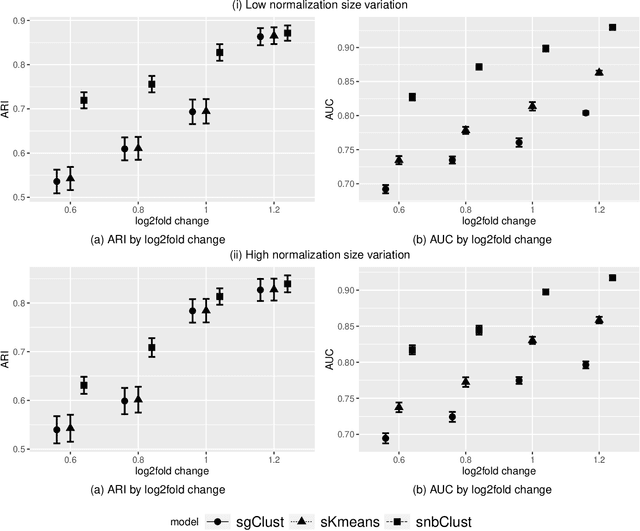

Abstract:Clustering with variable selection is a challenging but critical task for modern small-n-large-p data. Existing methods based on Gaussian mixture models or sparse K-means provide solutions to continuous data. With the prevalence of RNA-seq technology and lack of count data modeling for clustering, the current practice is to normalize count expression data into continuous measures and apply existing models with Gaussian assumption. In this paper, we develop a negative binomial mixture model with gene regularization to cluster samples (small $n$) with high-dimensional gene features (large $p$). EM algorithm and Bayesian information criterion are used for inference and determining tuning parameters. The method is compared with sparse Gaussian mixture model and sparse K-means using extensive simulations and two real transcriptomic applications in breast cancer and rat brain studies. The result shows superior performance of the proposed count data model in clustering accuracy, feature selection and biological interpretation by pathway enrichment analysis.
Prioritized Unit Propagation with Periodic Resetting is (Almost) All You Need for Random SAT Solving
Dec 04, 2019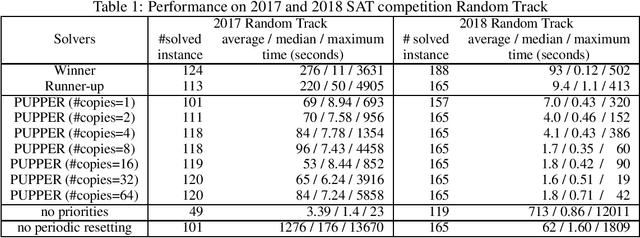
Abstract:We propose prioritized unit propagation with periodic resetting, which is a simple but surprisingly effective algorithm for solving random SAT instances that are meant to be hard. In particular, an evaluation on the Random Track of the 2017 and 2018 SAT competitions shows that a basic prototype of this simple idea already ranks at second place in both years. We share this observation in the hope that it helps the SAT community better understand the hardness of random instances used in competitions and inspire other interesting ideas on SAT solving.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge