Xinxing Cheng
Plasticine: A Traceable Diffusion Model for Medical Image Translation
Dec 20, 2025Abstract:Domain gaps arising from variations in imaging devices and population distributions pose significant challenges for machine learning in medical image analysis. Existing image-to-image translation methods primarily aim to learn mappings between domains, often generating diverse synthetic data with variations in anatomical scale and shape, but they usually overlook spatial correspondence during the translation process. For clinical applications, traceability, defined as the ability to provide pixel-level correspondences between original and translated images, is equally important. This property enhances clinical interpretability but has been largely overlooked in previous approaches. To address this gap, we propose Plasticine, which is, to the best of our knowledge, the first end-to-end image-to-image translation framework explicitly designed with traceability as a core objective. Our method combines intensity translation and spatial transformation within a denoising diffusion framework. This design enables the generation of synthetic images with interpretable intensity transitions and spatially coherent deformations, supporting pixel-wise traceability throughout the translation process.
TraceTrans: Translation and Spatial Tracing for Surgical Prediction
Oct 25, 2025Abstract:Image-to-image translation models have achieved notable success in converting images across visual domains and are increasingly used for medical tasks such as predicting post-operative outcomes and modeling disease progression. However, most existing methods primarily aim to match the target distribution and often neglect spatial correspondences between the source and translated images. This limitation can lead to structural inconsistencies and hallucinations, undermining the reliability and interpretability of the predictions. These challenges are accentuated in clinical applications by the stringent requirement for anatomical accuracy. In this work, we present TraceTrans, a novel deformable image translation model designed for post-operative prediction that generates images aligned with the target distribution while explicitly revealing spatial correspondences with the pre-operative input. The framework employs an encoder for feature extraction and dual decoders for predicting spatial deformations and synthesizing the translated image. The predicted deformation field imposes spatial constraints on the generated output, ensuring anatomical consistency with the source. Extensive experiments on medical cosmetology and brain MRI datasets demonstrate that TraceTrans delivers accurate and interpretable post-operative predictions, highlighting its potential for reliable clinical deployment.
MCM: Mamba-based Cardiac Motion Tracking using Sequential Images in MRI
Jul 23, 2025Abstract:Myocardial motion tracking is important for assessing cardiac function and diagnosing cardiovascular diseases, for which cine cardiac magnetic resonance (CMR) has been established as the gold standard imaging modality. Many existing methods learn motion from single image pairs consisting of a reference frame and a randomly selected target frame from the cardiac cycle. However, these methods overlook the continuous nature of cardiac motion and often yield inconsistent and non-smooth motion estimations. In this work, we propose a novel Mamba-based cardiac motion tracking network (MCM) that explicitly incorporates target image sequence from the cardiac cycle to achieve smooth and temporally consistent motion tracking. By developing a bi-directional Mamba block equipped with a bi-directional scanning mechanism, our method facilitates the estimation of plausible deformation fields. With our proposed motion decoder that integrates motion information from frames adjacent to the target frame, our method further enhances temporal coherence. Moreover, by taking advantage of Mamba's structured state-space formulation, the proposed method learns the continuous dynamics of the myocardium from sequential images without increasing computational complexity. We evaluate the proposed method on two public datasets. The experimental results demonstrate that the proposed method quantitatively and qualitatively outperforms both conventional and state-of-the-art learning-based cardiac motion tracking methods. The code is available at https://github.com/yjh-0104/MCM.
SACB-Net: Spatial-awareness Convolutions for Medical Image Registration
Mar 25, 2025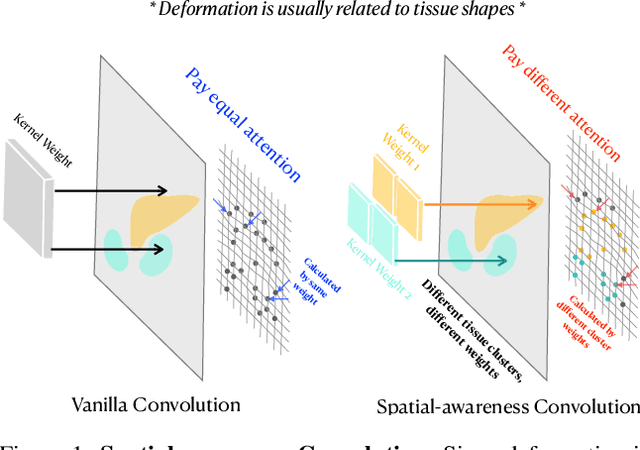
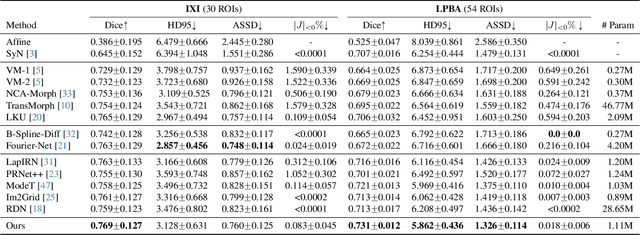
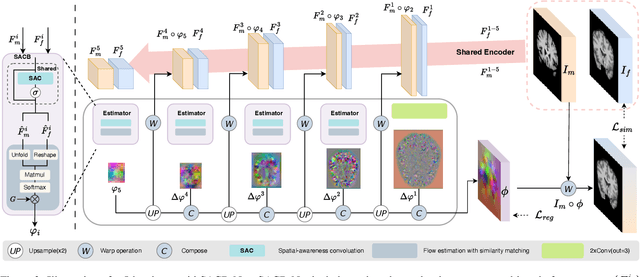
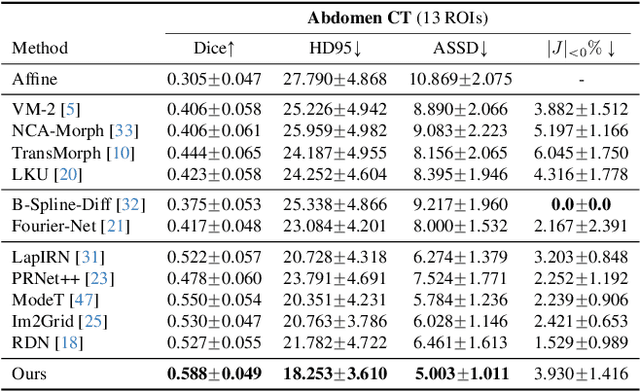
Abstract:Deep learning-based image registration methods have shown state-of-the-art performance and rapid inference speeds. Despite these advances, many existing approaches fall short in capturing spatially varying information in non-local regions of feature maps due to the reliance on spatially-shared convolution kernels. This limitation leads to suboptimal estimation of deformation fields. In this paper, we propose a 3D Spatial-Awareness Convolution Block (SACB) to enhance the spatial information within feature representations. Our SACB estimates the spatial clusters within feature maps by leveraging feature similarity and subsequently parameterizes the adaptive convolution kernels across diverse regions. This adaptive mechanism generates the convolution kernels (weights and biases) tailored to spatial variations, thereby enabling the network to effectively capture spatially varying information. Building on SACB, we introduce a pyramid flow estimator (named SACB-Net) that integrates SACBs to facilitate multi-scale flow composition, particularly addressing large deformations. Experimental results on the brain IXI and LPBA datasets as well as Abdomen CT datasets demonstrate the effectiveness of SACB and the superiority of SACB-Net over the state-of-the-art learning-based registration methods. The code is available at https://github.com/x-xc/SACB_Net .
WiNet: Wavelet-based Incremental Learning for Efficient Medical Image Registration
Jul 18, 2024Abstract:Deep image registration has demonstrated exceptional accuracy and fast inference. Recent advances have adopted either multiple cascades or pyramid architectures to estimate dense deformation fields in a coarse-to-fine manner. However, due to the cascaded nature and repeated composition/warping operations on feature maps, these methods negatively increase memory usage during training and testing. Moreover, such approaches lack explicit constraints on the learning process of small deformations at different scales, thus lacking explainability. In this study, we introduce a model-driven WiNet that incrementally estimates scale-wise wavelet coefficients for the displacement/velocity field across various scales, utilizing the wavelet coefficients derived from the original input image pair. By exploiting the properties of the wavelet transform, these estimated coefficients facilitate the seamless reconstruction of a full-resolution displacement/velocity field via our devised inverse discrete wavelet transform (IDWT) layer. This approach avoids the complexities of cascading networks or composition operations, making our WiNet an explainable and efficient competitor with other coarse-to-fine methods. Extensive experimental results from two 3D datasets show that our WiNet is accurate and GPU efficient. The code is available at https://github.com/x-xc/WiNet .
Decoder-Only Image Registration
Feb 05, 2024



Abstract:In unsupervised medical image registration, the predominant approaches involve the utilization of a encoder-decoder network architecture, allowing for precise prediction of dense, full-resolution displacement fields from given paired images. Despite its widespread use in the literature, we argue for the necessity of making both the encoder and decoder learnable in such an architecture. For this, we propose a novel network architecture, termed LessNet in this paper, which contains only a learnable decoder, while entirely omitting the utilization of a learnable encoder. LessNet substitutes the learnable encoder with simple, handcrafted features, eliminating the need to learn (optimize) network parameters in the encoder altogether. Consequently, this leads to a compact, efficient, and decoder-only architecture for 3D medical image registration. Evaluated on two publicly available brain MRI datasets, we demonstrate that our decoder-only LessNet can effectively and efficiently learn both dense displacement and diffeomorphic deformation fields in 3D. Furthermore, our decoder-only LessNet can achieve comparable registration performance to state-of-the-art methods such as VoxelMorph and TransMorph, while requiring significantly fewer computational resources. Our code and pre-trained models are available at https://github.com/xi-jia/LessNet.
Fourier-Net: Fast Image Registration with Band-limited Deformation
Nov 29, 2022



Abstract:Unsupervised image registration commonly adopts U-Net style networks to predict dense displacement fields in the full-resolution spatial domain. For high-resolution volumetric image data, this process is however resource intensive and time-consuming. To tackle this problem, we propose the Fourier-Net, replacing the expansive path in a U-Net style network with a parameter-free model-driven decoder. Specifically, instead of our Fourier-Net learning to output a full-resolution displacement field in the spatial domain, we learn its low-dimensional representation in a band-limited Fourier domain. This representation is then decoded by our devised model-driven decoder (consisting of a zero padding layer and an inverse discrete Fourier transform layer) to the dense, full-resolution displacement field in the spatial domain. These changes allow our unsupervised Fourier-Net to contain fewer parameters and computational operations, resulting in faster inference speeds. Fourier-Net is then evaluated on two public 3D brain datasets against various state-of-the-art approaches. For example, when compared to a recent transformer-based method, i.e., TransMorph, our Fourier-Net, only using 0.22$\%$ of its parameters and 6.66$\%$ of the mult-adds, achieves a 0.6\% higher Dice score and an 11.48$\times$ faster inference speed. Code is available at \url{https://github.com/xi-jia/Fourier-Net}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge