Wenqi Li
Exploring Speech Pattern Disorders in Autism using Machine Learning
May 03, 2024



Abstract:Diagnosing autism spectrum disorder (ASD) by identifying abnormal speech patterns from examiner-patient dialogues presents significant challenges due to the subtle and diverse manifestations of speech-related symptoms in affected individuals. This study presents a comprehensive approach to identify distinctive speech patterns through the analysis of examiner-patient dialogues. Utilizing a dataset of recorded dialogues, we extracted 40 speech-related features, categorized into frequency, zero-crossing rate, energy, spectral characteristics, Mel Frequency Cepstral Coefficients (MFCCs), and balance. These features encompass various aspects of speech such as intonation, volume, rhythm, and speech rate, reflecting the complex nature of communicative behaviors in ASD. We employed machine learning for both classification and regression tasks to analyze these speech features. The classification model aimed to differentiate between ASD and non-ASD cases, achieving an accuracy of 87.75%. Regression models were developed to predict speech pattern related variables and a composite score from all variables, facilitating a deeper understanding of the speech dynamics associated with ASD. The effectiveness of machine learning in interpreting intricate speech patterns and the high classification accuracy underscore the potential of computational methods in supporting the diagnostic processes for ASD. This approach not only aids in early detection but also contributes to personalized treatment planning by providing insights into the speech and communication profiles of individuals with ASD.
Disruptive Autoencoders: Leveraging Low-level features for 3D Medical Image Pre-training
Jul 31, 2023Abstract:Harnessing the power of pre-training on large-scale datasets like ImageNet forms a fundamental building block for the progress of representation learning-driven solutions in computer vision. Medical images are inherently different from natural images as they are acquired in the form of many modalities (CT, MR, PET, Ultrasound etc.) and contain granulated information like tissue, lesion, organs etc. These characteristics of medical images require special attention towards learning features representative of local context. In this work, we focus on designing an effective pre-training framework for 3D radiology images. First, we propose a new masking strategy called local masking where the masking is performed across channel embeddings instead of tokens to improve the learning of local feature representations. We combine this with classical low-level perturbations like adding noise and downsampling to further enable low-level representation learning. To this end, we introduce Disruptive Autoencoders, a pre-training framework that attempts to reconstruct the original image from disruptions created by a combination of local masking and low-level perturbations. Additionally, we also devise a cross-modal contrastive loss (CMCL) to accommodate the pre-training of multiple modalities in a single framework. We curate a large-scale dataset to enable pre-training of 3D medical radiology images (MRI and CT). The proposed pre-training framework is tested across multiple downstream tasks and achieves state-of-the-art performance. Notably, our proposed method tops the public test leaderboard of BTCV multi-organ segmentation challenge.
Communication-Efficient Vertical Federated Learning with Limited Overlapping Samples
Mar 30, 2023



Abstract:Federated learning is a popular collaborative learning approach that enables clients to train a global model without sharing their local data. Vertical federated learning (VFL) deals with scenarios in which the data on clients have different feature spaces but share some overlapping samples. Existing VFL approaches suffer from high communication costs and cannot deal efficiently with limited overlapping samples commonly seen in the real world. We propose a practical vertical federated learning (VFL) framework called \textbf{one-shot VFL} that can solve the communication bottleneck and the problem of limited overlapping samples simultaneously based on semi-supervised learning. We also propose \textbf{few-shot VFL} to improve the accuracy further with just one more communication round between the server and the clients. In our proposed framework, the clients only need to communicate with the server once or only a few times. We evaluate the proposed VFL framework on both image and tabular datasets. Our methods can improve the accuracy by more than 46.5\% and reduce the communication cost by more than 330$\times$ compared with state-of-the-art VFL methods when evaluated on CIFAR-10. Our code will be made publicly available at \url{https://nvidia.github.io/NVFlare/research/one-shot-vfl}.
Fair Federated Medical Image Segmentation via Client Contribution Estimation
Mar 29, 2023Abstract:How to ensure fairness is an important topic in federated learning (FL). Recent studies have investigated how to reward clients based on their contribution (collaboration fairness), and how to achieve uniformity of performance across clients (performance fairness). Despite achieving progress on either one, we argue that it is critical to consider them together, in order to engage and motivate more diverse clients joining FL to derive a high-quality global model. In this work, we propose a novel method to optimize both types of fairness simultaneously. Specifically, we propose to estimate client contribution in gradient and data space. In gradient space, we monitor the gradient direction differences of each client with respect to others. And in data space, we measure the prediction error on client data using an auxiliary model. Based on this contribution estimation, we propose a FL method, federated training via contribution estimation (FedCE), i.e., using estimation as global model aggregation weights. We have theoretically analyzed our method and empirically evaluated it on two real-world medical datasets. The effectiveness of our approach has been validated with significant performance improvements, better collaboration fairness, better performance fairness, and comprehensive analytical studies.
MONAI: An open-source framework for deep learning in healthcare
Nov 04, 2022



Abstract:Artificial Intelligence (AI) is having a tremendous impact across most areas of science. Applications of AI in healthcare have the potential to improve our ability to detect, diagnose, prognose, and intervene on human disease. For AI models to be used clinically, they need to be made safe, reproducible and robust, and the underlying software framework must be aware of the particularities (e.g. geometry, physiology, physics) of medical data being processed. This work introduces MONAI, a freely available, community-supported, and consortium-led PyTorch-based framework for deep learning in healthcare. MONAI extends PyTorch to support medical data, with a particular focus on imaging, and provide purpose-specific AI model architectures, transformations and utilities that streamline the development and deployment of medical AI models. MONAI follows best practices for software-development, providing an easy-to-use, robust, well-documented, and well-tested software framework. MONAI preserves the simple, additive, and compositional approach of its underlying PyTorch libraries. MONAI is being used by and receiving contributions from research, clinical and industrial teams from around the world, who are pursuing applications spanning nearly every aspect of healthcare.
NVIDIA FLARE: Federated Learning from Simulation to Real-World
Oct 24, 2022



Abstract:Federated learning (FL) enables building robust and generalizable AI models by leveraging diverse datasets from multiple collaborators without centralizing the data. We created NVIDIA FLARE as an open-source software development kit (SDK) to make it easier for data scientists to use FL in their research and real-world applications. The SDK includes solutions for state-of-the-art FL algorithms and federated machine learning approaches, which facilitate building workflows for distributed learning across enterprises and enable platform developers to create a secure, privacy-preserving offering for multiparty collaboration utilizing homomorphic encryption or differential privacy. The SDK is a lightweight, flexible, and scalable Python package, and allows researchers to bring their data science workflows implemented in any training libraries (PyTorch, TensorFlow, XGBoost, or even NumPy) and apply them in real-world FL settings. This paper introduces the key design principles of FLARE and illustrates some use cases (e.g., COVID analysis) with customizable FL workflows that implement different privacy-preserving algorithms. Code is available at https://github.com/NVIDIA/NVFlare.
Split-U-Net: Preventing Data Leakage in Split Learning for Collaborative Multi-Modal Brain Tumor Segmentation
Aug 22, 2022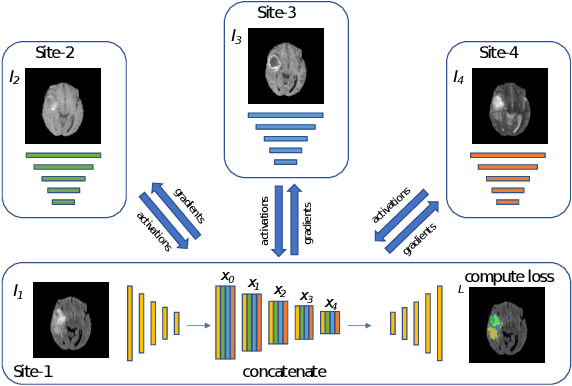
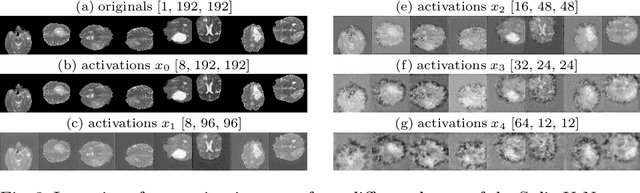

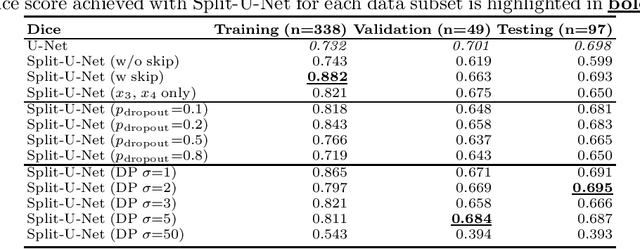
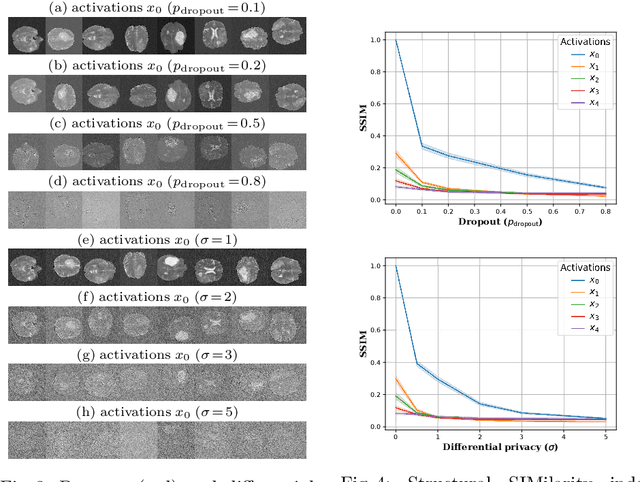
Abstract:Split learning (SL) has been proposed to train deep learning models in a decentralized manner. For decentralized healthcare applications with vertical data partitioning, SL can be beneficial as it allows institutes with complementary features or images for a shared set of patients to jointly develop more robust and generalizable models. In this work, we propose "Split-U-Net" and successfully apply SL for collaborative biomedical image segmentation. Nonetheless, SL requires the exchanging of intermediate activation maps and gradients to allow training models across different feature spaces, which might leak data and raise privacy concerns. Therefore, we also quantify the amount of data leakage in common SL scenarios for biomedical image segmentation and provide ways to counteract such leakage by applying appropriate defense strategies.
UNetFormer: A Unified Vision Transformer Model and Pre-Training Framework for 3D Medical Image Segmentation
Apr 05, 2022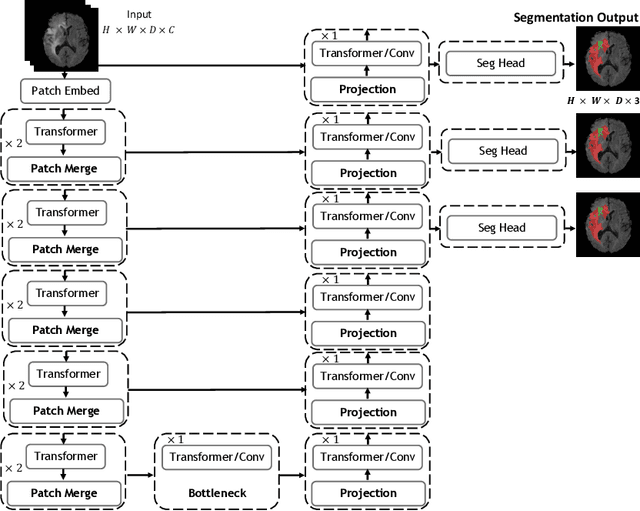
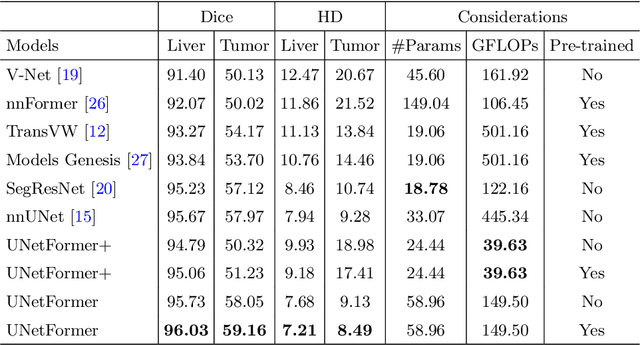
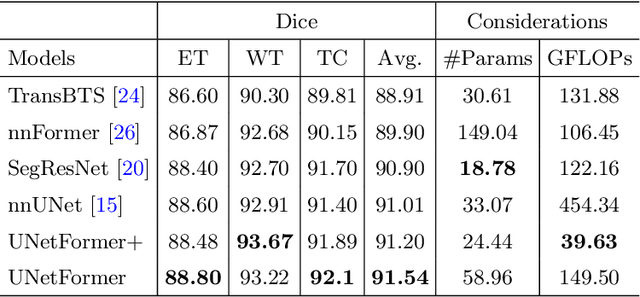
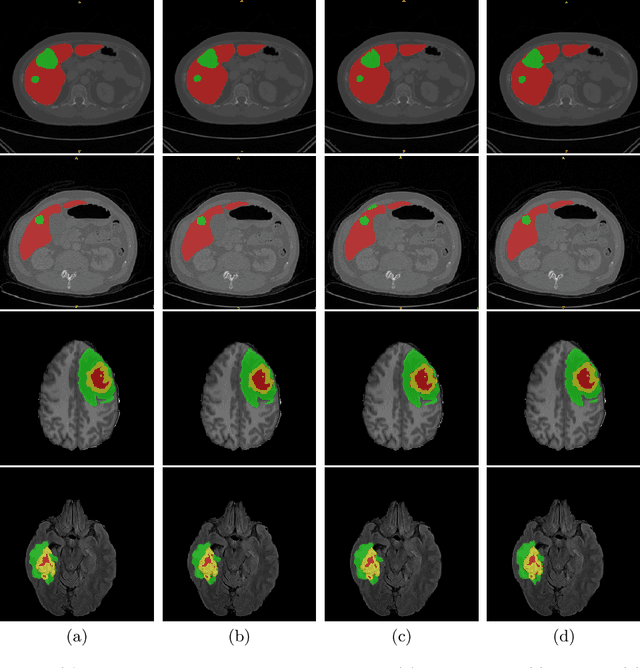
Abstract:Vision Transformers (ViT)s have recently become popular due to their outstanding modeling capabilities, in particular for capturing long-range information, and scalability to dataset and model sizes which has led to state-of-the-art performance in various computer vision and medical image analysis tasks. In this work, we introduce a unified framework consisting of two architectures, dubbed UNetFormer, with a 3D Swin Transformer-based encoder and Convolutional Neural Network (CNN) and transformer-based decoders. In the proposed model, the encoder is linked to the decoder via skip connections at five different resolutions with deep supervision. The design of proposed architecture allows for meeting a wide range of trade-off requirements between accuracy and computational cost. In addition, we present a methodology for self-supervised pre-training of the encoder backbone via learning to predict randomly masked volumetric tokens using contextual information of visible tokens. We pre-train our framework on a cohort of $5050$ CT images, gathered from publicly available CT datasets, and present a systematic investigation of various components such as masking ratio and patch size that affect the representation learning capability and performance of downstream tasks. We validate the effectiveness of our pre-training approach by fine-tuning and testing our model on liver and liver tumor segmentation task using the Medical Segmentation Decathlon (MSD) dataset and achieve state-of-the-art performance in terms of various segmentation metrics. To demonstrate its generalizability, we train and test the model on BraTS 21 dataset for brain tumor segmentation using MRI images and outperform other methods in terms of Dice score. Code: https://github.com/Project-MONAI/research-contributions
GradViT: Gradient Inversion of Vision Transformers
Mar 28, 2022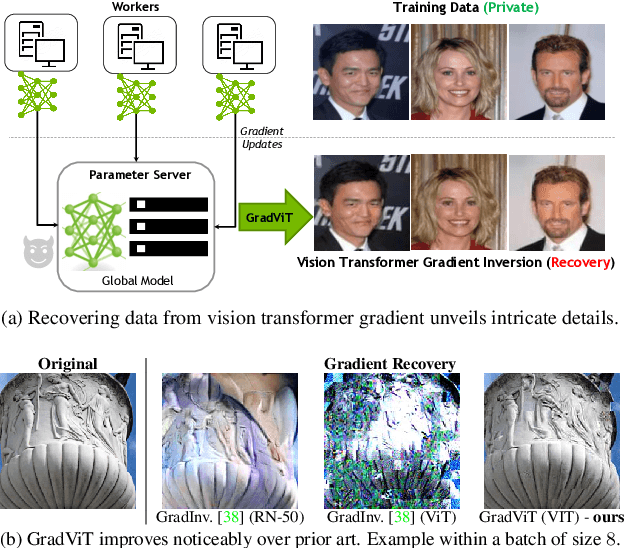
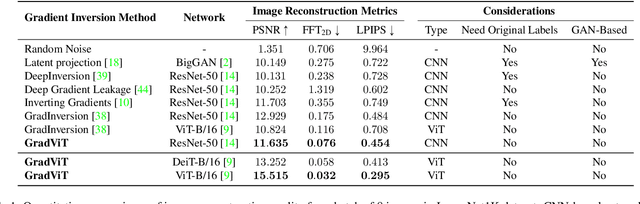
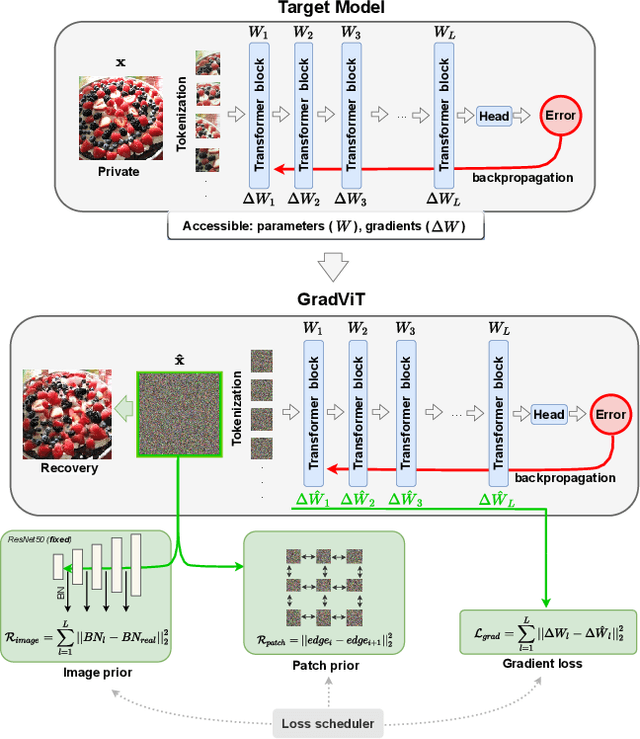
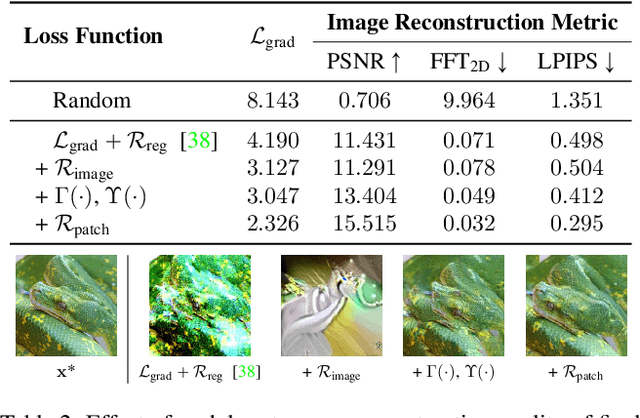
Abstract:In this work we demonstrate the vulnerability of vision transformers (ViTs) to gradient-based inversion attacks. During this attack, the original data batch is reconstructed given model weights and the corresponding gradients. We introduce a method, named GradViT, that optimizes random noise into naturally looking images via an iterative process. The optimization objective consists of (i) a loss on matching the gradients, (ii) image prior in the form of distance to batch-normalization statistics of a pretrained CNN model, and (iii) a total variation regularization on patches to guide correct recovery locations. We propose a unique loss scheduling function to overcome local minima during optimization. We evaluate GadViT on ImageNet1K and MS-Celeb-1M datasets, and observe unprecedentedly high fidelity and closeness to the original (hidden) data. During the analysis we find that vision transformers are significantly more vulnerable than previously studied CNNs due to the presence of the attention mechanism. Our method demonstrates new state-of-the-art results for gradient inversion in both qualitative and quantitative metrics. Project page at https://gradvit.github.io/.
MONAI Label: A framework for AI-assisted Interactive Labeling of 3D Medical Images
Mar 23, 2022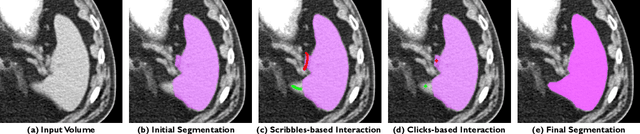
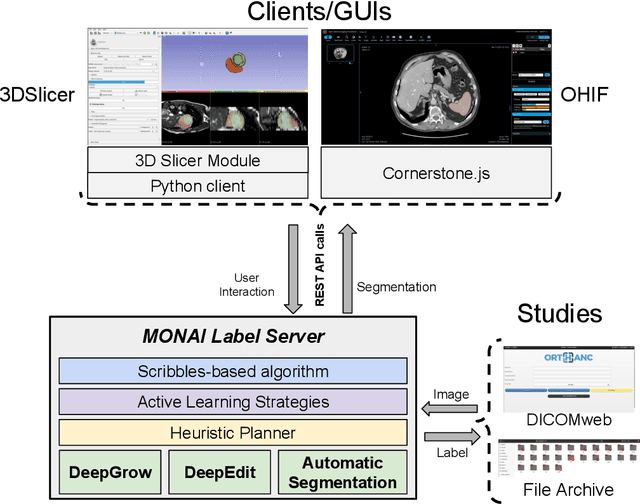
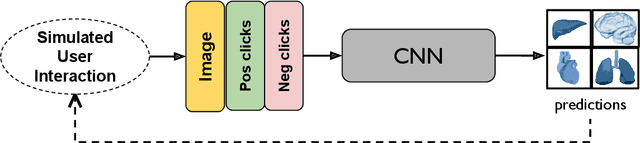
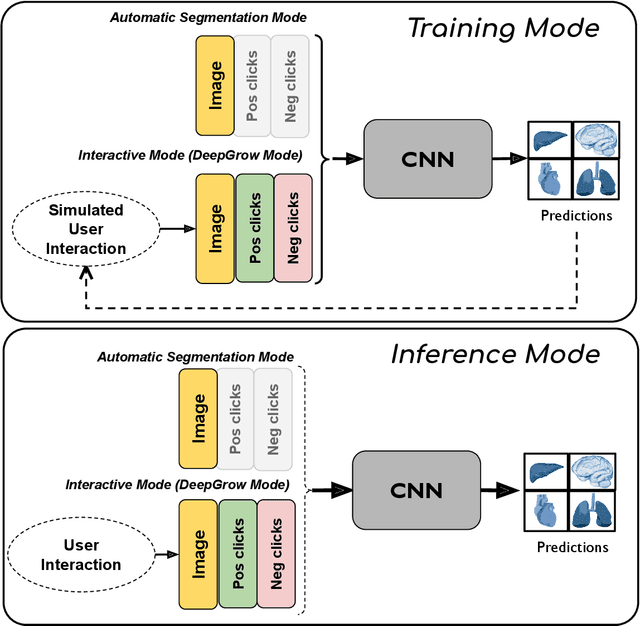
Abstract:The lack of annotated datasets is a major challenge in training new task-specific supervised AI algorithms as manual annotation is expensive and time-consuming. To address this problem, we present MONAI Label, a free and open-source platform that facilitates the development of AI-based applications that aim at reducing the time required to annotate 3D medical image datasets. Through MONAI Label researchers can develop annotation applications focusing on their domain of expertise. It allows researchers to readily deploy their apps as services, which can be made available to clinicians via their preferred user-interface. Currently, MONAI Label readily supports locally installed (3DSlicer) and web-based (OHIF) frontends, and offers two Active learning strategies to facilitate and speed up the training of segmentation algorithms. MONAI Label allows researchers to make incremental improvements to their labeling apps by making them available to other researchers and clinicians alike. Lastly, MONAI Label provides sample labeling apps, namely DeepEdit and DeepGrow, demonstrating dramatically reduced annotation times.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge