Pritesh Mehta
Confidence-aware Monocular Depth Estimation for Minimally Invasive Surgery
Mar 03, 2026Abstract:Purpose: Monocular depth estimation (MDE) is vital for scene understanding in minimally invasive surgery (MIS). However, endoscopic video sequences are often contaminated by smoke, specular reflections, blur, and occlusions, limiting the accuracy of MDE models. In addition, current MDE models do not output depth confidence, which could be a valuable tool for improving their clinical reliability. Methods: We propose a novel confidence-aware MDE framework featuring three significant contributions: (i) Calibrated confidence targets: an ensemble of fine-tuned stereo matching models is used to capture disparity variance into pixel-wise confidence probabilities; (ii) Confidence-aware loss: Baseline MDE models are optimized with confidence-aware loss functions, utilizing pixel-wise confidence probabilities such that reliable pixels dominate training; and (iii) Inference-time confidence: a confidence estimation head is proposed with two convolution layers to predict per-pixel confidence at inference, enabling assessment of depth reliability. Results: Comprehensive experimental validation across internal and public datasets demonstrates that our framework improves depth estimation accuracy and can robustly quantify the prediction's confidence. On the internal clinical endoscopic dataset (StereoKP), we improve dense depth estimation accuracy by ~8% as compared to the baseline model. Conclusion: Our confidence-aware framework enables improved accuracy of MDE models in MIS, addressing challenges posed by noise and artifacts in pre-clinical and clinical data, and allows MDE models to provide confidence maps that may be used to improve their reliability for clinical applications.
DeepEdit: Deep Editable Learning for Interactive Segmentation of 3D Medical Images
May 18, 2023Abstract:Automatic segmentation of medical images is a key step for diagnostic and interventional tasks. However, achieving this requires large amounts of annotated volumes, which can be tedious and time-consuming task for expert annotators. In this paper, we introduce DeepEdit, a deep learning-based method for volumetric medical image annotation, that allows automatic and semi-automatic segmentation, and click-based refinement. DeepEdit combines the power of two methods: a non-interactive (i.e. automatic segmentation using nnU-Net, UNET or UNETR) and an interactive segmentation method (i.e. DeepGrow), into a single deep learning model. It allows easy integration of uncertainty-based ranking strategies (i.e. aleatoric and epistemic uncertainty computation) and active learning. We propose and implement a method for training DeepEdit by using standard training combined with user interaction simulation. Once trained, DeepEdit allows clinicians to quickly segment their datasets by using the algorithm in auto segmentation mode or by providing clicks via a user interface (i.e. 3D Slicer, OHIF). We show the value of DeepEdit through evaluation on the PROSTATEx dataset for prostate/prostatic lesions and the Multi-Atlas Labeling Beyond the Cranial Vault (BTCV) dataset for abdominal CT segmentation, using state-of-the-art network architectures as baseline for comparison. DeepEdit could reduce the time and effort annotating 3D medical images compared to DeepGrow alone. Source code is available at https://github.com/Project-MONAI/MONAILabel
MONAI Label: A framework for AI-assisted Interactive Labeling of 3D Medical Images
Mar 23, 2022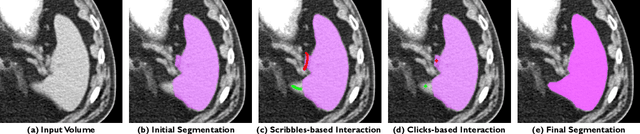
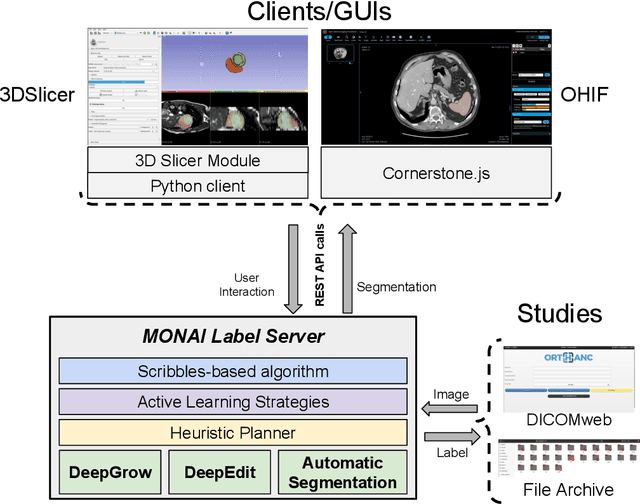
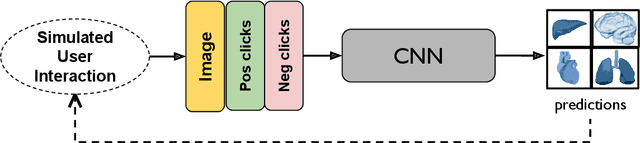
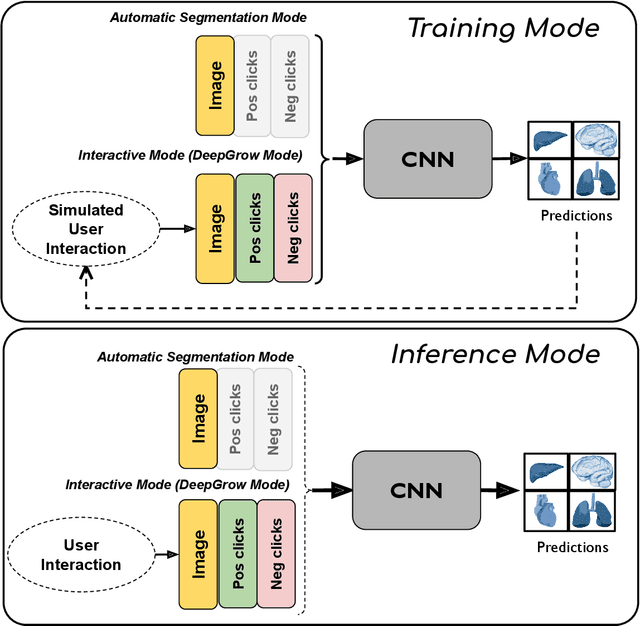
Abstract:The lack of annotated datasets is a major challenge in training new task-specific supervised AI algorithms as manual annotation is expensive and time-consuming. To address this problem, we present MONAI Label, a free and open-source platform that facilitates the development of AI-based applications that aim at reducing the time required to annotate 3D medical image datasets. Through MONAI Label researchers can develop annotation applications focusing on their domain of expertise. It allows researchers to readily deploy their apps as services, which can be made available to clinicians via their preferred user-interface. Currently, MONAI Label readily supports locally installed (3DSlicer) and web-based (OHIF) frontends, and offers two Active learning strategies to facilitate and speed up the training of segmentation algorithms. MONAI Label allows researchers to make incremental improvements to their labeling apps by making them available to other researchers and clinicians alike. Lastly, MONAI Label provides sample labeling apps, namely DeepEdit and DeepGrow, demonstrating dramatically reduced annotation times.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge