Wenjia Cai
Towards Medical Artificial General Intelligence via Knowledge-Enhanced Multimodal Pretraining
Apr 26, 2023



Abstract:Medical artificial general intelligence (MAGI) enables one foundation model to solve different medical tasks, which is very practical in the medical domain. It can significantly reduce the requirement of large amounts of task-specific data by sufficiently sharing medical knowledge among different tasks. However, due to the challenges of designing strongly generalizable models with limited and complex medical data, most existing approaches tend to develop task-specific models. To take a step towards MAGI, we propose a new paradigm called Medical-knOwledge-enhanced mulTimOdal pretRaining (MOTOR). In MOTOR, we combine two kinds of basic medical knowledge, i.e., general and specific knowledge, in a complementary manner to boost the general pretraining process. As a result, the foundation model with comprehensive basic knowledge can learn compact representations from pretraining radiographic data for better cross-modal alignment. MOTOR unifies the understanding and generation, which are two kinds of core intelligence of an AI system, into a single medical foundation model, to flexibly handle more diverse medical tasks. To enable a comprehensive evaluation and facilitate further research, we construct a medical multimodal benchmark including a wide range of downstream tasks, such as chest x-ray report generation and medical visual question answering. Extensive experiments on our benchmark show that MOTOR obtains promising results through simple task-oriented adaptation. The visualization shows that the injected knowledge successfully highlights key information in the medical data, demonstrating the excellent interpretability of MOTOR. Our MOTOR successfully mimics the human practice of fulfilling a "medical student" to accelerate the process of becoming a "specialist". We believe that our work makes a significant stride in realizing MAGI.
Cross-modal Clinical Graph Transformer for Ophthalmic Report Generation
Jun 04, 2022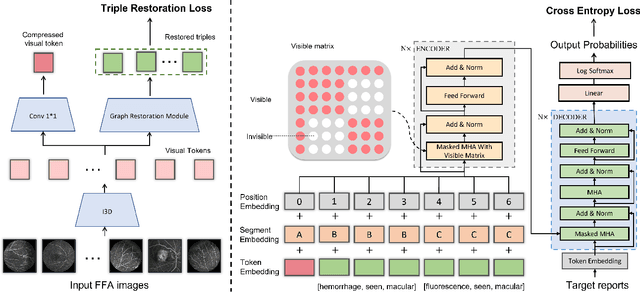

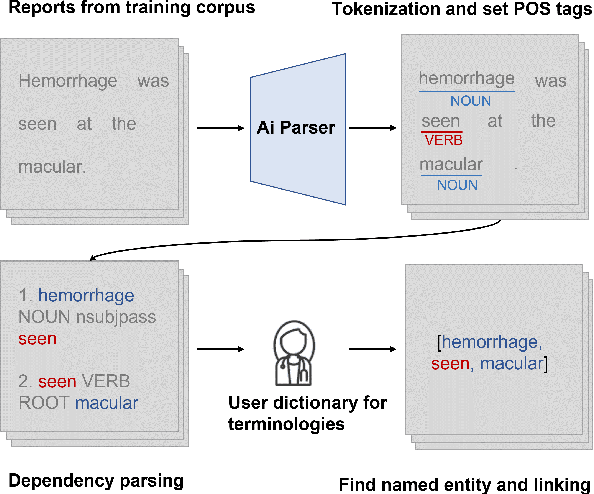
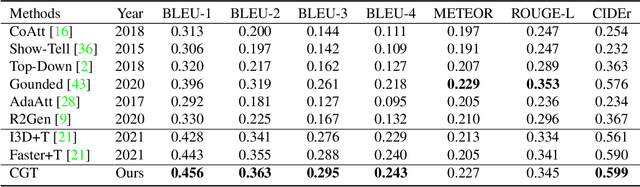
Abstract:Automatic generation of ophthalmic reports using data-driven neural networks has great potential in clinical practice. When writing a report, ophthalmologists make inferences with prior clinical knowledge. This knowledge has been neglected in prior medical report generation methods. To endow models with the capability of incorporating expert knowledge, we propose a Cross-modal clinical Graph Transformer (CGT) for ophthalmic report generation (ORG), in which clinical relation triples are injected into the visual features as prior knowledge to drive the decoding procedure. However, two major common Knowledge Noise (KN) issues may affect models' effectiveness. 1) Existing general biomedical knowledge bases such as the UMLS may not align meaningfully to the specific context and language of the report, limiting their utility for knowledge injection. 2) Incorporating too much knowledge may divert the visual features from their correct meaning. To overcome these limitations, we design an automatic information extraction scheme based on natural language processing to obtain clinical entities and relations directly from in-domain training reports. Given a set of ophthalmic images, our CGT first restores a sub-graph from the clinical graph and injects the restored triples into visual features. Then visible matrix is employed during the encoding procedure to limit the impact of knowledge. Finally, reports are predicted by the encoded cross-modal features via a Transformer decoder. Extensive experiments on the large-scale FFA-IR benchmark demonstrate that the proposed CGT is able to outperform previous benchmark methods and achieve state-of-the-art performances.
A Deep Learning-based Quality Assessment and Segmentation System with a Large-scale Benchmark Dataset for Optical Coherence Tomographic Angiography Image
Jul 22, 2021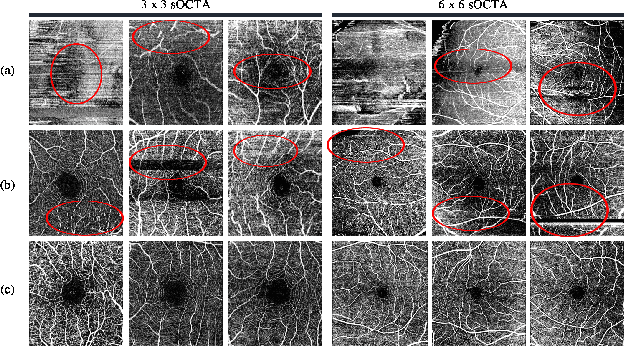
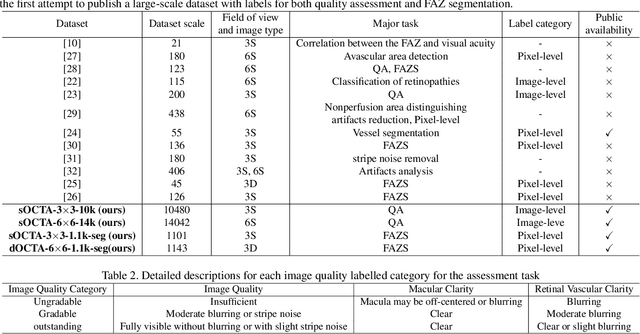
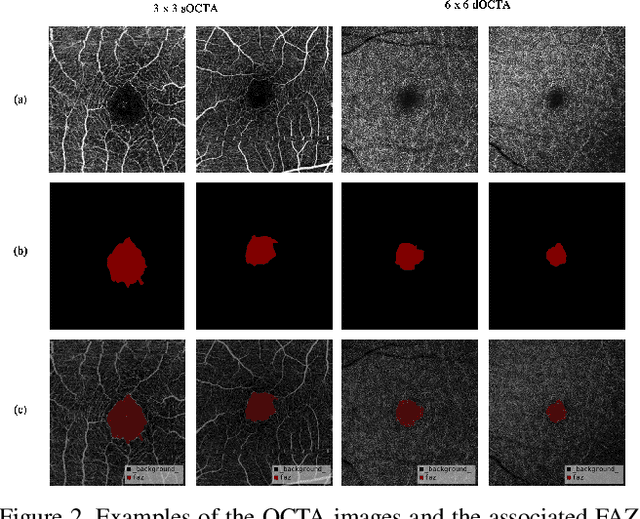
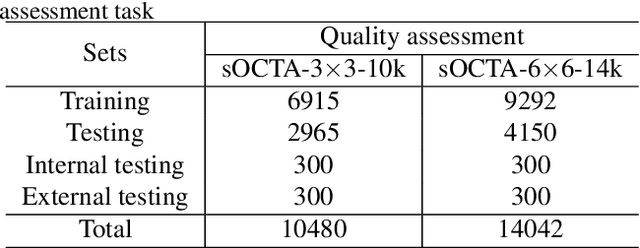
Abstract:Optical Coherence Tomography Angiography (OCTA) is a non-invasive and non-contacting imaging technique providing visualization of microvasculature of retina and optic nerve head in human eyes in vivo. The adequate image quality of OCTA is the prerequisite for the subsequent quantification of retinal microvasculature. Traditionally, the image quality score based on signal strength is used for discriminating low quality. However, it is insufficient for identifying artefacts such as motion and off-centration, which rely specialized knowledge and need tedious and time-consuming manual identification. One of the most primary issues in OCTA analysis is to sort out the foveal avascular zone (FAZ) region in the retina, which highly correlates with any visual acuity disease. However, the variations in OCTA visual quality affect the performance of deep learning in any downstream marginally. Moreover, filtering the low-quality OCTA images out is both labor-intensive and time-consuming. To address these issues, we develop an automated computer-aided OCTA image processing system using deep neural networks as the classifier and segmentor to help ophthalmologists in clinical diagnosis and research. This system can be an assistive tool as it can process OCTA images of different formats to assess the quality and segment the FAZ area. The source code is freely available at https://github.com/shanzha09/COIPS.git. Another major contribution is the large-scale OCTA dataset, namely OCTA-25K-IQA-SEG we publicize for performance evaluation. It is comprised of four subsets, namely sOCTA-3$\times$3-10k, sOCTA-6$\times$6-14k, sOCTA-3$\times$3-1.1k-seg, and dOCTA-6$\times$6-1.1k-seg, which contains a total number of 25,665 images. The large-scale OCTA dataset is available at https://doi.org/10.5281/zenodo.5111975, https://doi.org/10.5281/zenodo.5111972.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge