Wei Peng
Innovation Center for Pathogen Research Guangzhou Laboratory
Learning Hierarchical Procedural Memory for LLM Agents through Bayesian Selection and Contrastive Refinement
Dec 22, 2025Abstract:We present MACLA, a framework that decouples reasoning from learning by maintaining a frozen large language model while performing all adaptation in an external hierarchical procedural memory. MACLA extracts reusable procedures from trajectories, tracks reliability via Bayesian posteriors, selects actions through expected-utility scoring, and refines procedures by contrasting successes and failures. Across four benchmarks (ALFWorld, WebShop, TravelPlanner, InterCodeSQL), MACLA achieves 78.1 percent average performance, outperforming all baselines. On ALFWorld unseen tasks, MACLA reaches 90.3 percent with 3.1 percent positive generalization. The system constructs memory in 56 seconds, 2800 times faster than the state-of-the-art LLM parameter-training baseline, compressing 2851 trajectories into 187 procedures. Experimental results demonstrate that structured external memory with Bayesian selection and contrastive refinement enables sample-efficient, interpretable, and continually improving agents without LLM parameter updates.
OPBO: Order-Preserving Bayesian Optimization
Dec 22, 2025Abstract:Bayesian optimization is an effective method for solving expensive black-box optimization problems. Most existing methods use Gaussian processes (GP) as the surrogate model for approximating the black-box objective function, it is well-known that it can fail in high-dimensional space (e.g., dimension over 500). We argue that the reliance of GP on precise numerical fitting is fundamentally ill-suited in high-dimensional space, where it leads to prohibitive computational complexity. In order to address this, we propose a simple order-preserving Bayesian optimization (OPBO) method, where the surrogate model preserves the order, instead of the value, of the black-box objective function. Then we can use a simple but effective OP neural network (NN) to replace GP as the surrogate model. Moreover, instead of searching for the best solution from the acquisition model, we select good-enough solutions in the ordinal set to reduce computational cost. The experimental results show that for high-dimensional (over 500) black-box optimization problems, the proposed OPBO significantly outperforms traditional BO methods based on regression NN and GP. The source code is available at https://github.com/pengwei222/OPBO.
Are Vision Language Models Cross-Cultural Theory of Mind Reasoners?
Dec 19, 2025



Abstract:Theory of Mind (ToM) -- the ability to attribute beliefs, desires, and emotions to others -- is fundamental for human social intelligence, yet remains a major challenge for artificial agents. Existing Vision-Language Models (VLMs) are increasingly applied in socially grounded tasks, but their capacity for cross-cultural ToM reasoning is largely unexplored. In this work, we introduce CulturalToM-VQA, a new evaluation benchmark containing 5095 questions designed to probe ToM reasoning across diverse cultural contexts through visual question answering. The dataset captures culturally grounded cues such as rituals, attire, gestures, and interpersonal dynamics, enabling systematic evaluation of ToM reasoning beyond Western-centric benchmarks. Our dataset is built through a VLM-assisted human-in-the-loop pipeline, where human experts first curate culturally rich images across traditions, rituals, and social interactions; a VLM then assist in generating structured ToM-focused scene descriptions, which are refined into question-answer pairs spanning a taxonomy of six ToM tasks and four graded complexity levels. The resulting dataset covers diverse theory of mind facets such as mental state attribution, false belief reasoning, non-literal communication, social norm violations, perspective coordination, and multi-agent reasoning.
TAO-Net: Two-stage Adaptive OOD Classification Network for Fine-grained Encrypted Traffic Classification
Dec 11, 2025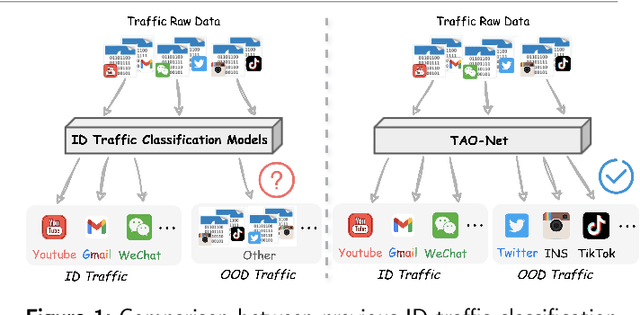

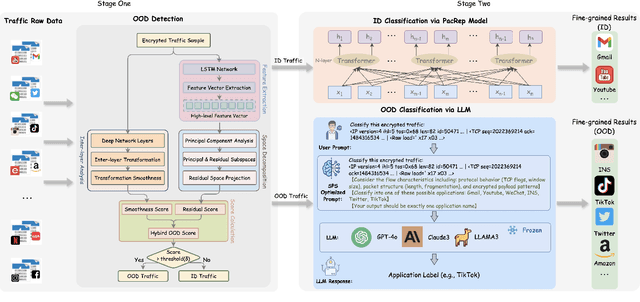
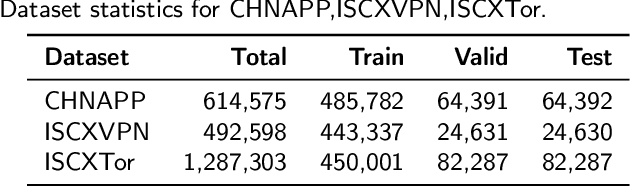
Abstract:Encrypted traffic classification aims to identify applications or services by analyzing network traffic data. One of the critical challenges is the continuous emergence of new applications, which generates Out-of-Distribution (OOD) traffic patterns that deviate from known categories and are not well represented by predefined models. Current approaches rely on predefined categories, which limits their effectiveness in handling unknown traffic types. Although some methods mitigate this limitation by simply classifying unknown traffic into a single "Other" category, they fail to make a fine-grained classification. In this paper, we propose a Two-stage Adaptive OOD classification Network (TAO-Net) that achieves accurate classification for both In-Distribution (ID) and OOD encrypted traffic. The method incorporates an innovative two-stage design: the first stage employs a hybrid OOD detection mechanism that integrates transformer-based inter-layer transformation smoothness and feature analysis to effectively distinguish between ID and OOD traffic, while the second stage leverages large language models with a novel semantic-enhanced prompt strategy to transform OOD traffic classification into a generation task, enabling flexible fine-grained classification without relying on predefined labels. Experiments on three datasets demonstrate that TAO-Net achieves 96.81-97.70% macro-precision and 96.77-97.68% macro-F1, outperforming previous methods that only reach 44.73-86.30% macro-precision, particularly in identifying emerging network applications.
Integrating Anatomical Priors into a Causal Diffusion Model
Sep 10, 2025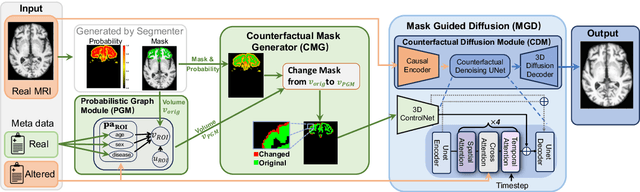
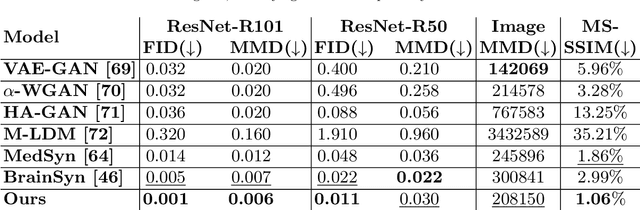
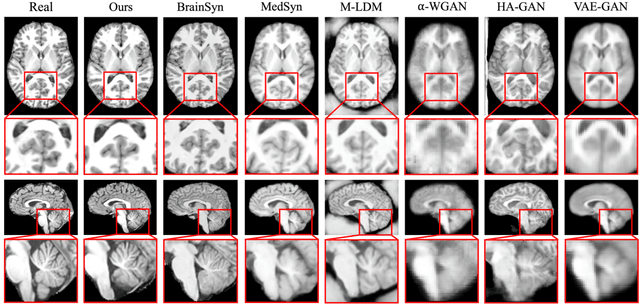
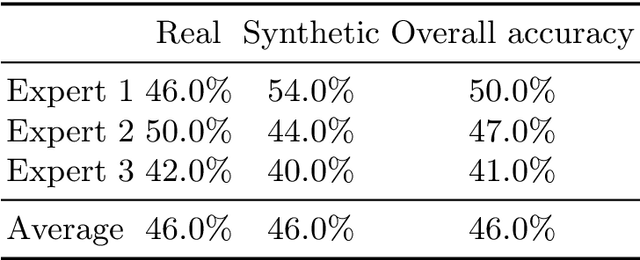
Abstract:3D brain MRI studies often examine subtle morphometric differences between cohorts that are hard to detect visually. Given the high cost of MRI acquisition, these studies could greatly benefit from image syntheses, particularly counterfactual image generation, as seen in other domains, such as computer vision. However, counterfactual models struggle to produce anatomically plausible MRIs due to the lack of explicit inductive biases to preserve fine-grained anatomical details. This shortcoming arises from the training of the models aiming to optimize for the overall appearance of the images (e.g., via cross-entropy) rather than preserving subtle, yet medically relevant, local variations across subjects. To preserve subtle variations, we propose to explicitly integrate anatomical constraints on a voxel-level as prior into a generative diffusion framework. Called Probabilistic Causal Graph Model (PCGM), the approach captures anatomical constraints via a probabilistic graph module and translates those constraints into spatial binary masks of regions where subtle variations occur. The masks (encoded by a 3D extension of ControlNet) constrain a novel counterfactual denoising UNet, whose encodings are then transferred into high-quality brain MRIs via our 3D diffusion decoder. Extensive experiments on multiple datasets demonstrate that PCGM generates structural brain MRIs of higher quality than several baseline approaches. Furthermore, we show for the first time that brain measurements extracted from counterfactuals (generated by PCGM) replicate the subtle effects of a disease on cortical brain regions previously reported in the neuroscience literature. This achievement is an important milestone in the use of synthetic MRIs in studies investigating subtle morphological differences.
TAMMs: Temporal-Aware Multimodal Model for Satellite Image Change Understanding and Forecasting
Jun 23, 2025



Abstract:Satellite image time-series analysis demands fine-grained spatial-temporal reasoning, which remains a challenge for existing multimodal large language models (MLLMs). In this work, we study the capabilities of MLLMs on a novel task that jointly targets temporal change understanding and future scene generation, aiming to assess their potential for modeling complex multimodal dynamics over time. We propose TAMMs, a Temporal-Aware Multimodal Model for satellite image change understanding and forecasting, which enhances frozen MLLMs with lightweight temporal modules for structured sequence encoding and contextual prompting. To guide future image generation, TAMMs introduces a Semantic-Fused Control Injection (SFCI) mechanism that adaptively combines high-level semantic reasoning and structural priors within an enhanced ControlNet. This dual-path conditioning enables temporally consistent and semantically grounded image synthesis. Experiments demonstrate that TAMMs outperforms strong MLLM baselines in both temporal change understanding and future image forecasting tasks, highlighting how carefully designed temporal reasoning and semantic fusion can unlock the full potential of MLLMs for spatio-temporal understanding.
Biomed-DPT: Dual Modality Prompt Tuning for Biomedical Vision-Language Models
May 08, 2025Abstract:Prompt learning is one of the most effective paradigms for adapting pre-trained vision-language models (VLMs) to the biomedical image classification tasks in few shot scenarios. However, most of the current prompt learning methods only used the text prompts and ignored the particular structures (such as the complex anatomical structures and subtle pathological features) in the biomedical images. In this work, we propose Biomed-DPT, a knowledge-enhanced dual modality prompt tuning technique. In designing the text prompt, Biomed-DPT constructs a dual prompt including the template-driven clinical prompts and the large language model (LLM)-driven domain-adapted prompts, then extracts the clinical knowledge from the domain-adapted prompts through the knowledge distillation technique. In designing the vision prompt, Biomed-DPT introduces the zero vector as a soft prompt to leverage attention re-weighting so that the focus on non-diagnostic regions and the recognition of non-critical pathological features are avoided. Biomed-DPT achieves an average classification accuracy of 66.14\% across 11 biomedical image datasets covering 9 modalities and 10 organs, with performance reaching 78.06\% in base classes and 75.97\% in novel classes, surpassing the Context Optimization (CoOp) method by 6.20\%, 3.78\%, and 8.04\%, respectively. Our code are available at \underline{https://github.com/Kanyooo/Biomed-DPT}.
Learning based convex approximation for constrained parametric optimization
May 07, 2025Abstract:We propose an input convex neural network (ICNN)-based self-supervised learning framework to solve continuous constrained optimization problems. By integrating the augmented Lagrangian method (ALM) with the constraint correction mechanism, our framework ensures \emph{non-strict constraint feasibility}, \emph{better optimality gap}, and \emph{best convergence rate} with respect to the state-of-the-art learning-based methods. We provide a rigorous convergence analysis, showing that the algorithm converges to a Karush-Kuhn-Tucker (KKT) point of the original problem even when the internal solver is a neural network, and the approximation error is bounded. We test our approach on a range of benchmark tasks including quadratic programming (QP), nonconvex programming, and large-scale AC optimal power flow problems. The results demonstrate that compared to existing solvers (e.g., \texttt{OSQP}, \texttt{IPOPT}) and the latest learning-based methods (e.g., DC3, PDL), our approach achieves a superior balance among accuracy, feasibility, and computational efficiency.
Anatomical Similarity as a New Metric to Evaluate Brain Generative Models
Apr 30, 2025Abstract:Generative models enhance neuroimaging through data augmentation, quality improvement, and rare condition studies. Despite advances in realistic synthetic MRIs, evaluations focus on texture and perception, lacking sensitivity to crucial anatomical fidelity. This study proposes a new metric, called WASABI (Wasserstein-Based Anatomical Brain Index), to assess the anatomical realism of synthetic brain MRIs. WASABI leverages \textit{SynthSeg}, a deep learning-based brain parcellation tool, to derive volumetric measures of brain regions in each MRI and uses the multivariate Wasserstein distance to compare distributions between real and synthetic anatomies. Based on controlled experiments on two real datasets and synthetic MRIs from five generative models, WASABI demonstrates higher sensitivity in quantifying anatomical discrepancies compared to traditional image-level metrics, even when synthetic images achieve near-perfect visual quality. Our findings advocate for shifting the evaluation paradigm beyond visual inspection and conventional metrics, emphasizing anatomical fidelity as a crucial benchmark for clinically meaningful brain MRI synthesis. Our code is available at https://github.com/BahramJafrasteh/wasabi-mri.
WorkTeam: Constructing Workflows from Natural Language with Multi-Agents
Mar 28, 2025



Abstract:Workflows play a crucial role in enhancing enterprise efficiency by orchestrating complex processes with multiple tools or components. However, hand-crafted workflow construction requires expert knowledge, presenting significant technical barriers. Recent advancements in Large Language Models (LLMs) have improved the generation of workflows from natural language instructions (aka NL2Workflow), yet existing single LLM agent-based methods face performance degradation on complex tasks due to the need for specialized knowledge and the strain of task-switching. To tackle these challenges, we propose WorkTeam, a multi-agent NL2Workflow framework comprising a supervisor, orchestrator, and filler agent, each with distinct roles that collaboratively enhance the conversion process. As there are currently no publicly available NL2Workflow benchmarks, we also introduce the HW-NL2Workflow dataset, which includes 3,695 real-world business samples for training and evaluation. Experimental results show that our approach significantly increases the success rate of workflow construction, providing a novel and effective solution for enterprise NL2Workflow services.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge