Hyeryun Park
Coding-Free and Privacy-Preserving MCP Framework for Clinical Agentic Research Intelligence System
Apr 14, 2026Abstract:Clinical research involves labor-intensive processes such as study design, cohort construction, model development, and documentation, requiring domain expertise, programming skills, and access to sensitive patient data. These demands create barriers for clinicians and external researchers conducting data-driven studies. To overcome these limitations, we developed a Clinical Agentic Research Intelligence System (CARIS) that automates the clinical research workflow while preserving data privacy, enabling comprehensive studies without direct access to raw data. CARIS integrates Large Language Models (LLMs) with modular tools via the Model Context Protocol (MCP), enabling natural language-driven orchestration of appropriate tools. Databases remain securely within the MCP server, and users access only the outputs and final research reports. Based on user intent, CARIS automatically executes the full pipeline: research planning, literature search, cohort construction, Institutional Review Board (IRB) documentation, Vibe Machine Learning (ML), and report generation, with iterative human-in-the-loop refinement. We evaluated CARIS on three heterogeneous datasets with distinct clinical tasks. Research plans and IRB documents were finalized within three to four iterations, using evidence from literature and data. The system supported Vibe ML by exploring feature-model combinations, ranking the top ten models, and generating performance visualizations. Final reports showed high completeness based on a checklist derived from the TRIPOD+AI framework, achieving 96% coverage in LLM evaluation and 82% in human evaluation. CARIS demonstrates that agentic AI can transform clinical hypotheses into executable research workflows across heterogeneous datasets. By eliminating the need for coding and direct data access, the system lowers barriers and bridges public and private clinical data environments.
Surgical Agent Orchestration Platform for Voice-directed Patient Data Interaction
Nov 11, 2025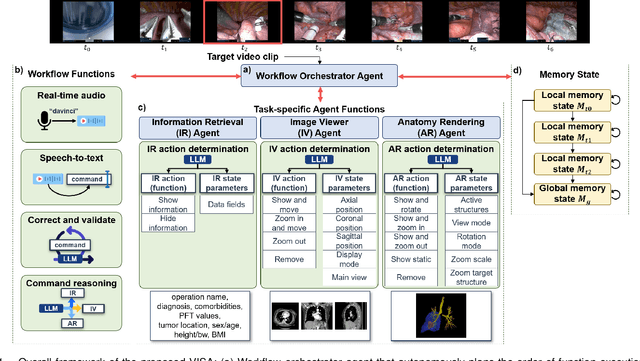
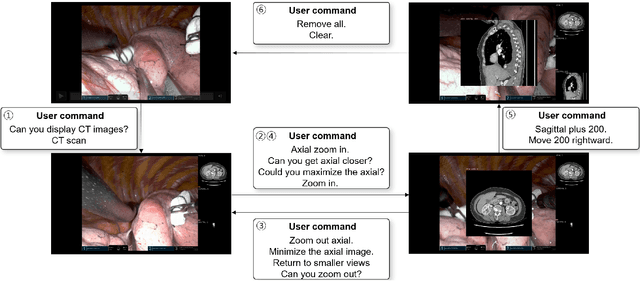
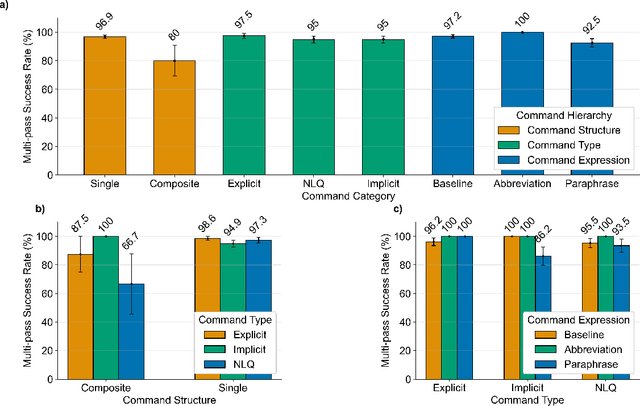
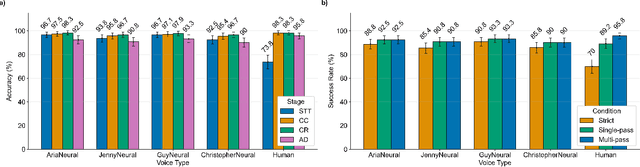

Abstract:In da Vinci robotic surgery, surgeons' hands and eyes are fully engaged in the procedure, making it difficult to access and manipulate multimodal patient data without interruption. We propose a voice-directed Surgical Agent Orchestrator Platform (SAOP) built on a hierarchical multi-agent framework, consisting of an orchestration agent and three task-specific agents driven by Large Language Models (LLMs). These LLM-based agents autonomously plan, refine, validate, and reason to map voice commands into specific tasks such as retrieving clinical information, manipulating CT scans, or navigating 3D anatomical models on the surgical video. We also introduce a Multi-level Orchestration Evaluation Metric (MOEM) to comprehensively assess the performance and robustness from command-level and category-level perspectives. The SAOP achieves high accuracy and success rates across 240 voice commands, while LLM-based agents improve robustness against speech recognition errors and diverse or ambiguous free-form commands, demonstrating strong potential to support minimally invasive da Vinci robotic surgery.
LoraMap: Harnessing the Power of LoRA Connections
Aug 29, 2024



Abstract:Large Language Models (LLMs) can benefit from mitigating hallucinations through fact-checking and overcoming substantial computational overhead with parameter-efficient techniques such as Low-Rank Adaptation (LoRA). While some studies have explored the parallel integration of multiple LoRAs, these approaches need attention to the connections between them. This paper investigates methods to establish connections among multiple LoRAs. We create three reasoning datasets tailored to fact-checking and fine-tune individual LoRAs, allowing them to view and reason from diverse perspectives. Then, we explore strategies for allocating these reasoning LoRAs and introduce LoraMap, an approach to map connections between them. The results on the fact-checking task demonstrate that the performance of LoraMap is superior to LoraHub, an existing LoRA composition method. LoraMap also outperforms with significantly fewer parameters than LoraConcat, which concatenates LoRAs and further fine-tunes them.
Automated Information Extraction from Thyroid Operation Narrative: A Comparative Study of GPT-4 and Fine-tuned KoELECTRA
Jun 12, 2024



Abstract:In the rapidly evolving field of healthcare, the integration of artificial intelligence (AI) has become a pivotal component in the automation of clinical workflows, ushering in a new era of efficiency and accuracy. This study focuses on the transformative capabilities of the fine-tuned KoELECTRA model in comparison to the GPT-4 model, aiming to facilitate automated information extraction from thyroid operation narratives. The current research landscape is dominated by traditional methods heavily reliant on regular expressions, which often face challenges in processing free-style text formats containing critical details of operation records, including frozen biopsy reports. Addressing this, the study leverages advanced natural language processing (NLP) techniques to foster a paradigm shift towards more sophisticated data processing systems. Through this comparative study, we aspire to unveil a more streamlined, precise, and efficient approach to document processing in the healthcare domain, potentially revolutionizing the way medical data is handled and analyzed.
* 9 pages, 2 figures, 3 tables
Clinical Text Generation through Leveraging Medical Concept and Relations
Oct 02, 2019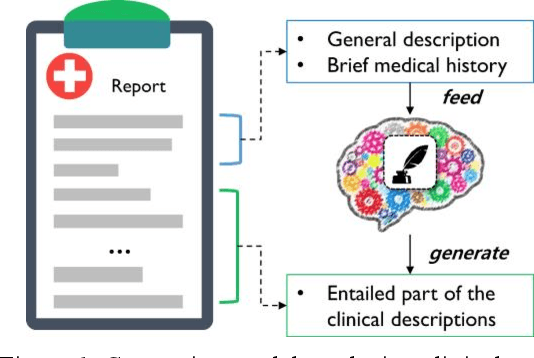
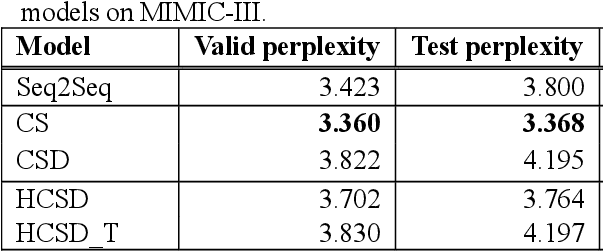
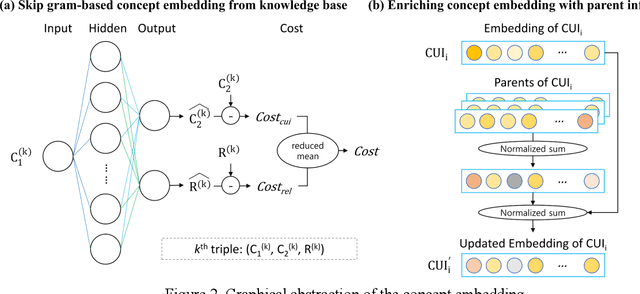
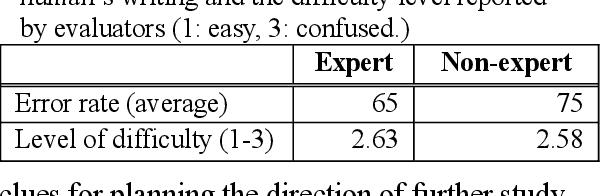
Abstract:With a neural sequence generation model, this study aims to develop a method of writing the patient clinical texts given a brief medical history. As a proof-of-a-concept, we have demonstrated that it can be workable to use medical concept embedding in clinical text generation. Our model was based on the Sequence-to-Sequence architecture and trained with a large set of de-identified clinical text data. The quantitative result shows that our concept embedding method decreased the perplexity of the baseline architecture. Also, we discuss the analyzed results from a human evaluation performed by medical doctors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge