Qingli Li
Dictionary-based Pathology Mining with Hard-instance-assisted Classifier Debiasing for Genetic Biomarker Prediction from WSIs
Mar 26, 2026Abstract:Prediction of genetic biomarkers, e.g., microsatellite instability in colorectal cancer is crucial for clinical decision making. But, two primary challenges hamper accurate prediction: (1) It is difficult to construct a pathology-aware representation involving the complex interconnections among pathological components. (2) WSIs contain a large proportion of areas unrelated to genetic biomarkers, which make the model easily overfit simple but irrelative instances. We hereby propose a Dictionary-based hierarchical pathology mining with hard-instance-assisted classifier Debiasing framework to address these challenges, dubbed as D2Bio. Our first module, dictionary-based hierarchical pathology mining, is able to mine diverse and very fine-grained pathological contextual interaction without the limit to the distances between patches. The second module, hard-instance-assisted classfier debiasing, learns a debiased classifier via focusing on hard but task-related features, without any additional annotations. Experimental results on five cohorts show the superiority of our method, with over 4% improvement in AUROC compared with the second best on the TCGA-CRC-MSI cohort. Our analysis further shows the clinical interpretability of D2Bio in genetic biomarker diagnosis and potential clinical utility in survival analysis. Code will be available at https://github.com/DeepMed-Lab-ECNU/D2Bio.
ReinPath: A Multimodal Reinforcement Learning Approach for Pathology
Jan 21, 2026Abstract:Interpretability is significant in computational pathology, leading to the development of multimodal information integration from histopathological image and corresponding text data.However, existing multimodal methods have limited interpretability due to the lack of high-quality dataset that support explicit reasoning and inference and simple reasoning process.To address the above problems, we introduce a novel multimodal pathology large language model with strong reasoning capabilities.To improve the generation of accurate and contextually relevant textual descriptions, we design a semantic reward strategy integrated with group relative policy optimization.We construct a high-quality pathology visual question answering (VQA) dataset, specifically designed to support complex reasoning tasks.Comprehensive experiments conducted on this dataset demonstrate that our method outperforms state-of-the-art methods, even when trained with only 20% of the data.Our method also achieves comparable performance on downstream zero-shot image classification task compared with CLIP.
Training Report of TeleChat3-MoE
Dec 30, 2025Abstract:TeleChat3-MoE is the latest series of TeleChat large language models, featuring a Mixture-of-Experts (MoE) architecture with parameter counts ranging from 105 billion to over one trillion,trained end-to-end on Ascend NPU cluster. This technical report mainly presents the underlying training infrastructure that enables reliable and efficient scaling to frontier model sizes. We detail systematic methodologies for operator-level and end-to-end numerical accuracy verification, ensuring consistency across hardware platforms and distributed parallelism strategies. Furthermore, we introduce a suite of performance optimizations, including interleaved pipeline scheduling, attention-aware data scheduling for long-sequence training,hierarchical and overlapped communication for expert parallelism, and DVM-based operator fusion. A systematic parallelization framework, leveraging analytical estimation and integer linear programming, is also proposed to optimize multi-dimensional parallelism configurations. Additionally, we present methodological approaches to cluster-level optimizations, addressing host- and device-bound bottlenecks during large-scale training tasks. These infrastructure advancements yield significant throughput improvements and near-linear scaling on clusters comprising thousands of devices, providing a robust foundation for large-scale language model development on hardware ecosystems.
DragLoRA: Online Optimization of LoRA Adapters for Drag-based Image Editing in Diffusion Model
May 18, 2025Abstract:Drag-based editing within pretrained diffusion model provides a precise and flexible way to manipulate foreground objects. Traditional methods optimize the input feature obtained from DDIM inversion directly, adjusting them iteratively to guide handle points towards target locations. However, these approaches often suffer from limited accuracy due to the low representation ability of the feature in motion supervision, as well as inefficiencies caused by the large search space required for point tracking. To address these limitations, we present DragLoRA, a novel framework that integrates LoRA (Low-Rank Adaptation) adapters into the drag-based editing pipeline. To enhance the training of LoRA adapters, we introduce an additional denoising score distillation loss which regularizes the online model by aligning its output with that of the original model. Additionally, we improve the consistency of motion supervision by adapting the input features using the updated LoRA, giving a more stable and accurate input feature for subsequent operations. Building on this, we design an adaptive optimization scheme that dynamically toggles between two modes, prioritizing efficiency without compromising precision. Extensive experiments demonstrate that DragLoRA significantly enhances the control precision and computational efficiency for drag-based image editing. The Codes of DragLoRA are available at: https://github.com/Sylvie-X/DragLoRA.
MDN: Mamba-Driven Dualstream Network For Medical Hyperspectral Image Segmentation
Feb 24, 2025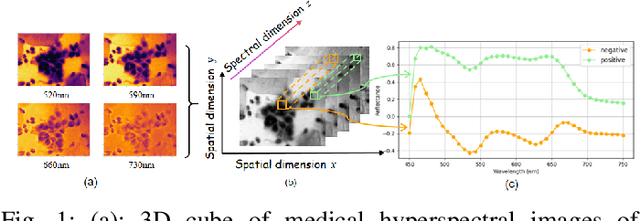
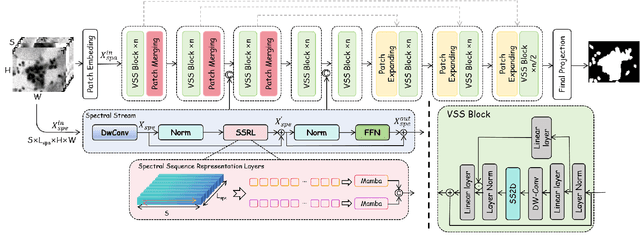
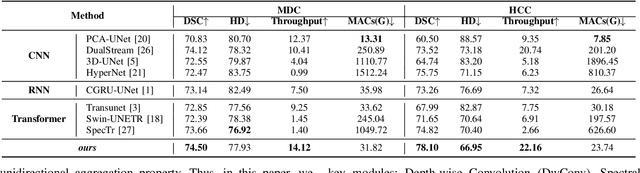
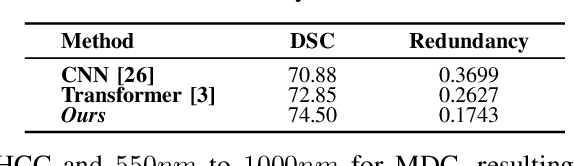
Abstract:Medical Hyperspectral Imaging (MHSI) offers potential for computational pathology and precision medicine. However, existing CNN and Transformer struggle to balance segmentation accuracy and speed due to high spatial-spectral dimensionality. In this study, we leverage Mamba's global context modeling to propose a dual-stream architecture for joint spatial-spectral feature extraction. To address the limitation of Mamba's unidirectional aggregation, we introduce a recurrent spectral sequence representation to capture low-redundancy global spectral features. Experiments on a public Multi-Dimensional Choledoch dataset and a private Cervical Cancer dataset show that our method outperforms state-of-the-art approaches in segmentation accuracy while minimizing resource usage and achieving the fastest inference speed. Our code will be available at https://github.com/DeepMed-Lab-ECNU/MDN.
Progressive Vision-Language Prompt for Multi-Organ Multi-Class Cell Semantic Segmentation with Single Branch
Dec 04, 2024



Abstract:Pathological cell semantic segmentation is a fundamental technology in computational pathology, essential for applications like cancer diagnosis and effective treatment. Given that multiple cell types exist across various organs, with subtle differences in cell size and shape, multi-organ, multi-class cell segmentation is particularly challenging. Most existing methods employ multi-branch frameworks to enhance feature extraction, but often result in complex architectures. Moreover, reliance on visual information limits performance in multi-class analysis due to intricate textural details. To address these challenges, we propose a Multi-OrgaN multi-Class cell semantic segmentation method with a single brancH (MONCH) that leverages vision-language input. Specifically, we design a hierarchical feature extraction mechanism to provide coarse-to-fine-grained features for segmenting cells of various shapes, including high-frequency, convolutional, and topological features. Inspired by the synergy of textual and multi-grained visual features, we introduce a progressive prompt decoder to harmonize multimodal information, integrating features from fine to coarse granularity for better context capture. Extensive experiments on the PanNuke dataset, which has significant class imbalance and subtle cell size and shape variations, demonstrate that MONCH outperforms state-of-the-art cell segmentation methods and vision-language models. Codes and implementations will be made publicly available.
A Semi-Supervised Approach with Error Reflection for Echocardiography Segmentation
Dec 01, 2024



Abstract:Segmenting internal structure from echocardiography is essential for the diagnosis and treatment of various heart diseases. Semi-supervised learning shows its ability in alleviating annotations scarcity. While existing semi-supervised methods have been successful in image segmentation across various medical imaging modalities, few have attempted to design methods specifically addressing the challenges posed by the poor contrast, blurred edge details and noise of echocardiography. These characteristics pose challenges to the generation of high-quality pseudo-labels in semi-supervised segmentation based on Mean Teacher. Inspired by human reflection on erroneous practices, we devise an error reflection strategy for echocardiography semi-supervised segmentation architecture. The process triggers the model to reflect on inaccuracies in unlabeled image segmentation, thereby enhancing the robustness of pseudo-label generation. Specifically, the strategy is divided into two steps. The first step is called reconstruction reflection. The network is tasked with reconstructing authentic proxy images from the semantic masks of unlabeled images and their auxiliary sketches, while maximizing the structural similarity between the original inputs and the proxies. The second step is called guidance correction. Reconstruction error maps decouple unreliable segmentation regions. Then, reliable data that are more likely to occur near high-density areas are leveraged to guide the optimization of unreliable data potentially located around decision boundaries. Additionally, we introduce an effective data augmentation strategy, termed as multi-scale mixing up strategy, to minimize the empirical distribution gap between labeled and unlabeled images and perceive diverse scales of cardiac anatomical structures. Extensive experiments demonstrate the competitiveness of the proposed method.
Label Filling via Mixed Supervision for Medical Image Segmentation from Noisy Annotations
Oct 21, 2024Abstract:The success of medical image segmentation usually requires a large number of high-quality labels. But since the labeling process is usually affected by the raters' varying skill levels and characteristics, the estimated masks provided by different raters usually suffer from high inter-rater variability. In this paper, we propose a simple yet effective Label Filling framework, termed as LF-Net, predicting the groundtruth segmentation label given only noisy annotations during training. The fundamental idea of label filling is to supervise the segmentation model by a subset of pixels with trustworthy labels, meanwhile filling labels of other pixels by mixed supervision. More concretely, we propose a qualified majority voting strategy, i.e., a threshold voting scheme is designed to model agreement among raters and the majority-voted labels of the selected subset of pixels are regarded as supervision. To fill labels of other pixels, two types of mixed auxiliary supervision are proposed: a soft label learned from intrinsic structures of noisy annotations, and raters' characteristics labels which propagate individual rater's characteristics information. LF-Net has two main advantages. 1) Training with trustworthy pixels incorporates training with confident supervision, guiding the direction of groundtruth label learning. 2) Two types of mixed supervision prevent over-fitting issues when the network is supervised by a subset of pixels, and guarantee high fidelity with the true label. Results on five datasets of diverse imaging modalities show that our LF-Net boosts segmentation accuracy in all datasets compared with state-of-the-art methods, with even a 7% improvement in DSC for MS lesion segmentation.
RefineStyle: Dynamic Convolution Refinement for StyleGAN
Oct 08, 2024Abstract:In StyleGAN, convolution kernels are shaped by both static parameters shared across images and dynamic modulation factors $w^+\in\mathcal{W}^+$ specific to each image. Therefore, $\mathcal{W}^+$ space is often used for image inversion and editing. However, pre-trained model struggles with synthesizing out-of-domain images due to the limited capabilities of $\mathcal{W}^+$ and its resultant kernels, necessitating full fine-tuning or adaptation through a complex hypernetwork. This paper proposes an efficient refining strategy for dynamic kernels. The key idea is to modify kernels by low-rank residuals, learned from input image or domain guidance. These residuals are generated by matrix multiplication between two sets of tokens with the same number, which controls the complexity. We validate the refining scheme in image inversion and domain adaptation. In the former task, we design grouped transformer blocks to learn these token sets by one- or two-stage training. In the latter task, token sets are directly optimized to support synthesis in the target domain while preserving original content. Extensive experiments show that our method achieves low distortions for image inversion and high quality for out-of-domain editing.
Boosting Open-Domain Continual Learning via Leveraging Intra-domain Category-aware Prototype
Aug 19, 2024Abstract:Despite recent progress in enhancing the efficacy of Open-Domain Continual Learning (ODCL) in Vision-Language Models (VLM), failing to (1) correctly identify the Task-ID of a test image and (2) use only the category set corresponding to the Task-ID, while preserving the knowledge related to each domain, cannot address the two primary challenges of ODCL: forgetting old knowledge and maintaining zero-shot capabilities, as well as the confusions caused by category-relatedness between domains. In this paper, we propose a simple yet effective solution: leveraging intra-domain category-aware prototypes for ODCL in CLIP (DPeCLIP), where the prototype is the key to bridging the above two processes. Concretely, we propose a training-free Task-ID discriminator method, by utilizing prototypes as classifiers for identifying Task-IDs. Furthermore, to maintain the knowledge corresponding to each domain, we incorporate intra-domain category-aware prototypes as domain prior prompts into the training process. Extensive experiments conducted on 11 different datasets demonstrate the effectiveness of our approach, achieving 2.37% and 1.14% average improvement in class-incremental and task-incremental settings, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge