Mohammad Yaqub
Advancing Fetal Ultrasound Image Quality Assessment in Low-Resource Settings
Jul 30, 2025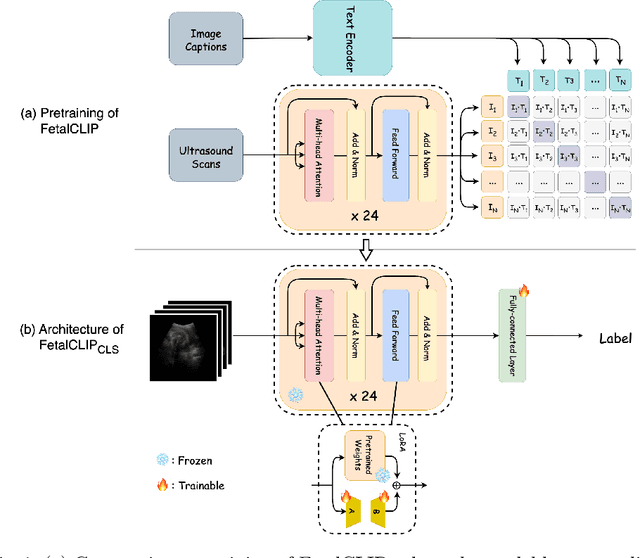
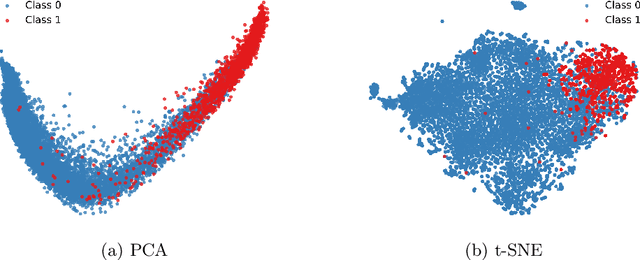
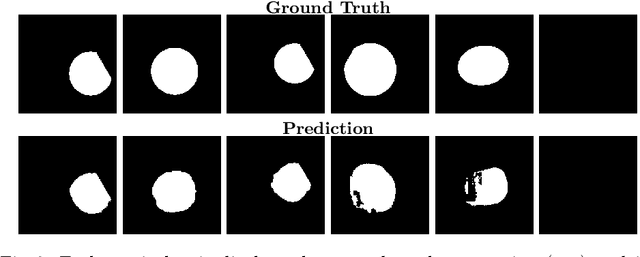
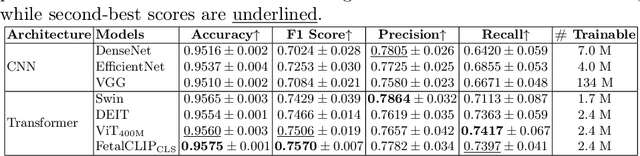


Abstract:Accurate fetal biometric measurements, such as abdominal circumference, play a vital role in prenatal care. However, obtaining high-quality ultrasound images for these measurements heavily depends on the expertise of sonographers, posing a significant challenge in low-income countries due to the scarcity of trained personnel. To address this issue, we leverage FetalCLIP, a vision-language model pretrained on a curated dataset of over 210,000 fetal ultrasound image-caption pairs, to perform automated fetal ultrasound image quality assessment (IQA) on blind-sweep ultrasound data. We introduce FetalCLIP$_{CLS}$, an IQA model adapted from FetalCLIP using Low-Rank Adaptation (LoRA), and evaluate it on the ACOUSLIC-AI dataset against six CNN and Transformer baselines. FetalCLIP$_{CLS}$ achieves the highest F1 score of 0.757. Moreover, we show that an adapted segmentation model, when repurposed for classification, further improves performance, achieving an F1 score of 0.771. Our work demonstrates how parameter-efficient fine-tuning of fetal ultrasound foundation models can enable task-specific adaptations, advancing prenatal care in resource-limited settings. The experimental code is available at: https://github.com/donglihe-hub/FetalCLIP-IQA.
TransPrune: Token Transition Pruning for Efficient Large Vision-Language Model
Jul 28, 2025Abstract:Large Vision-Language Models (LVLMs) have advanced multimodal learning but face high computational costs due to the large number of visual tokens, motivating token pruning to improve inference efficiency. The key challenge lies in identifying which tokens are truly important. Most existing approaches rely on attention-based criteria to estimate token importance. However, they inherently suffer from certain limitations, such as positional bias. In this work, we explore a new perspective on token importance based on token transitions in LVLMs. We observe that the transition of token representations provides a meaningful signal of semantic information. Based on this insight, we propose TransPrune, a training-free and efficient token pruning method. Specifically, TransPrune progressively prunes tokens by assessing their importance through a combination of Token Transition Variation (TTV)-which measures changes in both the magnitude and direction of token representations-and Instruction-Guided Attention (IGA), which measures how strongly the instruction attends to image tokens via attention. Extensive experiments demonstrate that TransPrune achieves comparable multimodal performance to original LVLMs, such as LLaVA-v1.5 and LLaVA-Next, across eight benchmarks, while reducing inference TFLOPs by more than half. Moreover, TTV alone can serve as an effective criterion without relying on attention, achieving performance comparable to attention-based methods. The code will be made publicly available upon acceptance of the paper at https://github.com/liaolea/TransPrune.
Not Only Grey Matter: OmniBrain for Robust Multimodal Classification of Alzheimer's Disease
Jul 28, 2025



Abstract:Alzheimer's disease affects over 55 million people worldwide and is projected to more than double by 2050, necessitating rapid, accurate, and scalable diagnostics. However, existing approaches are limited because they cannot achieve clinically acceptable accuracy, generalization across datasets, robustness to missing modalities, and explainability all at the same time. This inability to satisfy all these requirements simultaneously undermines their reliability in clinical settings. We propose OmniBrain, a multimodal framework that integrates brain MRI, radiomics, gene expression, and clinical data using a unified model with cross-attention and modality dropout. OmniBrain achieves $92.2 \pm 2.4\%$accuracy on the ANMerge dataset and generalizes to the MRI-only ADNI dataset with $70.4 \pm 2.7\%$ accuracy, outperforming unimodal and prior multimodal approaches. Explainability analyses highlight neuropathologically relevant brain regions and genes, enhancing clinical trust. OmniBrain offers a robust, interpretable, and practical solution for real-world Alzheimer's diagnosis.
crossMoDA Challenge: Evolution of Cross-Modality Domain Adaptation Techniques for Vestibular Schwannoma and Cochlea Segmentation from 2021 to 2023
Jun 13, 2025Abstract:The cross-Modality Domain Adaptation (crossMoDA) challenge series, initiated in 2021 in conjunction with the International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI), focuses on unsupervised cross-modality segmentation, learning from contrast-enhanced T1 (ceT1) and transferring to T2 MRI. The task is an extreme example of domain shift chosen to serve as a meaningful and illustrative benchmark. From a clinical application perspective, it aims to automate Vestibular Schwannoma (VS) and cochlea segmentation on T2 scans for more cost-effective VS management. Over time, the challenge objectives have evolved to enhance its clinical relevance. The challenge evolved from using single-institutional data and basic segmentation in 2021 to incorporating multi-institutional data and Koos grading in 2022, and by 2023, it included heterogeneous routine data and sub-segmentation of intra- and extra-meatal tumour components. In this work, we report the findings of the 2022 and 2023 editions and perform a retrospective analysis of the challenge progression over the years. The observations from the successive challenge contributions indicate that the number of outliers decreases with an expanding dataset. This is notable since the diversity of scanning protocols of the datasets concurrently increased. The winning approach of the 2023 edition reduced the number of outliers on the 2021 and 2022 testing data, demonstrating how increased data heterogeneity can enhance segmentation performance even on homogeneous data. However, the cochlea Dice score declined in 2023, likely due to the added complexity from tumour sub-annotations affecting overall segmentation performance. While progress is still needed for clinically acceptable VS segmentation, the plateauing performance suggests that a more challenging cross-modal task may better serve future benchmarking.
On the Robustness of Medical Vision-Language Models: Are they Truly Generalizable?
May 21, 2025Abstract:Medical Vision-Language Models (MVLMs) have achieved par excellence generalization in medical image analysis, yet their performance under noisy, corrupted conditions remains largely untested. Clinical imaging is inherently susceptible to acquisition artifacts and noise; however, existing evaluations predominantly assess generally clean datasets, overlooking robustness -- i.e., the model's ability to perform under real-world distortions. To address this gap, we first introduce MediMeta-C, a corruption benchmark that systematically applies several perturbations across multiple medical imaging datasets. Combined with MedMNIST-C, this establishes a comprehensive robustness evaluation framework for MVLMs. We further propose RobustMedCLIP, a visual encoder adaptation of a pretrained MVLM that incorporates few-shot tuning to enhance resilience against corruptions. Through extensive experiments, we benchmark 5 major MVLMs across 5 medical imaging modalities, revealing that existing models exhibit severe degradation under corruption and struggle with domain-modality tradeoffs. Our findings highlight the necessity of diverse training and robust adaptation strategies, demonstrating that efficient low-rank adaptation when paired with few-shot tuning, improves robustness while preserving generalization across modalities.
Kalman Filter Enhanced GRPO for Reinforcement Learning-Based Language Model Reasoning
May 12, 2025Abstract:Reward baseline is important for Reinforcement Learning (RL) algorithms to reduce variance in policy gradient estimates. Recently, for language modeling, Group Relative Policy Optimization (GRPO) is proposed to compute the advantage for each output by subtracting the mean reward, as the baseline, for all outputs in the group. However, it can lead to inaccurate advantage estimates in environments with highly noisy rewards, potentially introducing bias. In this work, we propose a model, called Kalman Filter Enhanced Group Relative Policy Optimization (KRPO), by using lightweight Kalman filtering to dynamically estimate the latent reward mean and variance. This filtering technique replaces the naive batch mean baseline, enabling more adaptive advantage normalization. Our method does not require additional learned parameters over GRPO. This approach offers a simple yet effective way to incorporate multiple outputs of GRPO into advantage estimation, improving policy optimization in settings where highly dynamic reward signals are difficult to model for language models. Through experiments and analyses, we show that using a more adaptive advantage estimation model, KRPO can improve the stability and performance of GRPO. The code is available at https://github.com/billhhh/KRPO_LLMs_RL
Efficient Parameter Adaptation for Multi-Modal Medical Image Segmentation and Prognosis
Apr 18, 2025



Abstract:Cancer detection and prognosis relies heavily on medical imaging, particularly CT and PET scans. Deep Neural Networks (DNNs) have shown promise in tumor segmentation by fusing information from these modalities. However, a critical bottleneck exists: the dependency on CT-PET data concurrently for training and inference, posing a challenge due to the limited availability of PET scans. Hence, there is a clear need for a flexible and efficient framework that can be trained with the widely available CT scans and can be still adapted for PET scans when they become available. In this work, we propose a parameter-efficient multi-modal adaptation (PEMMA) framework for lightweight upgrading of a transformer-based segmentation model trained only on CT scans such that it can be efficiently adapted for use with PET scans when they become available. This framework is further extended to perform prognosis task maintaining the same efficient cross-modal fine-tuning approach. The proposed approach is tested with two well-known segementation backbones, namely UNETR and Swin UNETR. Our approach offers two main advantages. Firstly, we leverage the inherent modularity of the transformer architecture and perform low-rank adaptation (LoRA) as well as decomposed low-rank adaptation (DoRA) of the attention weights to achieve parameter-efficient adaptation. Secondly, by minimizing cross-modal entanglement, PEMMA allows updates using only one modality without causing catastrophic forgetting in the other. Our method achieves comparable performance to early fusion, but with only 8% of the trainable parameters, and demonstrates a significant +28% Dice score improvement on PET scans when trained with a single modality. Furthermore, in prognosis, our method improves the concordance index by +10% when adapting a CT-pretrained model to include PET scans, and by +23% when adapting for both PET and EHR data.
SALT: Singular Value Adaptation with Low-Rank Transformation
Mar 20, 2025



Abstract:The complex nature of medical image segmentation calls for models that are specifically designed to capture detailed, domain-specific features. Large foundation models offer considerable flexibility, yet the cost of fine-tuning these models remains a significant barrier. Parameter-Efficient Fine-Tuning (PEFT) methods, such as Low-Rank Adaptation (LoRA), efficiently update model weights with low-rank matrices but may suffer from underfitting when the chosen rank is insufficient to capture domain-specific nuances. Conversely, full-rank Singular Value Decomposition (SVD) based methods provide comprehensive updates by modifying all singular values, yet they often lack flexibility and exhibit variable performance across datasets. We propose SALT (Singular Value Adaptation with Low-Rank Transformation), a method that selectively adapts the most influential singular values using trainable scale and shift parameters while complementing this with a low-rank update for the remaining subspace. This hybrid approach harnesses the advantages of both LoRA and SVD, enabling effective adaptation without relying on increasing model size or depth. Evaluated on 5 challenging medical datasets, ranging from as few as 20 samples to 1000, SALT outperforms state-of-the-art PEFT (LoRA and SVD) by 2% to 5% in Dice with only 3.9% trainable parameters, demonstrating robust adaptation even in low-resource settings. The code for SALT is available at: https://github.com/BioMedIA-MBZUAI/SALT
Segmentation-Guided CT Synthesis with Pixel-Wise Conformal Uncertainty Bounds
Mar 11, 2025Abstract:Accurate dose calculations in proton therapy rely on high-quality CT images. While planning CTs (pCTs) serve as a reference for dosimetric planning, Cone Beam CT (CBCT) is used throughout Adaptive Radiotherapy (ART) to generate sCTs for improved dose calculations. Despite its lower cost and reduced radiation exposure advantages, CBCT suffers from severe artefacts and poor image quality, making it unsuitable for precise dosimetry. Deep learning-based CBCT-to-CT translation has emerged as a promising approach. Still, existing methods often introduce anatomical inconsistencies and lack reliable uncertainty estimates, limiting their clinical adoption. To bridge this gap, we propose STF-RUE, a novel framework integrating two key components. First, STF, a segmentation-guided CBCT-to-CT translation method that enhances anatomical consistency by leveraging segmentation priors extracted from pCTs. Second, RUE, a conformal prediction method that augments predicted CTs with pixel-wise conformal prediction intervals, providing clinicians with robust reliability indicator. Comprehensive experiments using UNet++ and Fast-DDPM on two benchmark datasets demonstrate that STF-RUE significantly improves translation accuracy, as measured by a novel soft-tissue-focused metric designed for precise dose computation. Additionally, STF-RUE provides better-calibrated uncertainty sets for synthetic CT, reinforcing trust in synthetic CTs. By addressing both anatomical fidelity and uncertainty quantification, STF-RUE marks a crucial step toward safer and more effective adaptive proton therapy. Code is available at https://anonymous.4open.science/r/cbct2ct_translation-B2D9/.
In-Model Merging for Enhancing the Robustness of Medical Imaging Classification Models
Feb 27, 2025



Abstract:Model merging is an effective strategy to merge multiple models for enhancing model performances, and more efficient than ensemble learning as it will not introduce extra computation into inference. However, limited research explores if the merging process can occur within one model and enhance the model's robustness, which is particularly critical in the medical image domain. In the paper, we are the first to propose in-model merging (InMerge), a novel approach that enhances the model's robustness by selectively merging similar convolutional kernels in the deep layers of a single convolutional neural network (CNN) during the training process for classification. We also analytically reveal important characteristics that affect how in-model merging should be performed, serving as an insightful reference for the community. We demonstrate the feasibility and effectiveness of this technique for different CNN architectures on 4 prevalent datasets. The proposed InMerge-trained model surpasses the typically-trained model by a substantial margin. The code will be made public.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge