Michael Baumgartner
Revisiting 2D Foundation Models for Scalable 3D Medical Image Classification
Dec 15, 2025



Abstract:3D medical image classification is essential for modern clinical workflows. Medical foundation models (FMs) have emerged as a promising approach for scaling to new tasks, yet current research suffers from three critical pitfalls: data-regime bias, suboptimal adaptation, and insufficient task coverage. In this paper, we address these pitfalls and introduce AnyMC3D, a scalable 3D classifier adapted from 2D FMs. Our method scales efficiently to new tasks by adding only lightweight plugins (about 1M parameters per task) on top of a single frozen backbone. This versatile framework also supports multi-view inputs, auxiliary pixel-level supervision, and interpretable heatmap generation. We establish a comprehensive benchmark of 12 tasks covering diverse pathologies, anatomies, and modalities, and systematically analyze state-of-the-art 3D classification techniques. Our analysis reveals key insights: (1) effective adaptation is essential to unlock FM potential, (2) general-purpose FMs can match medical-specific FMs if properly adapted, and (3) 2D-based methods surpass 3D architectures for 3D classification. For the first time, we demonstrate the feasibility of achieving state-of-the-art performance across diverse applications using a single scalable framework (including 1st place in the VLM3D challenge), eliminating the need for separate task-specific models.
The Missing Piece: A Case for Pre-Training in 3D Medical Object Detection
Sep 19, 2025Abstract:Large-scale pre-training holds the promise to advance 3D medical object detection, a crucial component of accurate computer-aided diagnosis. Yet, it remains underexplored compared to segmentation, where pre-training has already demonstrated significant benefits. Existing pre-training approaches for 3D object detection rely on 2D medical data or natural image pre-training, failing to fully leverage 3D volumetric information. In this work, we present the first systematic study of how existing pre-training methods can be integrated into state-of-the-art detection architectures, covering both CNNs and Transformers. Our results show that pre-training consistently improves detection performance across various tasks and datasets. Notably, reconstruction-based self-supervised pre-training outperforms supervised pre-training, while contrastive pre-training provides no clear benefit for 3D medical object detection. Our code is publicly available at: https://github.com/MIC-DKFZ/nnDetection-finetuning.
* MICCAI 2025
Advances in Automated Fetal Brain MRI Segmentation and Biometry: Insights from the FeTA 2024 Challenge
May 05, 2025


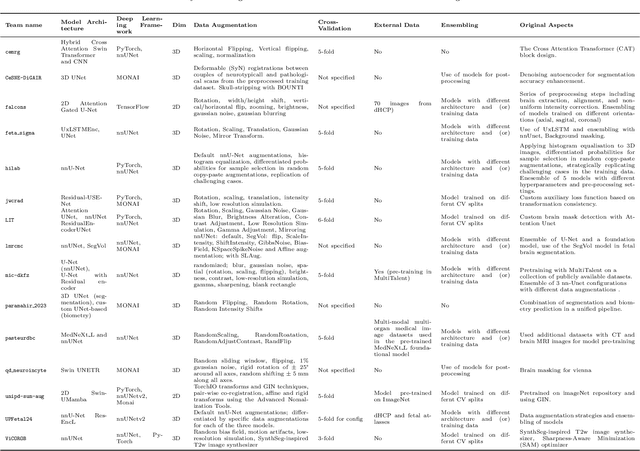
Abstract:Accurate fetal brain tissue segmentation and biometric analysis are essential for studying brain development in utero. The FeTA Challenge 2024 advanced automated fetal brain MRI analysis by introducing biometry prediction as a new task alongside tissue segmentation. For the first time, our diverse multi-centric test set included data from a new low-field (0.55T) MRI dataset. Evaluation metrics were also expanded to include the topology-specific Euler characteristic difference (ED). Sixteen teams submitted segmentation methods, most of which performed consistently across both high- and low-field scans. However, longitudinal trends indicate that segmentation accuracy may be reaching a plateau, with results now approaching inter-rater variability. The ED metric uncovered topological differences that were missed by conventional metrics, while the low-field dataset achieved the highest segmentation scores, highlighting the potential of affordable imaging systems when paired with high-quality reconstruction. Seven teams participated in the biometry task, but most methods failed to outperform a simple baseline that predicted measurements based solely on gestational age, underscoring the challenge of extracting reliable biometric estimates from image data alone. Domain shift analysis identified image quality as the most significant factor affecting model generalization, with super-resolution pipelines also playing a substantial role. Other factors, such as gestational age, pathology, and acquisition site, had smaller, though still measurable, effects. Overall, FeTA 2024 offers a comprehensive benchmark for multi-class segmentation and biometry estimation in fetal brain MRI, underscoring the need for data-centric approaches, improved topological evaluation, and greater dataset diversity to enable clinically robust and generalizable AI tools.
Primus: Enforcing Attention Usage for 3D Medical Image Segmentation
Mar 03, 2025



Abstract:Transformers have achieved remarkable success across multiple fields, yet their impact on 3D medical image segmentation remains limited with convolutional networks still dominating major benchmarks. In this work, we a) analyze current Transformer-based segmentation models and identify critical shortcomings, particularly their over-reliance on convolutional blocks. Further, we demonstrate that in some architectures, performance is unaffected by the absence of the Transformer, thereby demonstrating their limited effectiveness. To address these challenges, we move away from hybrid architectures and b) introduce a fully Transformer-based segmentation architecture, termed Primus. Primus leverages high-resolution tokens, combined with advances in positional embeddings and block design, to maximally leverage its Transformer blocks. Through these adaptations Primus surpasses current Transformer-based methods and competes with state-of-the-art convolutional models on multiple public datasets. By doing so, we create the first pure Transformer architecture and take a significant step towards making Transformers state-of-the-art for 3D medical image segmentation.
Multi-Class Segmentation of Aortic Branches and Zones in Computed Tomography Angiography: The AortaSeg24 Challenge
Feb 07, 2025



Abstract:Multi-class segmentation of the aorta in computed tomography angiography (CTA) scans is essential for diagnosing and planning complex endovascular treatments for patients with aortic dissections. However, existing methods reduce aortic segmentation to a binary problem, limiting their ability to measure diameters across different branches and zones. Furthermore, no open-source dataset is currently available to support the development of multi-class aortic segmentation methods. To address this gap, we organized the AortaSeg24 MICCAI Challenge, introducing the first dataset of 100 CTA volumes annotated for 23 clinically relevant aortic branches and zones. This dataset was designed to facilitate both model development and validation. The challenge attracted 121 teams worldwide, with participants leveraging state-of-the-art frameworks such as nnU-Net and exploring novel techniques, including cascaded models, data augmentation strategies, and custom loss functions. We evaluated the submitted algorithms using the Dice Similarity Coefficient (DSC) and Normalized Surface Distance (NSD), highlighting the approaches adopted by the top five performing teams. This paper presents the challenge design, dataset details, evaluation metrics, and an in-depth analysis of the top-performing algorithms. The annotated dataset, evaluation code, and implementations of the leading methods are publicly available to support further research. All resources can be accessed at https://aortaseg24.grand-challenge.org.
Tumor Detection, Segmentation and Classification Challenge on Automated 3D Breast Ultrasound: The TDSC-ABUS Challenge
Jan 26, 2025



Abstract:Breast cancer is one of the most common causes of death among women worldwide. Early detection helps in reducing the number of deaths. Automated 3D Breast Ultrasound (ABUS) is a newer approach for breast screening, which has many advantages over handheld mammography such as safety, speed, and higher detection rate of breast cancer. Tumor detection, segmentation, and classification are key components in the analysis of medical images, especially challenging in the context of 3D ABUS due to the significant variability in tumor size and shape, unclear tumor boundaries, and a low signal-to-noise ratio. The lack of publicly accessible, well-labeled ABUS datasets further hinders the advancement of systems for breast tumor analysis. Addressing this gap, we have organized the inaugural Tumor Detection, Segmentation, and Classification Challenge on Automated 3D Breast Ultrasound 2023 (TDSC-ABUS2023). This initiative aims to spearhead research in this field and create a definitive benchmark for tasks associated with 3D ABUS image analysis. In this paper, we summarize the top-performing algorithms from the challenge and provide critical analysis for ABUS image examination. We offer the TDSC-ABUS challenge as an open-access platform at https://tdsc-abus2023.grand-challenge.org/ to benchmark and inspire future developments in algorithmic research.
Unlocking the Potential of Digital Pathology: Novel Baselines for Compression
Dec 17, 2024



Abstract:Digital pathology offers a groundbreaking opportunity to transform clinical practice in histopathological image analysis, yet faces a significant hurdle: the substantial file sizes of pathological Whole Slide Images (WSI). While current digital pathology solutions rely on lossy JPEG compression to address this issue, lossy compression can introduce color and texture disparities, potentially impacting clinical decision-making. While prior research addresses perceptual image quality and downstream performance independently of each other, we jointly evaluate compression schemes for perceptual and downstream task quality on four different datasets. In addition, we collect an initially uncompressed dataset for an unbiased perceptual evaluation of compression schemes. Our results show that deep learning models fine-tuned for perceptual quality outperform conventional compression schemes like JPEG-XL or WebP for further compression of WSI. However, they exhibit a significant bias towards the compression artifacts present in the training data and struggle to generalize across various compression schemes. We introduce a novel evaluation metric based on feature similarity between original files and compressed files that aligns very well with the actual downstream performance on the compressed WSI. Our metric allows for a general and standardized evaluation of lossy compression schemes and mitigates the requirement to independently assess different downstream tasks. Our study provides novel insights for the assessment of lossy compression schemes for WSI and encourages a unified evaluation of lossy compression schemes to accelerate the clinical uptake of digital pathology.
Touchstone Benchmark: Are We on the Right Way for Evaluating AI Algorithms for Medical Segmentation?
Nov 06, 2024



Abstract:How can we test AI performance? This question seems trivial, but it isn't. Standard benchmarks often have problems such as in-distribution and small-size test sets, oversimplified metrics, unfair comparisons, and short-term outcome pressure. As a consequence, good performance on standard benchmarks does not guarantee success in real-world scenarios. To address these problems, we present Touchstone, a large-scale collaborative segmentation benchmark of 9 types of abdominal organs. This benchmark is based on 5,195 training CT scans from 76 hospitals around the world and 5,903 testing CT scans from 11 additional hospitals. This diverse test set enhances the statistical significance of benchmark results and rigorously evaluates AI algorithms across various out-of-distribution scenarios. We invited 14 inventors of 19 AI algorithms to train their algorithms, while our team, as a third party, independently evaluated these algorithms on three test sets. In addition, we also evaluated pre-existing AI frameworks--which, differing from algorithms, are more flexible and can support different algorithms--including MONAI from NVIDIA, nnU-Net from DKFZ, and numerous other open-source frameworks. We are committed to expanding this benchmark to encourage more innovation of AI algorithms for the medical domain.
Decoupling Semantic Similarity from Spatial Alignment for Neural Networks
Oct 30, 2024



Abstract:What representation do deep neural networks learn? How similar are images to each other for neural networks? Despite the overwhelming success of deep learning methods key questions about their internal workings still remain largely unanswered, due to their internal high dimensionality and complexity. To address this, one approach is to measure the similarity of activation responses to various inputs. Representational Similarity Matrices (RSMs) distill this similarity into scalar values for each input pair. These matrices encapsulate the entire similarity structure of a system, indicating which input leads to similar responses. While the similarity between images is ambiguous, we argue that the spatial location of semantic objects does neither influence human perception nor deep learning classifiers. Thus this should be reflected in the definition of similarity between image responses for computer vision systems. Revisiting the established similarity calculations for RSMs we expose their sensitivity to spatial alignment. In this paper, we propose to solve this through semantic RSMs, which are invariant to spatial permutation. We measure semantic similarity between input responses by formulating it as a set-matching problem. Further, we quantify the superiority of semantic RSMs over spatio-semantic RSMs through image retrieval and by comparing the similarity between representations to the similarity between predicted class probabilities.
Overcoming Common Flaws in the Evaluation of Selective Classification Systems
Jul 01, 2024



Abstract:Selective Classification, wherein models can reject low-confidence predictions, promises reliable translation of machine-learning based classification systems to real-world scenarios such as clinical diagnostics. While current evaluation of these systems typically assumes fixed working points based on pre-defined rejection thresholds, methodological progress requires benchmarking the general performance of systems akin to the $\mathrm{AUROC}$ in standard classification. In this work, we define 5 requirements for multi-threshold metrics in selective classification regarding task alignment, interpretability, and flexibility, and show how current approaches fail to meet them. We propose the Area under the Generalized Risk Coverage curve ($\mathrm{AUGRC}$), which meets all requirements and can be directly interpreted as the average risk of undetected failures. We empirically demonstrate the relevance of $\mathrm{AUGRC}$ on a comprehensive benchmark spanning 6 data sets and 13 confidence scoring functions. We find that the proposed metric substantially changes metric rankings on 5 out of the 6 data sets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge