Kai Ma
School of Electrical Engineering, Yanshan University, Qinhuangdao, China
Double-Uncertainty Assisted Spatial and Temporal Regularization Weighting for Learning-based Registration
Jul 15, 2021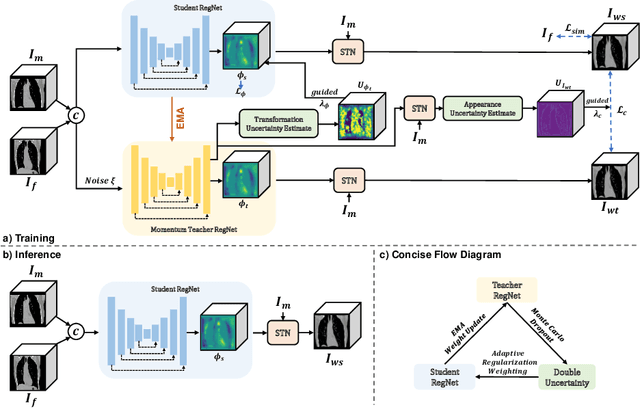
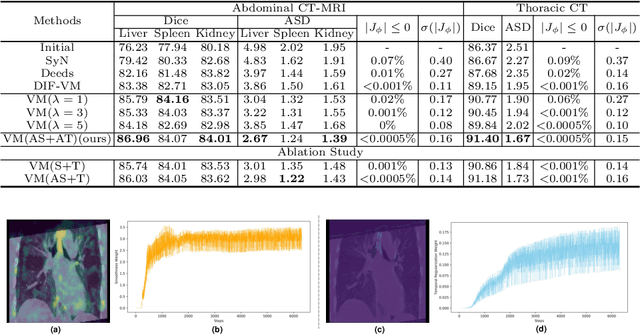
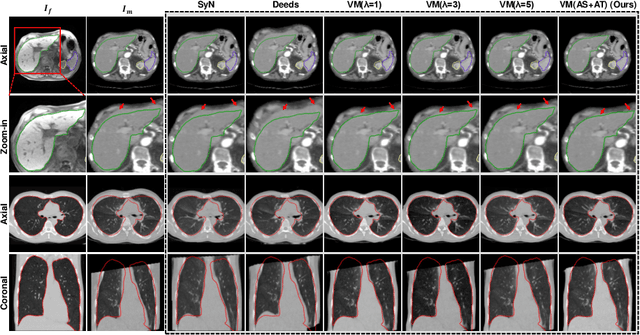
Abstract:In order to tackle the difficulty associated with the ill-posed nature of the image registration problem, researchers use regularization to constrain the solution space. For most learning-based registration approaches, the regularization usually has a fixed weight and only constrains the spatial transformation. Such convention has two limitations: (1) The regularization strength of a specific image pair should be associated with the content of the images, thus the ``one value fits all'' scheme is not ideal; (2) Only spatially regularizing the transformation (but overlooking the temporal consistency of different estimations) may not be the best strategy to cope with the ill-posedness. In this study, we propose a mean-teacher based registration framework. This framework incorporates an additional \textit{temporal regularization} term by encouraging the teacher model's temporal ensemble prediction to be consistent with that of the student model. At each training step, it also automatically adjusts the weights of the \textit{spatial regularization} and the \textit{temporal regularization} by taking account of the transformation uncertainty and appearance uncertainty derived from the perturbed teacher model. We perform experiments on multi- and uni-modal registration tasks, and the results show that our strategy outperforms the traditional and learning-based benchmark methods.
Mutual-GAN: Towards Unsupervised Cross-Weather Adaptation with Mutual Information Constraint
Jun 30, 2021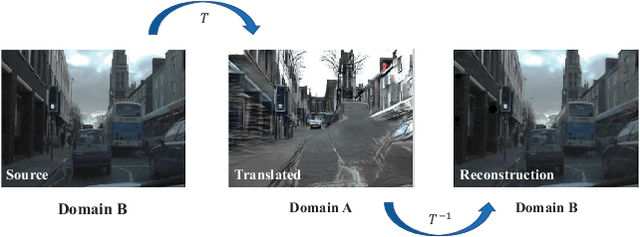
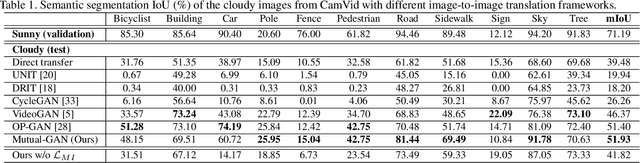
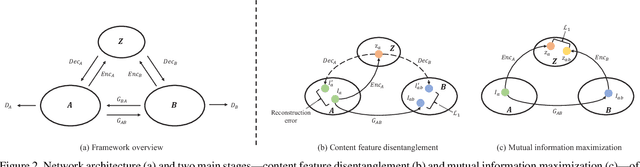
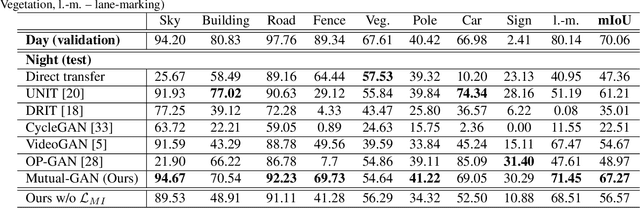
Abstract:Convolutional neural network (CNN) have proven its success for semantic segmentation, which is a core task of emerging industrial applications such as autonomous driving. However, most progress in semantic segmentation of urban scenes is reported on standard scenarios, i.e., daytime scenes with favorable illumination conditions. In practical applications, the outdoor weather and illumination are changeable, e.g., cloudy and nighttime, which results in a significant drop of semantic segmentation accuracy of CNN only trained with daytime data. In this paper, we propose a novel generative adversarial network (namely Mutual-GAN) to alleviate the accuracy decline when daytime-trained neural network is applied to videos captured under adverse weather conditions. The proposed Mutual-GAN adopts mutual information constraint to preserve image-objects during cross-weather adaptation, which is an unsolved problem for most unsupervised image-to-image translation approaches (e.g., CycleGAN). The proposed Mutual-GAN is evaluated on two publicly available driving video datasets (i.e., CamVid and SYNTHIA). The experimental results demonstrate that our Mutual-GAN can yield visually plausible translated images and significantly improve the semantic segmentation accuracy of daytime-trained deep learning network while processing videos under challenging weathers.
Residual Moment Loss for Medical Image Segmentation
Jun 27, 2021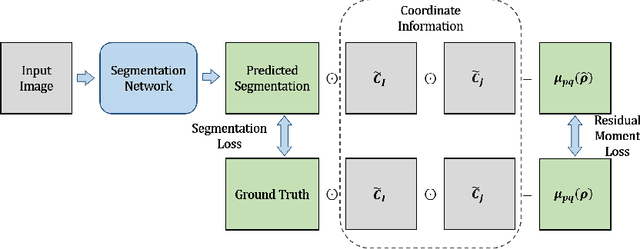
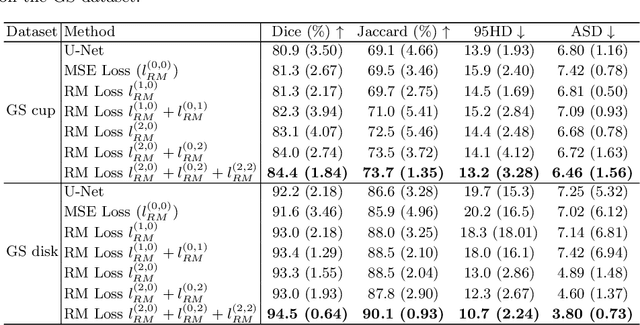
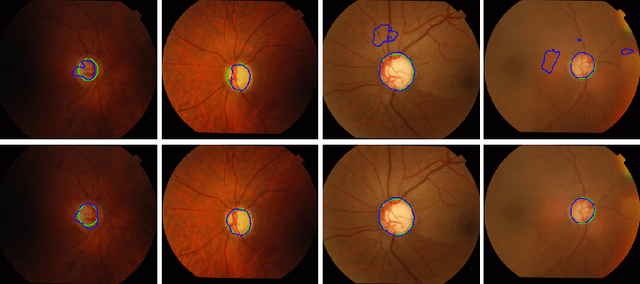
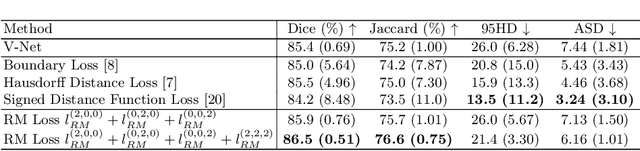
Abstract:Location information is proven to benefit the deep learning models on capturing the manifold structure of target objects, and accordingly boosts the accuracy of medical image segmentation. However, most existing methods encode the location information in an implicit way, e.g. the distance transform maps, which describe the relative distance from each pixel to the contour boundary, for the network to learn. These implicit approaches do not fully exploit the position information (i.e. absolute location) of targets. In this paper, we propose a novel loss function, namely residual moment (RM) loss, to explicitly embed the location information of segmentation targets during the training of deep learning networks. Particularly, motivated by image moments, the segmentation prediction map and ground-truth map are weighted by coordinate information. Then our RM loss encourages the networks to maintain the consistency between the two weighted maps, which promotes the segmentation networks to easily locate the targets and extract manifold-structure-related features. We validate the proposed RM loss by conducting extensive experiments on two publicly available datasets, i.e., 2D optic cup and disk segmentation and 3D left atrial segmentation. The experimental results demonstrate the effectiveness of our RM loss, which significantly boosts the accuracy of segmentation networks.
LE-NAS: Learning-based Ensenble with NAS for Dose Prediction
Jun 12, 2021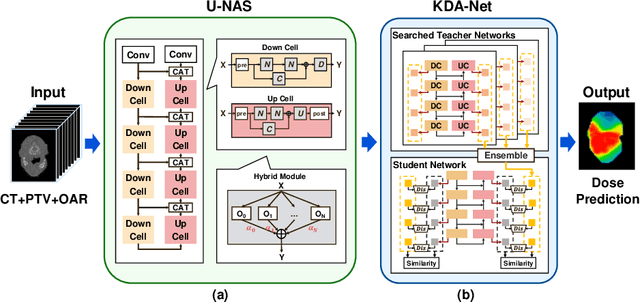
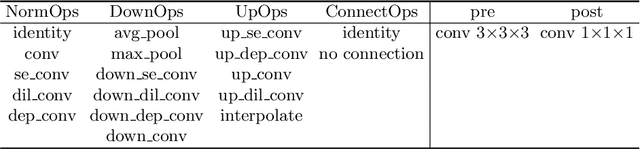

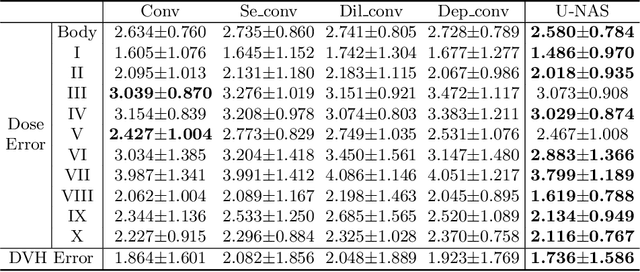
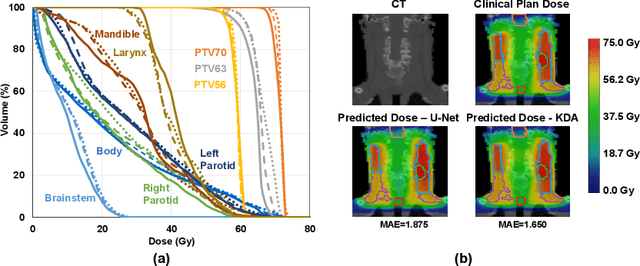
Abstract:Radiation therapy treatment planning is a complex process, as the target dose prescription and normal tissue sparing are conflicting objectives. Automated and accurate dose prediction for radiation therapy planning is in high demand. In this study, we propose a novel learning-based ensemble approach, named LE-NAS, which integrates neural architecture search (NAS) with knowledge distillation for 3D radiotherapy dose prediction. Specifically, the prediction network first exhaustively searches each block from enormous architecture space. Then, multiple architectures are selected with promising performance and diversity. To reduce the inference time, we adopt the teacher-student paradigm by treating the combination of diverse outputs from multiple searched networks as supervisions to guide the student network training. In addition, we apply adversarial learning to optimize the student network to recover the knowledge in teacher networks. To the best of our knowledge, we are the first to investigate the combination of NAS and knowledge distillation. The proposed method has been evaluated on the public OpenKBP dataset, and experimental results demonstrate the effectiveness of our method and its superior performance to the state-of-the-art method.
Noisy Labels are Treasure: Mean-Teacher-Assisted Confident Learning for Hepatic Vessel Segmentation
Jun 03, 2021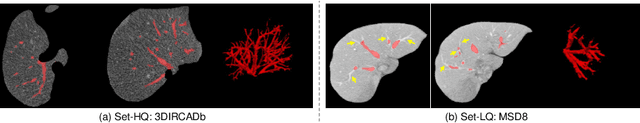
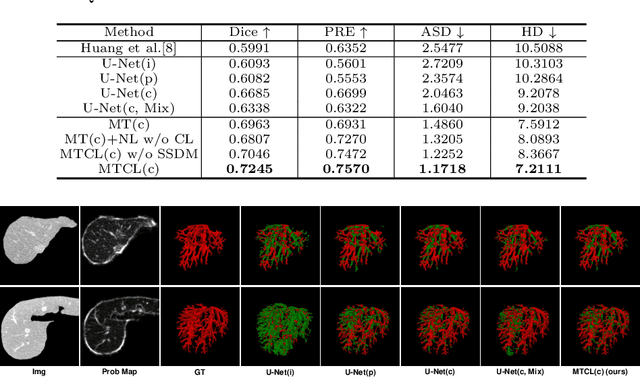
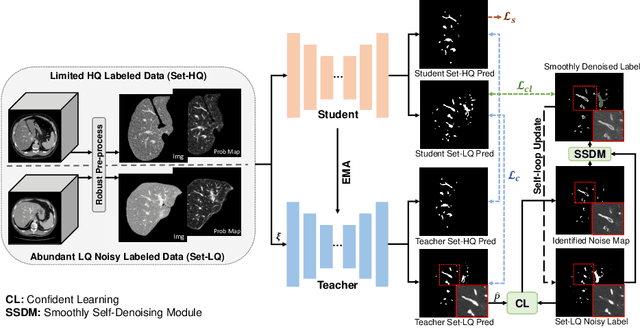
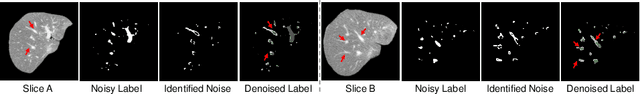
Abstract:Manually segmenting the hepatic vessels from Computer Tomography (CT) is far more expertise-demanding and laborious than other structures due to the low-contrast and complex morphology of vessels, resulting in the extreme lack of high-quality labeled data. Without sufficient high-quality annotations, the usual data-driven learning-based approaches struggle with deficient training. On the other hand, directly introducing additional data with low-quality annotations may confuse the network, leading to undesirable performance degradation. To address this issue, we propose a novel mean-teacher-assisted confident learning framework to robustly exploit the noisy labeled data for the challenging hepatic vessel segmentation task. Specifically, with the adapted confident learning assisted by a third party, i.e., the weight-averaged teacher model, the noisy labels in the additional low-quality dataset can be transformed from "encumbrance" to "treasure" via progressive pixel-wise soft-correction, thus providing productive guidance. Extensive experiments using two public datasets demonstrate the superiority of the proposed framework as well as the effectiveness of each component.
Generalized Organ Segmentation by Imitating One-shot Reasoning using Anatomical Correlation
Mar 30, 2021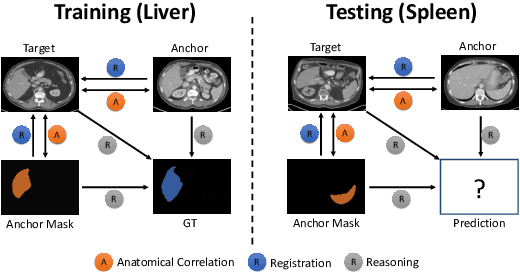
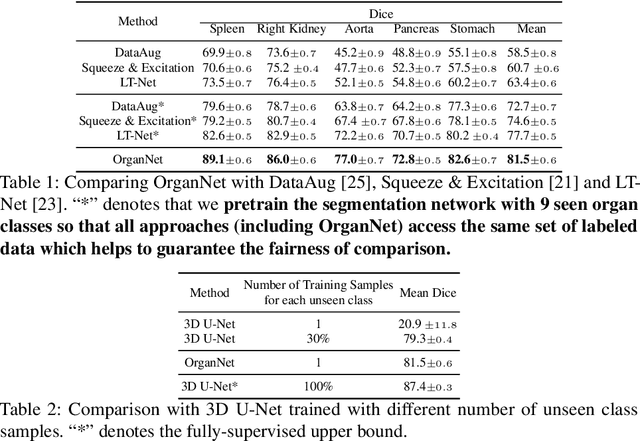
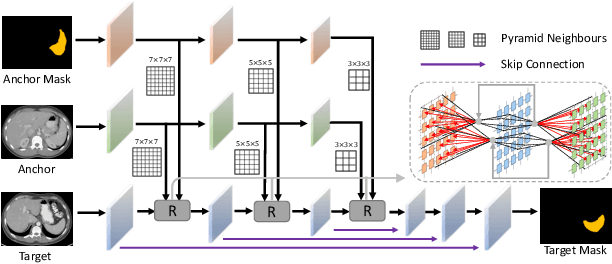
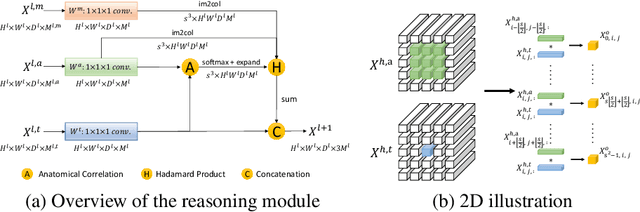
Abstract:Learning by imitation is one of the most significant abilities of human beings and plays a vital role in human's computational neural system. In medical image analysis, given several exemplars (anchors), experienced radiologist has the ability to delineate unfamiliar organs by imitating the reasoning process learned from existing types of organs. Inspired by this observation, we propose OrganNet which learns a generalized organ concept from a set of annotated organ classes and then transfer this concept to unseen classes. In this paper, we show that such process can be integrated into the one-shot segmentation task which is a very challenging but meaningful topic. We propose pyramid reasoning modules (PRMs) to model the anatomical correlation between anchor and target volumes. In practice, the proposed module first computes a correlation matrix between target and anchor computerized tomography (CT) volumes. Then, this matrix is used to transform the feature representations of both anchor volume and its segmentation mask. Finally, OrganNet learns to fuse the representations from various inputs and predicts segmentation results for target volume. Extensive experiments show that OrganNet can effectively resist the wide variations in organ morphology and produce state-of-the-art results in one-shot segmentation task. Moreover, even when compared with fully-supervised segmentation models, OrganNet is still able to produce satisfying segmentation results.
Stabilized Medical Image Attacks
Mar 09, 2021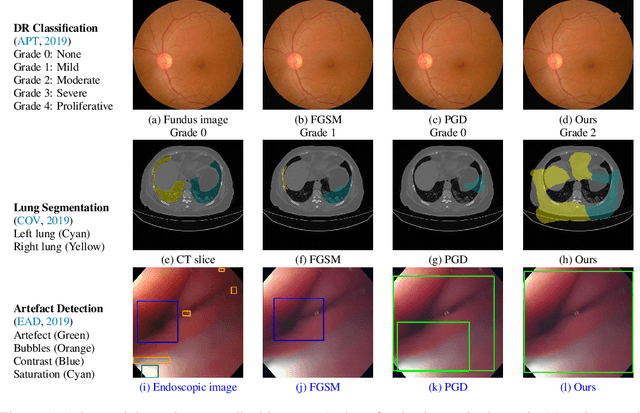
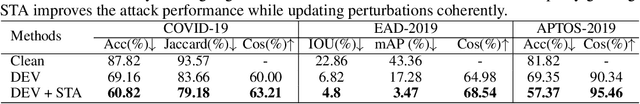
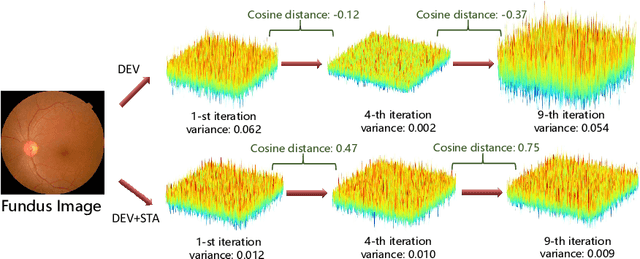
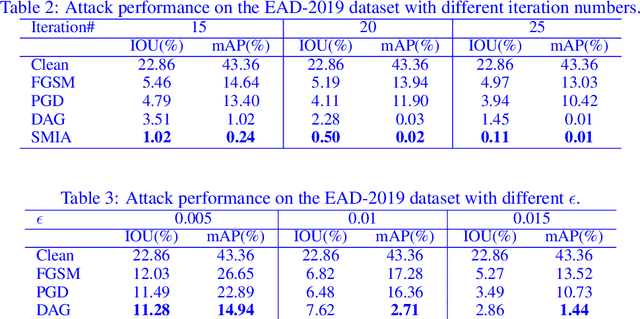
Abstract:Convolutional Neural Networks (CNNs) have advanced existing medical systems for automatic disease diagnosis. However, a threat to these systems arises that adversarial attacks make CNNs vulnerable. Inaccurate diagnosis results make a negative influence on human healthcare. There is a need to investigate potential adversarial attacks to robustify deep medical diagnosis systems. On the other side, there are several modalities of medical images (e.g., CT, fundus, and endoscopic image) of which each type is significantly different from others. It is more challenging to generate adversarial perturbations for different types of medical images. In this paper, we propose an image-based medical adversarial attack method to consistently produce adversarial perturbations on medical images. The objective function of our method consists of a loss deviation term and a loss stabilization term. The loss deviation term increases the divergence between the CNN prediction of an adversarial example and its ground truth label. Meanwhile, the loss stabilization term ensures similar CNN predictions of this example and its smoothed input. From the perspective of the whole iterations for perturbation generation, the proposed loss stabilization term exhaustively searches the perturbation space to smooth the single spot for local optimum escape. We further analyze the KL-divergence of the proposed loss function and find that the loss stabilization term makes the perturbations updated towards a fixed objective spot while deviating from the ground truth. This stabilization ensures the proposed medical attack effective for different types of medical images while producing perturbations in small variance. Experiments on several medical image analysis benchmarks including the recent COVID-19 dataset show the stability of the proposed method.
MixSearch: Searching for Domain Generalized Medical Image Segmentation Architectures
Feb 26, 2021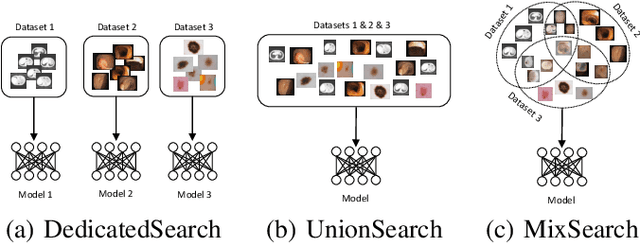
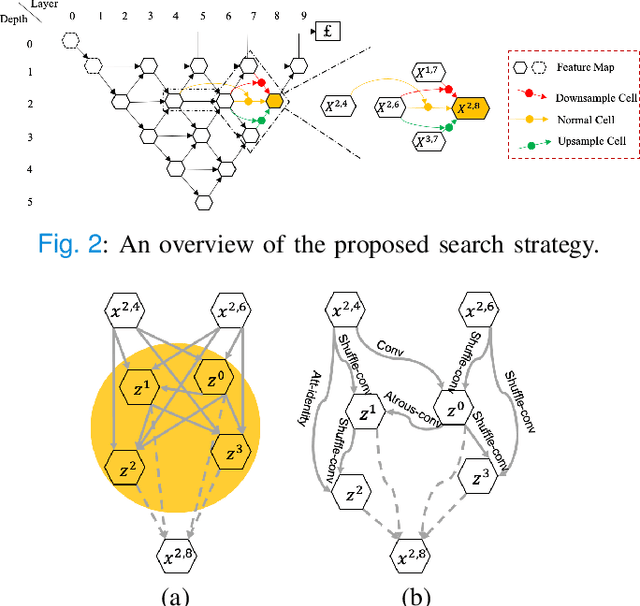
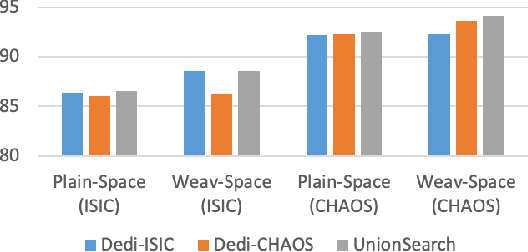
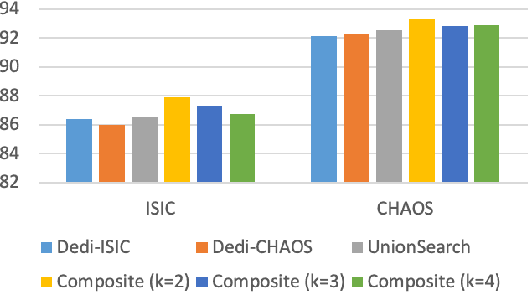
Abstract:Considering the scarcity of medical data, most datasets in medical image analysis are an order of magnitude smaller than those of natural images. However, most Network Architecture Search (NAS) approaches in medical images focused on specific datasets and did not take into account the generalization ability of the learned architectures on unseen datasets as well as different domains. In this paper, we address this point by proposing to search for generalizable U-shape architectures on a composited dataset that mixes medical images from multiple segmentation tasks and domains creatively, which is named MixSearch. Specifically, we propose a novel approach to mix multiple small-scale datasets from multiple domains and segmentation tasks to produce a large-scale dataset. Then, a novel weaved encoder-decoder structure is designed to search for a generalized segmentation network in both cell-level and network-level. The network produced by the proposed MixSearch framework achieves state-of-the-art results compared with advanced encoder-decoder networks across various datasets.
Ensembled ResUnet for Anatomical Brain Barriers Segmentation
Jan 04, 2021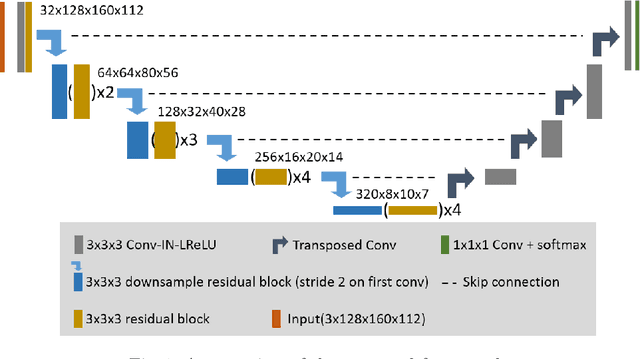
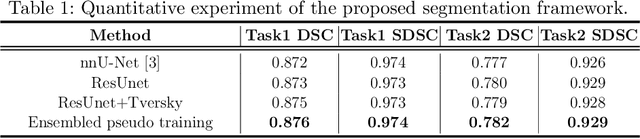
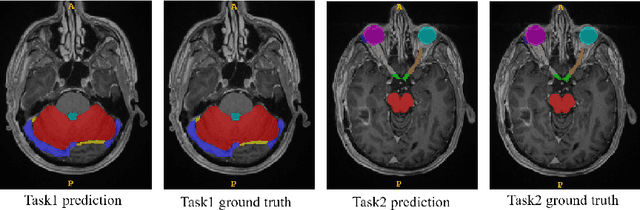
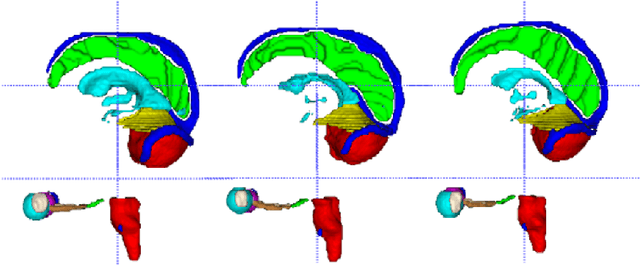
Abstract:Accuracy segmentation of brain structures could be helpful for glioma and radiotherapy planning. However, due to the visual and anatomical differences between different modalities, the accurate segmentation of brain structures becomes challenging. To address this problem, we first construct a residual block based U-shape network with a deep encoder and shallow decoder, which can trade off the framework performance and efficiency. Then, we introduce the Tversky loss to address the issue of the class imbalance between different foreground and the background classes. Finally, a model ensemble strategy is utilized to remove outliers and further boost performance.
A Hierarchical Feature Constraint to Camouflage Medical Adversarial Attacks
Dec 17, 2020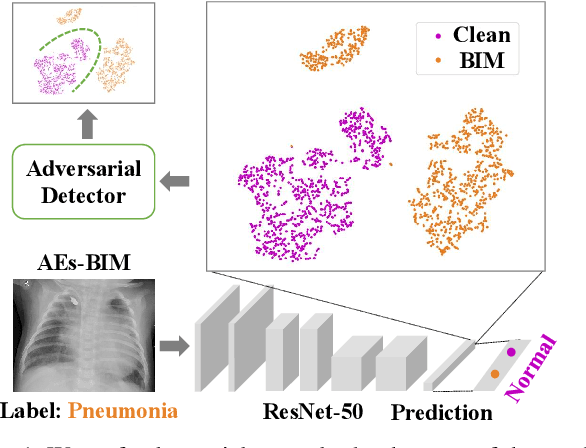
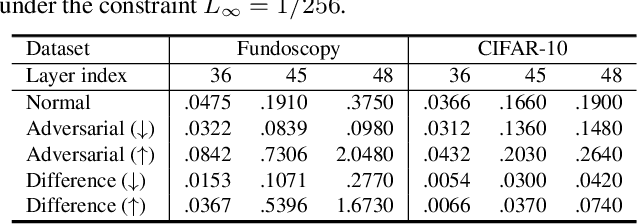
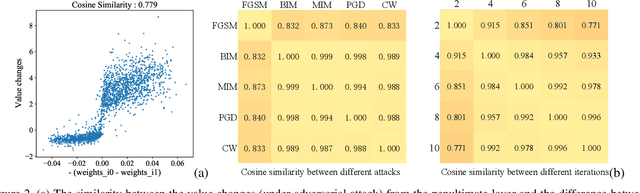
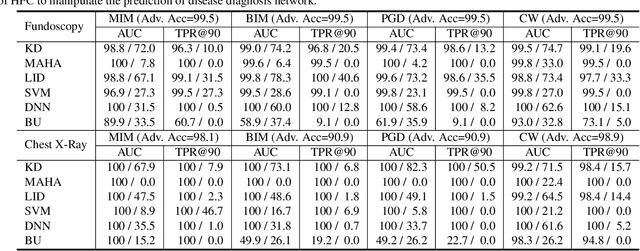
Abstract:Deep neural networks (DNNs) for medical images are extremely vulnerable to adversarial examples (AEs), which poses security concerns on clinical decision making. Luckily, medical AEs are also easy to detect in hierarchical feature space per our study herein. To better understand this phenomenon, we thoroughly investigate the intrinsic characteristic of medical AEs in feature space, providing both empirical evidence and theoretical explanations for the question: why are medical adversarial attacks easy to detect? We first perform a stress test to reveal the vulnerability of deep representations of medical images, in contrast to natural images. We then theoretically prove that typical adversarial attacks to binary disease diagnosis network manipulate the prediction by continuously optimizing the vulnerable representations in a fixed direction, resulting in outlier features that make medical AEs easy to detect. However, this vulnerability can also be exploited to hide the AEs in the feature space. We propose a novel hierarchical feature constraint (HFC) as an add-on to existing adversarial attacks, which encourages the hiding of the adversarial representation within the normal feature distribution. We evaluate the proposed method on two public medical image datasets, namely {Fundoscopy} and {Chest X-Ray}. Experimental results demonstrate the superiority of our adversarial attack method as it bypasses an array of state-of-the-art adversarial detectors more easily than competing attack methods, supporting that the great vulnerability of medical features allows an attacker more room to manipulate the adversarial representations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge