Jiawei Yang
Parallelized Spatiotemporal Binding
Feb 26, 2024Abstract:While modern best practices advocate for scalable architectures that support long-range interactions, object-centric models are yet to fully embrace these architectures. In particular, existing object-centric models for handling sequential inputs, due to their reliance on RNN-based implementation, show poor stability and capacity and are slow to train on long sequences. We introduce Parallelizable Spatiotemporal Binder or PSB, the first temporally-parallelizable slot learning architecture for sequential inputs. Unlike conventional RNN-based approaches, PSB produces object-centric representations, known as slots, for all time-steps in parallel. This is achieved by refining the initial slots across all time-steps through a fixed number of layers equipped with causal attention. By capitalizing on the parallelism induced by our architecture, the proposed model exhibits a significant boost in efficiency. In experiments, we test PSB extensively as an encoder within an auto-encoding framework paired with a wide variety of decoder options. Compared to the state-of-the-art, our architecture demonstrates stable training on longer sequences, achieves parallelization that results in a 60% increase in training speed, and yields performance that is on par with or better on unsupervised 2D and 3D object-centric scene decomposition and understanding.
Denoising Vision Transformers
Jan 05, 2024



Abstract:We delve into a nuanced but significant challenge inherent to Vision Transformers (ViTs): feature maps of these models exhibit grid-like artifacts, which detrimentally hurt the performance of ViTs in downstream tasks. Our investigations trace this fundamental issue down to the positional embeddings at the input stage. To address this, we propose a novel noise model, which is universally applicable to all ViTs. Specifically, the noise model dissects ViT outputs into three components: a semantics term free from noise artifacts and two artifact-related terms that are conditioned on pixel locations. Such a decomposition is achieved by enforcing cross-view feature consistency with neural fields in a per-image basis. This per-image optimization process extracts artifact-free features from raw ViT outputs, providing clean features for offline applications. Expanding the scope of our solution to support online functionality, we introduce a learnable denoiser to predict artifact-free features directly from unprocessed ViT outputs, which shows remarkable generalization capabilities to novel data without the need for per-image optimization. Our two-stage approach, termed Denoising Vision Transformers (DVT), does not require re-training existing pre-trained ViTs and is immediately applicable to any Transformer-based architecture. We evaluate our method on a variety of representative ViTs (DINO, MAE, DeiT-III, EVA02, CLIP, DINOv2, DINOv2-reg). Extensive evaluations demonstrate that our DVT consistently and significantly improves existing state-of-the-art general-purpose models in semantic and geometric tasks across multiple datasets (e.g., +3.84 mIoU). We hope our study will encourage a re-evaluation of ViT design, especially regarding the naive use of positional embeddings.
Rethinking Directional Integration in Neural Radiance Fields
Nov 28, 2023



Abstract:Recent works use the Neural radiance field (NeRF) to perform multi-view 3D reconstruction, providing a significant leap in rendering photorealistic scenes. However, despite its efficacy, NeRF exhibits limited capability of learning view-dependent effects compared to light field rendering or image-based view synthesis. To that end, we introduce a modification to the NeRF rendering equation which is as simple as a few lines of code change for any NeRF variations, while greatly improving the rendering quality of view-dependent effects. By swapping the integration operator and the direction decoder network, we only integrate the positional features along the ray and move the directional terms out of the integration, resulting in a disentanglement of the view-dependent and independent components. The modified equation is equivalent to the classical volumetric rendering in ideal cases on object surfaces with Dirac densities. Furthermore, we prove that with the errors caused by network approximation and numerical integration, our rendering equation exhibits better convergence properties with lower error accumulations compared to the classical NeRF. We also show that the modified equation can be interpreted as light field rendering with learned ray embeddings. Experiments on different NeRF variations show consistent improvements in the quality of view-dependent effects with our simple modification.
EmerNeRF: Emergent Spatial-Temporal Scene Decomposition via Self-Supervision
Nov 03, 2023Abstract:We present EmerNeRF, a simple yet powerful approach for learning spatial-temporal representations of dynamic driving scenes. Grounded in neural fields, EmerNeRF simultaneously captures scene geometry, appearance, motion, and semantics via self-bootstrapping. EmerNeRF hinges upon two core components: First, it stratifies scenes into static and dynamic fields. This decomposition emerges purely from self-supervision, enabling our model to learn from general, in-the-wild data sources. Second, EmerNeRF parameterizes an induced flow field from the dynamic field and uses this flow field to further aggregate multi-frame features, amplifying the rendering precision of dynamic objects. Coupling these three fields (static, dynamic, and flow) enables EmerNeRF to represent highly-dynamic scenes self-sufficiently, without relying on ground truth object annotations or pre-trained models for dynamic object segmentation or optical flow estimation. Our method achieves state-of-the-art performance in sensor simulation, significantly outperforming previous methods when reconstructing static (+2.93 PSNR) and dynamic (+3.70 PSNR) scenes. In addition, to bolster EmerNeRF's semantic generalization, we lift 2D visual foundation model features into 4D space-time and address a general positional bias in modern Transformers, significantly boosting 3D perception performance (e.g., 37.50% relative improvement in occupancy prediction accuracy on average). Finally, we construct a diverse and challenging 120-sequence dataset to benchmark neural fields under extreme and highly-dynamic settings.
CodeFuse-13B: A Pretrained Multi-lingual Code Large Language Model
Oct 10, 2023Abstract:Code Large Language Models (Code LLMs) have gained significant attention in the industry due to their wide applications in the full lifecycle of software engineering. However, the effectiveness of existing models in understanding non-English inputs for multi-lingual code-related tasks is still far from well studied. This paper introduces CodeFuse-13B, an open-sourced pre-trained code LLM. It is specifically designed for code-related tasks with both English and Chinese prompts and supports over 40 programming languages. CodeFuse achieves its effectiveness by utilizing a high quality pre-training dataset that is carefully filtered by program analyzers and optimized during the training process. Extensive experiments are conducted using real-world usage scenarios, the industry-standard benchmark HumanEval-x, and the specially designed CodeFuseEval for Chinese prompts. To assess the effectiveness of CodeFuse, we actively collected valuable human feedback from the AntGroup's software development process where CodeFuse has been successfully deployed. The results demonstrate that CodeFuse-13B achieves a HumanEval pass@1 score of 37.10%, positioning it as one of the top multi-lingual code LLMs with similar parameter sizes. In practical scenarios, such as code generation, code translation, code comments, and testcase generation, CodeFuse performs better than other models when confronted with Chinese prompts.
STNet: Spatial and Temporal feature fusion network for change detection in remote sensing images
Apr 22, 2023Abstract:As an important task in remote sensing image analysis, remote sensing change detection (RSCD) aims to identify changes of interest in a region from spatially co-registered multi-temporal remote sensing images, so as to monitor the local development. Existing RSCD methods usually formulate RSCD as a binary classification task, representing changes of interest by merely feature concatenation or feature subtraction and recovering the spatial details via densely connected change representations, whose performances need further improvement. In this paper, we propose STNet, a RSCD network based on spatial and temporal feature fusions. Specifically, we design a temporal feature fusion (TFF) module to combine bi-temporal features using a cross-temporal gating mechanism for emphasizing changes of interest; a spatial feature fusion module is deployed to capture fine-grained information using a cross-scale attention mechanism for recovering the spatial details of change representations. Experimental results on three benchmark datasets for RSCD demonstrate that the proposed method achieves the state-of-the-art performance. Code is available at https://github.com/xwmaxwma/rschange.
Improving Autoencoder-based Outlier Detection with Adjustable Probabilistic Reconstruction Error and Mean-shift Outlier Scoring
Apr 03, 2023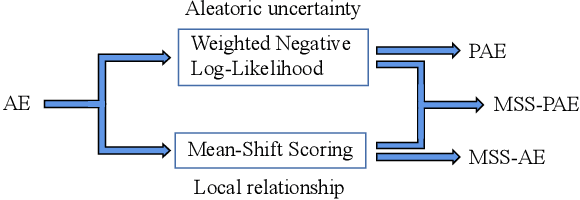
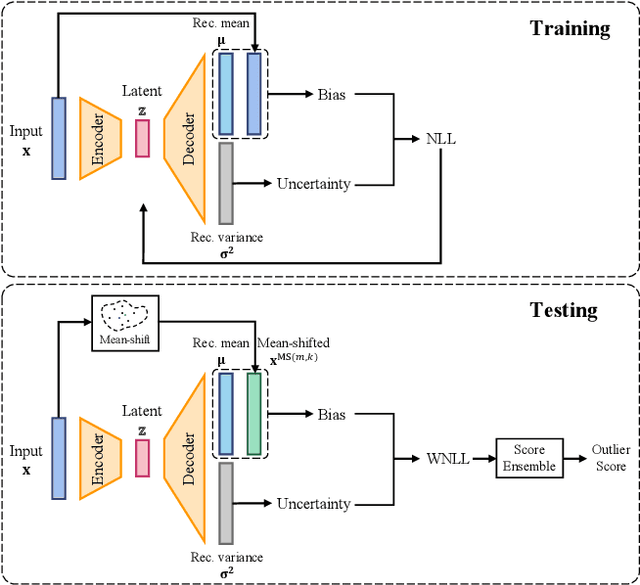
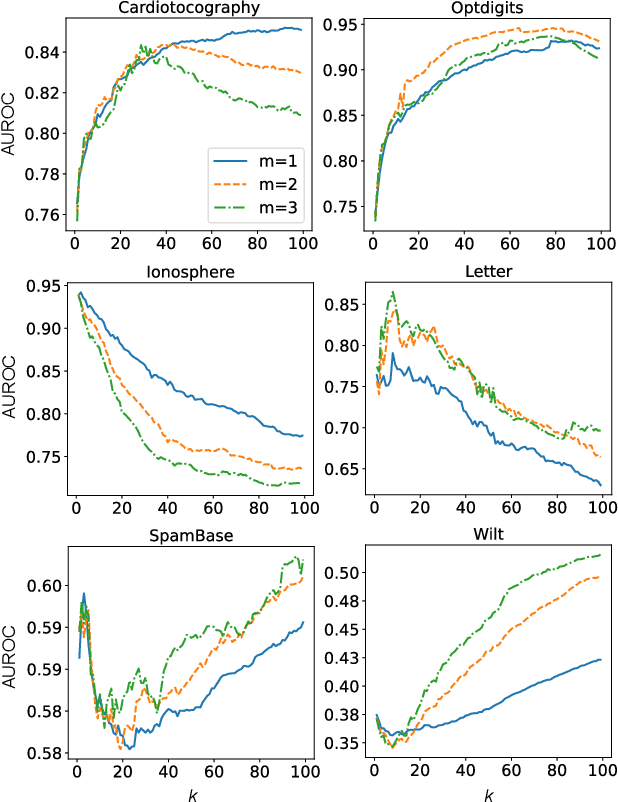
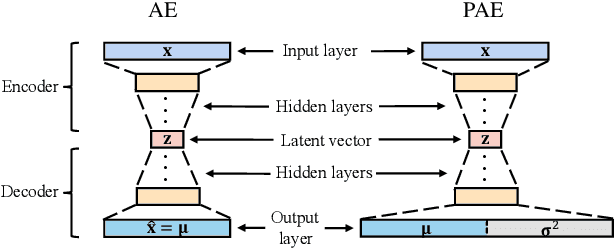
Abstract:Autoencoders were widely used in many machine learning tasks thanks to their strong learning ability which has drawn great interest among researchers in the field of outlier detection. However, conventional autoencoder-based methods lacked considerations in two aspects. This limited their performance in outlier detection. First, the mean squared error used in conventional autoencoders ignored the judgment uncertainty of the autoencoder, which limited their representation ability. Second, autoencoders suffered from the abnormal reconstruction problem: some outliers can be unexpectedly reconstructed well, making them difficult to identify from the inliers. To mitigate the aforementioned issues, two novel methods were proposed in this paper. First, a novel loss function named Probabilistic Reconstruction Error (PRE) was constructed to factor in both reconstruction bias and judgment uncertainty. To further control the trade-off of these two factors, two weights were introduced in PRE producing Adjustable Probabilistic Reconstruction Error (APRE), which benefited the outlier detection in different applications. Second, a conceptually new outlier scoring method based on mean-shift (MSS) was proposed to reduce the false inliers caused by the autoencoder. Experiments on 32 real-world outlier detection datasets proved the effectiveness of the proposed methods. The combination of the proposed methods achieved 41% of the relative performance improvement compared to the best baseline. The MSS improved the performance of multiple autoencoder-based outlier detectors by an average of 20%. The proposed two methods have the potential to advance autoencoder's development in outlier detection. The code is available on www.OutlierNet.com for reproducibility.
Oral-NeXF: 3D Oral Reconstruction with Neural X-ray Field from Panoramic Imaging
Mar 21, 2023Abstract:3D reconstruction of medical images from 2D images has increasingly become a challenging research topic with the advanced development of deep learning methods. Previous work in 3D reconstruction from limited (generally one or two) X-ray images mainly relies on learning from paired 2D and 3D images. In 3D oral reconstruction from panoramic imaging, the model also relies on some prior individual information, such as the dental arch curve or voxel-wise annotations, to restore the curved shape of the mandible during reconstruction. These limitations have hindered the use of single X-ray tomography in clinical applications. To address these challenges, we propose a new model that relies solely on projection data, including imaging direction and projection image, during panoramic scans to reconstruct the 3D oral structure. Our model builds on the neural radiance field by introducing multi-head prediction, dynamic sampling, and adaptive rendering, which accommodates the projection process of panoramic X-ray in dental imaging. Compared to end-to-end learning methods, our method achieves state-of-the-art performance without requiring additional supervision or prior knowledge.
Neighborhood Averaging for Improving Outlier Detectors
Mar 17, 2023



Abstract:We hypothesize that similar objects should have similar outlier scores. To our knowledge, all existing outlier detectors calculate the outlier score for each object independently regardless of the outlier scores of the other objects. Therefore, they do not guarantee that similar objects have similar outlier scores. To verify our proposed hypothesis, we propose an outlier score post-processing technique for outlier detectors, called neighborhood averaging(NA), which pays attention to objects and their neighbors and guarantees them to have more similar outlier scores than their original scores. Given an object and its outlier score from any outlier detector, NA modifies its outlier score by combining it with its k nearest neighbors' scores. We demonstrate the effectivity of NA by using the well-known k-nearest neighbors (k-NN). Experimental results show that NA improves all 10 tested baseline detectors by 13% (from 0.70 to 0.79 AUC) on average evaluated on nine real-world datasets. Moreover, even outlier detectors that are already based on k-NN are also improved. The experiments also show that in some applications, the choice of detector is no more significant when detectors are jointly used with NA, which may pose a challenge to the generally considered idea that the data model is the most important factor. We open our code on www.outlierNet.com for reproducibility.
FreeNeRF: Improving Few-shot Neural Rendering with Free Frequency Regularization
Mar 13, 2023Abstract:Novel view synthesis with sparse inputs is a challenging problem for neural radiance fields (NeRF). Recent efforts alleviate this challenge by introducing external supervision, such as pre-trained models and extra depth signals, and by non-trivial patch-based rendering. In this paper, we present Frequency regularized NeRF (FreeNeRF), a surprisingly simple baseline that outperforms previous methods with minimal modifications to the plain NeRF. We analyze the key challenges in few-shot neural rendering and find that frequency plays an important role in NeRF's training. Based on the analysis, we propose two regularization terms. One is to regularize the frequency range of NeRF's inputs, while the other is to penalize the near-camera density fields. Both techniques are ``free lunches'' at no additional computational cost. We demonstrate that even with one line of code change, the original NeRF can achieve similar performance as other complicated methods in the few-shot setting. FreeNeRF achieves state-of-the-art performance across diverse datasets, including Blender, DTU, and LLFF. We hope this simple baseline will motivate a rethinking of the fundamental role of frequency in NeRF's training under the low-data regime and beyond.
* Project page: https://jiawei-yang.github.io/FreeNeRF/, Code at: https://github.com/Jiawei-Yang/FreeNeRF
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge