Haoxin Zheng
Think Twice Before You Act: Improving Inverse Problem Solving With MCMC
Sep 13, 2024Abstract:Recent studies demonstrate that diffusion models can serve as a strong prior for solving inverse problems. A prominent example is Diffusion Posterior Sampling (DPS), which approximates the posterior distribution of data given the measure using Tweedie's formula. Despite the merits of being versatile in solving various inverse problems without re-training, the performance of DPS is hindered by the fact that this posterior approximation can be inaccurate especially for high noise levels. Therefore, we propose \textbf{D}iffusion \textbf{P}osterior \textbf{MC}MC (\textbf{DPMC}), a novel inference algorithm based on Annealed MCMC to solve inverse problems with pretrained diffusion models. We define a series of intermediate distributions inspired by the approximated conditional distributions used by DPS. Through annealed MCMC sampling, we encourage the samples to follow each intermediate distribution more closely before moving to the next distribution at a lower noise level, and therefore reduce the accumulated error along the path. We test our algorithm in various inverse problems, including super resolution, Gaussian deblurring, motion deblurring, inpainting, and phase retrieval. Our algorithm outperforms DPS with less number of evaluations across nearly all tasks, and is competitive among existing approaches.
Cross-Slice Attention and Evidential Critical Loss for Uncertainty-Aware Prostate Cancer Detection
Jul 01, 2024



Abstract:Current deep learning-based models typically analyze medical images in either 2D or 3D albeit disregarding volumetric information or suffering sub-optimal performance due to the anisotropic resolution of MR data. Furthermore, providing an accurate uncertainty estimation is beneficial to clinicians, as it indicates how confident a model is about its prediction. We propose a novel 2.5D cross-slice attention model that utilizes both global and local information, along with an evidential critical loss, to perform evidential deep learning for the detection in MR images of prostate cancer, one of the most common cancers and a leading cause of cancer-related death in men. We perform extensive experiments with our model on two different datasets and achieve state-of-the-art performance in prostate cancer detection along with improved epistemic uncertainty estimation. The implementation of the model is available at https://github.com/aL3x-O-o-Hung/GLCSA_ECLoss.
CSAM: A 2.5D Cross-Slice Attention Module for Anisotropic Volumetric Medical Image Segmentation
Nov 08, 2023



Abstract:A large portion of volumetric medical data, especially magnetic resonance imaging (MRI) data, is anisotropic, as the through-plane resolution is typically much lower than the in-plane resolution. Both 3D and purely 2D deep learning-based segmentation methods are deficient in dealing with such volumetric data since the performance of 3D methods suffers when confronting anisotropic data, and 2D methods disregard crucial volumetric information. Insufficient work has been done on 2.5D methods, in which 2D convolution is mainly used in concert with volumetric information. These models focus on learning the relationship across slices, but typically have many parameters to train. We offer a Cross-Slice Attention Module (CSAM) with minimal trainable parameters, which captures information across all the slices in the volume by applying semantic, positional, and slice attention on deep feature maps at different scales. Our extensive experiments using different network architectures and tasks demonstrate the usefulness and generalizability of CSAM. Associated code is available at https://github.com/aL3x-O-o-Hung/CSAM.
PartDiff: Image Super-resolution with Partial Diffusion Models
Jul 21, 2023



Abstract:Denoising diffusion probabilistic models (DDPMs) have achieved impressive performance on various image generation tasks, including image super-resolution. By learning to reverse the process of gradually diffusing the data distribution into Gaussian noise, DDPMs generate new data by iteratively denoising from random noise. Despite their impressive performance, diffusion-based generative models suffer from high computational costs due to the large number of denoising steps.In this paper, we first observed that the intermediate latent states gradually converge and become indistinguishable when diffusing a pair of low- and high-resolution images. This observation inspired us to propose the Partial Diffusion Model (PartDiff), which diffuses the image to an intermediate latent state instead of pure random noise, where the intermediate latent state is approximated by the latent of diffusing the low-resolution image. During generation, Partial Diffusion Models start denoising from the intermediate distribution and perform only a part of the denoising steps. Additionally, to mitigate the error caused by the approximation, we introduce "latent alignment", which aligns the latent between low- and high-resolution images during training. Experiments on both magnetic resonance imaging (MRI) and natural images show that, compared to plain diffusion-based super-resolution methods, Partial Diffusion Models significantly reduce the number of denoising steps without sacrificing the quality of generation.
Oral-NeXF: 3D Oral Reconstruction with Neural X-ray Field from Panoramic Imaging
Mar 21, 2023



Abstract:3D reconstruction of medical images from 2D images has increasingly become a challenging research topic with the advanced development of deep learning methods. Previous work in 3D reconstruction from limited (generally one or two) X-ray images mainly relies on learning from paired 2D and 3D images. In 3D oral reconstruction from panoramic imaging, the model also relies on some prior individual information, such as the dental arch curve or voxel-wise annotations, to restore the curved shape of the mandible during reconstruction. These limitations have hindered the use of single X-ray tomography in clinical applications. To address these challenges, we propose a new model that relies solely on projection data, including imaging direction and projection image, during panoramic scans to reconstruct the 3D oral structure. Our model builds on the neural radiance field by introducing multi-head prediction, dynamic sampling, and adaptive rendering, which accommodates the projection process of panoramic X-ray in dental imaging. Compared to end-to-end learning methods, our method achieves state-of-the-art performance without requiring additional supervision or prior knowledge.
CAT-Net: A Cross-Slice Attention Transformer Model for Prostate Zonal Segmentation in MRI
Mar 29, 2022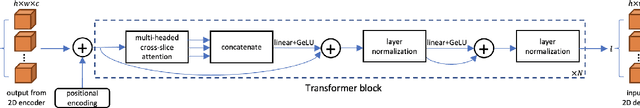
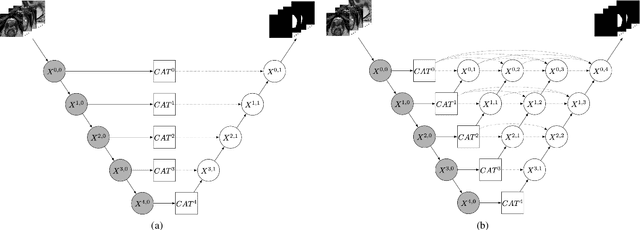
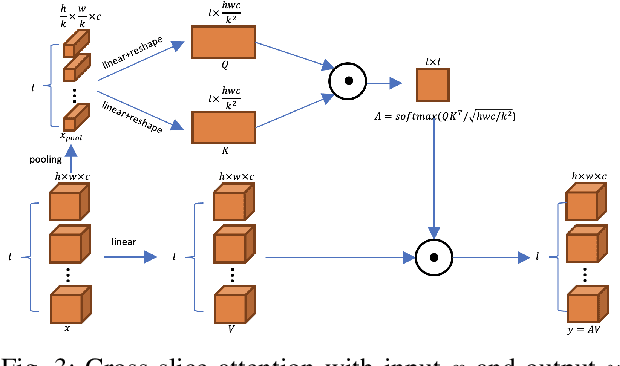
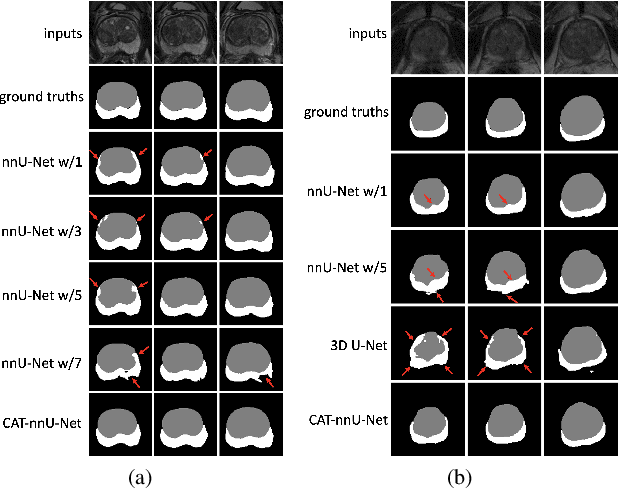
Abstract:Prostate cancer is the second leading cause of cancer death among men in the United States. The diagnosis of prostate MRI often relies on the accurate prostate zonal segmentation. However, state-of-the-art automatic segmentation methods often fail to produce well-contained volumetric segmentation of the prostate zones since certain slices of prostate MRI, such as base and apex slices, are harder to segment than other slices. This difficulty can be overcome by accounting for the cross-slice relationship of adjacent slices, but current methods do not fully learn and exploit such relationships. In this paper, we propose a novel cross-slice attention mechanism, which we use in a Transformer module to systematically learn the cross-slice relationship at different scales. The module can be utilized in any existing learning-based segmentation framework with skip connections. Experiments show that our cross-slice attention is able to capture the cross-slice information in prostate zonal segmentation and improve the performance of current state-of-the-art methods. Our method significantly improves segmentation accuracy in the peripheral zone, such that the segmentation results are consistent across all the prostate slices (apex, mid-gland, and base).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge