Jiahang Xu
Beyond Prompt Content: Enhancing LLM Performance via Content-Format Integrated Prompt Optimization
Feb 10, 2025Abstract:Large Language Models (LLMs) have shown significant capability across various tasks, with their real-world effectiveness often driven by prompt design. While recent research has focused on optimizing prompt content, the role of prompt formatting, a critical but often overlooked dimension, has received limited systematic investigation. In this paper, we introduce Content-Format Integrated Prompt Optimization (CFPO), an innovative methodology that jointly optimizes both prompt content and formatting through an iterative refinement process. CFPO leverages natural language mutations to explore content variations and employs a dynamic format exploration strategy that systematically evaluates diverse format options. Our extensive evaluations across multiple tasks and open-source LLMs demonstrate that CFPO demonstrates measurable performance improvements compared to content-only optimization methods. This highlights the importance of integrated content-format optimization and offers a practical, model-agnostic approach to enhancing LLM performance. Code is available at https://github.com/HenryLau7/CFPO.
Mutual Reasoning Makes Smaller LLMs Stronger Problem-Solvers
Aug 12, 2024Abstract:This paper introduces rStar, a self-play mutual reasoning approach that significantly improves reasoning capabilities of small language models (SLMs) without fine-tuning or superior models. rStar decouples reasoning into a self-play mutual generation-discrimination process. First, a target SLM augments the Monte Carlo Tree Search (MCTS) with a rich set of human-like reasoning actions to construct higher quality reasoning trajectories. Next, another SLM, with capabilities similar to the target SLM, acts as a discriminator to verify each trajectory generated by the target SLM. The mutually agreed reasoning trajectories are considered mutual consistent, thus are more likely to be correct. Extensive experiments across five SLMs demonstrate rStar can effectively solve diverse reasoning problems, including GSM8K, GSM-Hard, MATH, SVAMP, and StrategyQA. Remarkably, rStar boosts GSM8K accuracy from 12.51% to 63.91% for LLaMA2-7B, from 36.46% to 81.88% for Mistral-7B, from 74.53% to 91.13% for LLaMA3-8B-Instruct. Code will be available at https://github.com/zhentingqi/rStar.
VisEval: A Benchmark for Data Visualization in the Era of Large Language Models
Jul 01, 2024Abstract:Translating natural language to visualization (NL2VIS) has shown great promise for visual data analysis, but it remains a challenging task that requires multiple low-level implementations, such as natural language processing and visualization design. Recent advancements in pre-trained large language models (LLMs) are opening new avenues for generating visualizations from natural language. However, the lack of a comprehensive and reliable benchmark hinders our understanding of LLMs' capabilities in visualization generation. In this paper, we address this gap by proposing a new NL2VIS benchmark called VisEval. Firstly, we introduce a high-quality and large-scale dataset. This dataset includes 2,524 representative queries covering 146 databases, paired with accurately labeled ground truths. Secondly, we advocate for a comprehensive automated evaluation methodology covering multiple dimensions, including validity, legality, and readability. By systematically scanning for potential issues with a number of heterogeneous checkers, VisEval provides reliable and trustworthy evaluation outcomes. We run VisEval on a series of state-of-the-art LLMs. Our evaluation reveals prevalent challenges and delivers essential insights for future advancements.
Phi-3 Technical Report: A Highly Capable Language Model Locally on Your Phone
Apr 23, 2024



Abstract:We introduce phi-3-mini, a 3.8 billion parameter language model trained on 3.3 trillion tokens, whose overall performance, as measured by both academic benchmarks and internal testing, rivals that of models such as Mixtral 8x7B and GPT-3.5 (e.g., phi-3-mini achieves 69% on MMLU and 8.38 on MT-bench), despite being small enough to be deployed on a phone. The innovation lies entirely in our dataset for training, a scaled-up version of the one used for phi-2, composed of heavily filtered web data and synthetic data. The model is also further aligned for robustness, safety, and chat format. We also provide some initial parameter-scaling results with a 7B and 14B models trained for 4.8T tokens, called phi-3-small and phi-3-medium, both significantly more capable than phi-3-mini (e.g., respectively 75% and 78% on MMLU, and 8.7 and 8.9 on MT-bench).
LongRoPE: Extending LLM Context Window Beyond 2 Million Tokens
Feb 21, 2024



Abstract:Large context window is a desirable feature in large language models (LLMs). However, due to high fine-tuning costs, scarcity of long texts, and catastrophic values introduced by new token positions, current extended context windows are limited to around 128k tokens. This paper introduces LongRoPE that, for the first time, extends the context window of pre-trained LLMs to an impressive 2048k tokens, with up to only 1k fine-tuning steps at within 256k training lengths, while maintaining performance at the original short context window. This is achieved by three key innovations: (i) we identify and exploit two forms of non-uniformities in positional interpolation through an efficient search, providing a better initialization for fine-tuning and enabling an 8x extension in non-fine-tuning scenarios; (ii) we introduce a progressive extension strategy that first fine-tunes a 256k length LLM and then conducts a second positional interpolation on the fine-tuned extended LLM to achieve a 2048k context window; (iii) we readjust LongRoPE on 8k length to recover the short context window performance. Extensive experiments on LLaMA2 and Mistral across various tasks demonstrate the effectiveness of our method. Models extended via LongRoPE retain the original architecture with minor modifications to the positional embedding, and can reuse most pre-existing optimizations.
Compresso: Structured Pruning with Collaborative Prompting Learns Compact Large Language Models
Oct 11, 2023



Abstract:Despite the remarkable success of Large Language Models (LLMs), the massive size poses significant deployment challenges, particularly on resource-constrained hardware. While existing LLM compression methods focus on quantization, pruning remains relatively unexplored due to the high cost of training-based approaches and data collection challenges. One-shot pruning methods, although cost-effective and data-free, have become dominant in LLM pruning, but lead to performance decline under the structured pruning setting. In this work, we introduce a new paradigm for structurally pruning LLMs, called Compresso. Our approach, through the collaboration of the proposed resource-efficient pruning algorithm and the LLM itself, learns optimal pruning decisions during the training process. Compresso addresses the challenges of expensive training costs and data collection by incorporating Low-Rank Adaptation (LoRA) into the $L_0$ regularization during the instruction tuning process. Then, we further augment the pruning algorithm by introducing a collaborative prompt that fosters collaboration between the LLM and the pruning algorithm, significantly boosting the overall performance. To this end, Compresso prunes LLaMA-7B to 5.4B, maintaining original performance and even surpassing LLaMA-7B in reading comprehension by 2.62%. Extensive experiments demonstrate that Compresso significantly outperforms one-shot pruning baselines across various sparsity ratios, achieving up to 2.21%, 11.43%, 7.04%, and 4.81% higher scores on the commonsense reasoning, reading comprehension, MMLU, and BBH benchmarks, respectively.
Constraint-aware and Ranking-distilled Token Pruning for Efficient Transformer Inference
Jun 26, 2023Abstract:Deploying pre-trained transformer models like BERT on downstream tasks in resource-constrained scenarios is challenging due to their high inference cost, which grows rapidly with input sequence length. In this work, we propose a constraint-aware and ranking-distilled token pruning method ToP, which selectively removes unnecessary tokens as input sequence passes through layers, allowing the model to improve online inference speed while preserving accuracy. ToP overcomes the limitation of inaccurate token importance ranking in the conventional self-attention mechanism through a ranking-distilled token distillation technique, which distills effective token rankings from the final layer of unpruned models to early layers of pruned models. Then, ToP introduces a coarse-to-fine pruning approach that automatically selects the optimal subset of transformer layers and optimizes token pruning decisions within these layers through improved $L_0$ regularization. Extensive experiments on GLUE benchmark and SQuAD tasks demonstrate that ToP outperforms state-of-the-art token pruning and model compression methods with improved accuracy and speedups. ToP reduces the average FLOPs of BERT by 8.1x while achieving competitive accuracy on GLUE, and provides a real latency speedup of up to 7.4x on an Intel CPU.
ElasticViT: Conflict-aware Supernet Training for Deploying Fast Vision Transformer on Diverse Mobile Devices
Mar 21, 2023



Abstract:Neural Architecture Search (NAS) has shown promising performance in the automatic design of vision transformers (ViT) exceeding 1G FLOPs. However, designing lightweight and low-latency ViT models for diverse mobile devices remains a big challenge. In this work, we propose ElasticViT, a two-stage NAS approach that trains a high-quality ViT supernet over a very large search space that supports a wide range of mobile devices, and then searches an optimal sub-network (subnet) for direct deployment. However, prior supernet training methods that rely on uniform sampling suffer from the gradient conflict issue: the sampled subnets can have vastly different model sizes (e.g., 50M vs. 2G FLOPs), leading to different optimization directions and inferior performance. To address this challenge, we propose two novel sampling techniques: complexity-aware sampling and performance-aware sampling. Complexity-aware sampling limits the FLOPs difference among the subnets sampled across adjacent training steps, while covering different-sized subnets in the search space. Performance-aware sampling further selects subnets that have good accuracy, which can reduce gradient conflicts and improve supernet quality. Our discovered models, ElasticViT models, achieve top-1 accuracy from 67.2% to 80.0% on ImageNet from 60M to 800M FLOPs without extra retraining, outperforming all prior CNNs and ViTs in terms of accuracy and latency. Our tiny and small models are also the first ViT models that surpass state-of-the-art CNNs with significantly lower latency on mobile devices. For instance, ElasticViT-S1 runs 2.62x faster than EfficientNet-B0 with 0.1% higher accuracy.
SpaceEvo: Hardware-Friendly Search Space Design for Efficient INT8 Inference
Mar 15, 2023Abstract:The combination of Neural Architecture Search (NAS) and quantization has proven successful in automatically designing low-FLOPs INT8 quantized neural networks (QNN). However, directly applying NAS to design accurate QNN models that achieve low latency on real-world devices leads to inferior performance. In this work, we find that the poor INT8 latency is due to the quantization-unfriendly issue: the operator and configuration (e.g., channel width) choices in prior art search spaces lead to diverse quantization efficiency and can slow down the INT8 inference speed. To address this challenge, we propose SpaceEvo, an automatic method for designing a dedicated, quantization-friendly search space for each target hardware. The key idea of SpaceEvo is to automatically search hardware-preferred operators and configurations to construct the search space, guided by a metric called Q-T score to quantify how quantization-friendly a candidate search space is. We further train a quantized-for-all supernet over our discovered search space, enabling the searched models to be directly deployed without extra retraining or quantization. Our discovered models establish new SOTA INT8 quantized accuracy under various latency constraints, achieving up to 10.1% accuracy improvement on ImageNet than prior art CNNs under the same latency. Extensive experiments on diverse edge devices demonstrate that SpaceEvo consistently outperforms existing manually-designed search spaces with up to 2.5x faster speed while achieving the same accuracy.
Improving Hypernasality Estimation with Automatic Speech Recognition in Cleft Palate Speech
Aug 10, 2022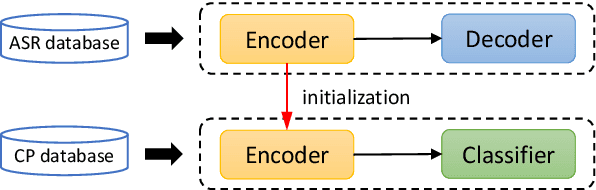
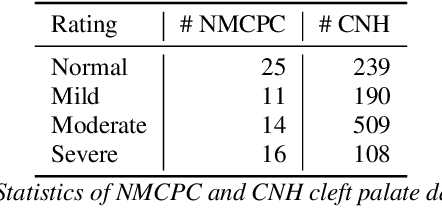
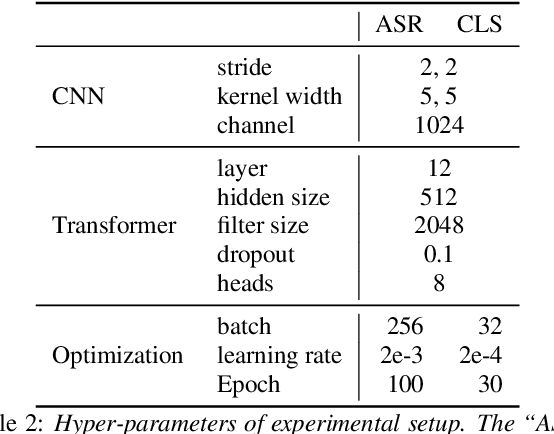
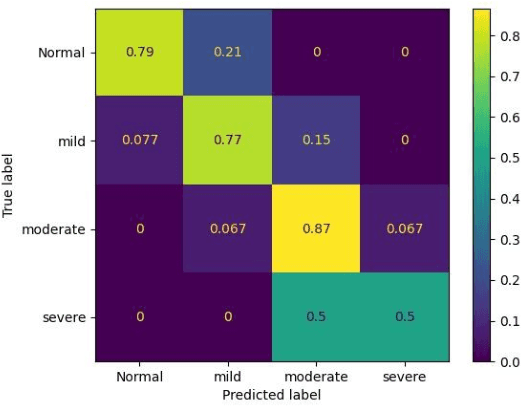
Abstract:Hypernasality is an abnormal resonance in human speech production, especially in patients with craniofacial anomalies such as cleft palate. In clinical application, hypernasality estimation is crucial in cleft palate diagnosis, as its results determine the subsequent surgery and additional speech therapy. Therefore, designing an automatic hypernasality assessment method will facilitate speech-language pathologists to make precise diagnoses. Existing methods for hypernasality estimation only conduct acoustic analysis based on low-resource cleft palate dataset, by using statistical or neural network-based features. In this paper, we propose a novel approach that uses automatic speech recognition model to improve hypernasality estimation. Specifically, we first pre-train an encoder-decoder framework in an automatic speech recognition (ASR) objective by using speech-to-text dataset, and then fine-tune ASR encoder on the cleft palate dataset for hypernasality estimation. Benefiting from such design, our model for hypernasality estimation can enjoy the advantages of ASR model: 1) compared with low-resource cleft palate dataset, the ASR task usually includes large-scale speech data in the general domain, which enables better model generalization; 2) the text annotations in ASR dataset guide model to extract better acoustic features. Experimental results on two cleft palate datasets demonstrate that our method achieves superior performance compared with previous approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge