Hanspeter Pfister
Robert Krueger and Jared Jessup contributed equally to this work
MST++: Multi-stage Spectral-wise Transformer for Efficient Spectral Reconstruction
Apr 17, 2022
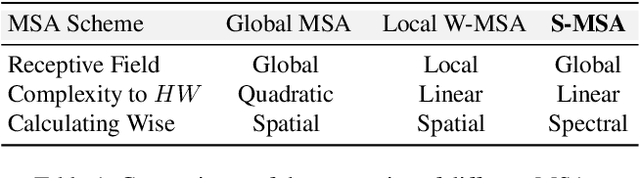
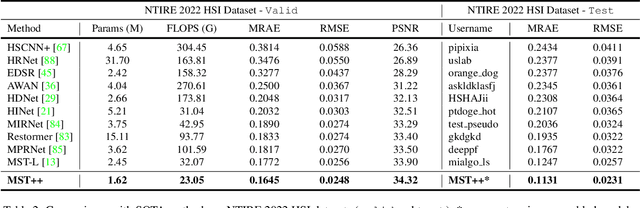
Abstract:Existing leading methods for spectral reconstruction (SR) focus on designing deeper or wider convolutional neural networks (CNNs) to learn the end-to-end mapping from the RGB image to its hyperspectral image (HSI). These CNN-based methods achieve impressive restoration performance while showing limitations in capturing the long-range dependencies and self-similarity prior. To cope with this problem, we propose a novel Transformer-based method, Multi-stage Spectral-wise Transformer (MST++), for efficient spectral reconstruction. In particular, we employ Spectral-wise Multi-head Self-attention (S-MSA) that is based on the HSI spatially sparse while spectrally self-similar nature to compose the basic unit, Spectral-wise Attention Block (SAB). Then SABs build up Single-stage Spectral-wise Transformer (SST) that exploits a U-shaped structure to extract multi-resolution contextual information. Finally, our MST++, cascaded by several SSTs, progressively improves the reconstruction quality from coarse to fine. Comprehensive experiments show that our MST++ significantly outperforms other state-of-the-art methods. In the NTIRE 2022 Spectral Reconstruction Challenge, our approach won the First place. Code and pre-trained models are publicly available at https://github.com/caiyuanhao1998/MST-plus-plus.
* Winner of NTIRE 2022 Challenge on Spectral Reconstruction from RGB; The First Transformer-based Method for Spectral Reconstruction
Instance Segmentation of Unlabeled Modalities via Cyclic Segmentation GAN
Apr 06, 2022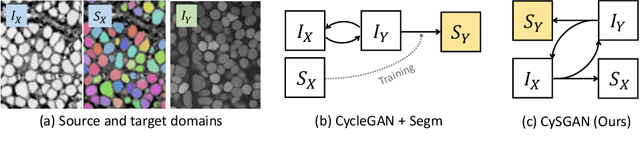
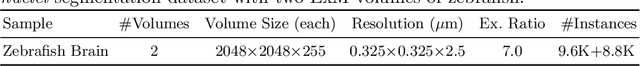
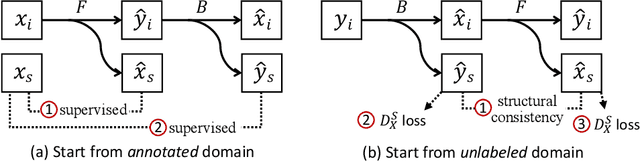
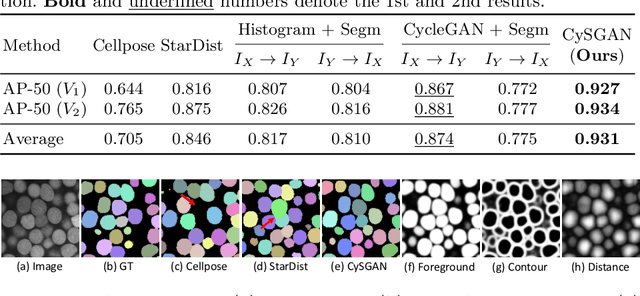
Abstract:Instance segmentation for unlabeled imaging modalities is a challenging but essential task as collecting expert annotation can be expensive and time-consuming. Existing works segment a new modality by either deploying a pre-trained model optimized on diverse training data or conducting domain translation and image segmentation as two independent steps. In this work, we propose a novel Cyclic Segmentation Generative Adversarial Network (CySGAN) that conducts image translation and instance segmentation jointly using a unified framework. Besides the CycleGAN losses for image translation and supervised losses for the annotated source domain, we introduce additional self-supervised and segmentation-based adversarial objectives to improve the model performance by leveraging unlabeled target domain images. We benchmark our approach on the task of 3D neuronal nuclei segmentation with annotated electron microscopy (EM) images and unlabeled expansion microscopy (ExM) data. Our CySGAN outperforms both pretrained generalist models and the baselines that sequentially conduct image translation and segmentation. Our implementation and the newly collected, densely annotated ExM nuclei dataset, named NucExM, are available at https://connectomics-bazaar.github.io/proj/CySGAN/index.html.
Learning to Generate Realistic Noisy Images via Pixel-level Noise-aware Adversarial Training
Apr 06, 2022
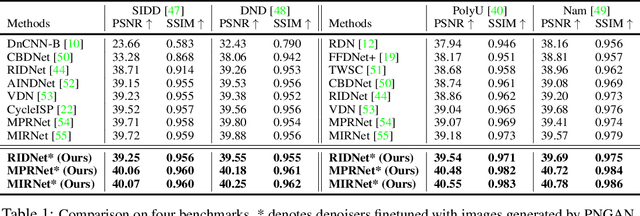
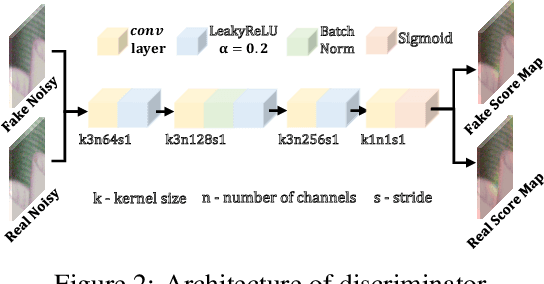
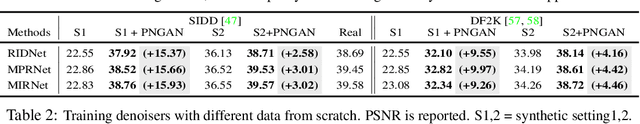
Abstract:Existing deep learning real denoising methods require a large amount of noisy-clean image pairs for supervision. Nonetheless, capturing a real noisy-clean dataset is an unacceptable expensive and cumbersome procedure. To alleviate this problem, this work investigates how to generate realistic noisy images. Firstly, we formulate a simple yet reasonable noise model that treats each real noisy pixel as a random variable. This model splits the noisy image generation problem into two sub-problems: image domain alignment and noise domain alignment. Subsequently, we propose a novel framework, namely Pixel-level Noise-aware Generative Adversarial Network (PNGAN). PNGAN employs a pre-trained real denoiser to map the fake and real noisy images into a nearly noise-free solution space to perform image domain alignment. Simultaneously, PNGAN establishes a pixel-level adversarial training to conduct noise domain alignment. Additionally, for better noise fitting, we present an efficient architecture Simple Multi-scale Network (SMNet) as the generator. Qualitative validation shows that noise generated by PNGAN is highly similar to real noise in terms of intensity and distribution. Quantitative experiments demonstrate that a series of denoisers trained with the generated noisy images achieve state-of-the-art (SOTA) results on four real denoising benchmarks.
Revisiting RCAN: Improved Training for Image Super-Resolution
Jan 27, 2022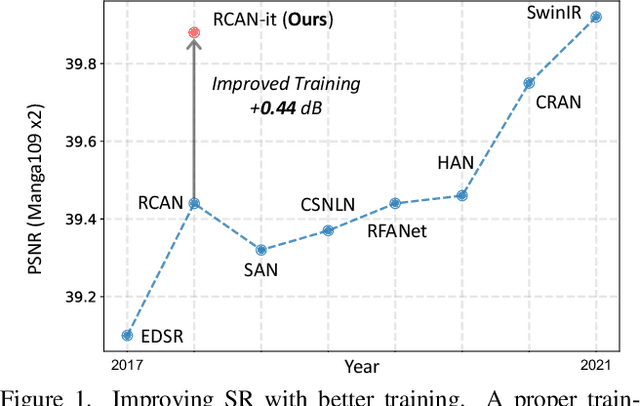
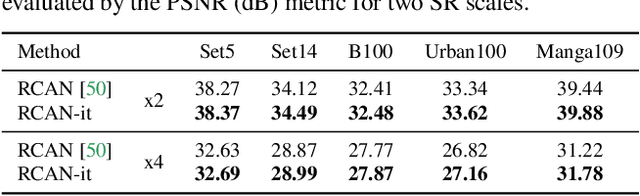
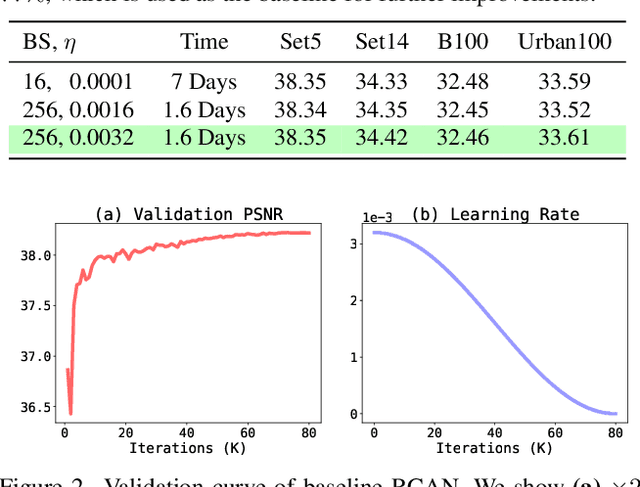
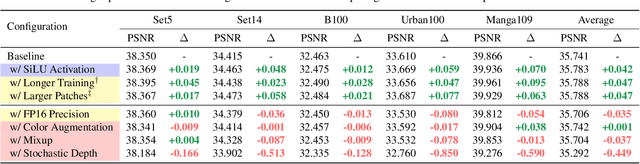
Abstract:Image super-resolution (SR) is a fast-moving field with novel architectures attracting the spotlight. However, most SR models were optimized with dated training strategies. In this work, we revisit the popular RCAN model and examine the effect of different training options in SR. Surprisingly (or perhaps as expected), we show that RCAN can outperform or match nearly all the CNN-based SR architectures published after RCAN on standard benchmarks with a proper training strategy and minimal architecture change. Besides, although RCAN is a very large SR architecture with more than four hundred convolutional layers, we draw a notable conclusion that underfitting is still the main problem restricting the model capability instead of overfitting. We observe supportive evidence that increasing training iterations clearly improves the model performance while applying regularization techniques generally degrades the predictions. We denote our simply revised RCAN as RCAN-it and recommend practitioners to use it as baselines for future research. Code is publicly available at https://github.com/zudi-lin/rcan-it.
PyTorch Connectomics: A Scalable and Flexible Segmentation Framework for EM Connectomics
Dec 10, 2021
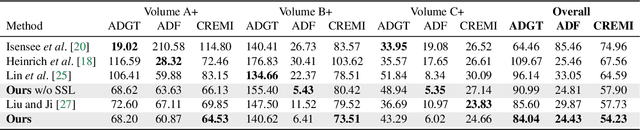
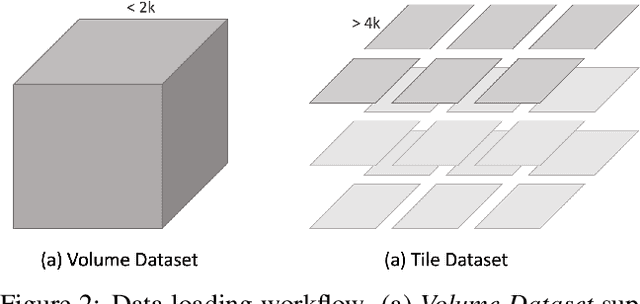
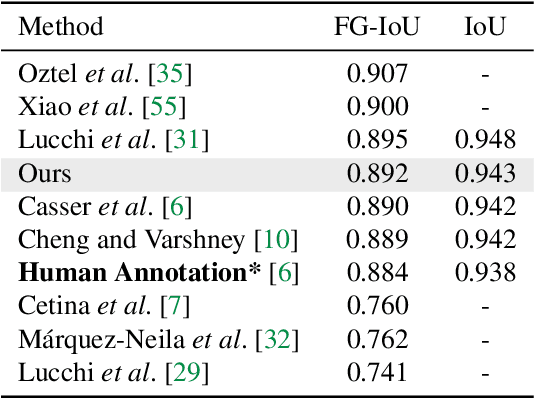
Abstract:We present PyTorch Connectomics (PyTC), an open-source deep-learning framework for the semantic and instance segmentation of volumetric microscopy images, built upon PyTorch. We demonstrate the effectiveness of PyTC in the field of connectomics, which aims to segment and reconstruct neurons, synapses, and other organelles like mitochondria at nanometer resolution for understanding neuronal communication, metabolism, and development in animal brains. PyTC is a scalable and flexible toolbox that tackles datasets at different scales and supports multi-task and semi-supervised learning to better exploit expensive expert annotations and the vast amount of unlabeled data during training. Those functionalities can be easily realized in PyTC by changing the configuration options without coding and adapted to other 2D and 3D segmentation tasks for different tissues and imaging modalities. Quantitatively, our framework achieves the best performance in the CREMI challenge for synaptic cleft segmentation (outperforms existing best result by relatively 6.1$\%$) and competitive performance on mitochondria and neuronal nuclei segmentation. Code and tutorials are publicly available at https://connectomics.readthedocs.io.
Object Propagation via Inter-Frame Attentions for Temporally Stable Video Instance Segmentation
Nov 15, 2021
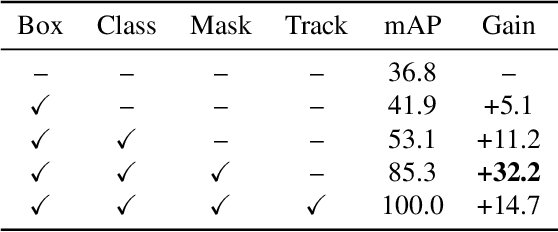
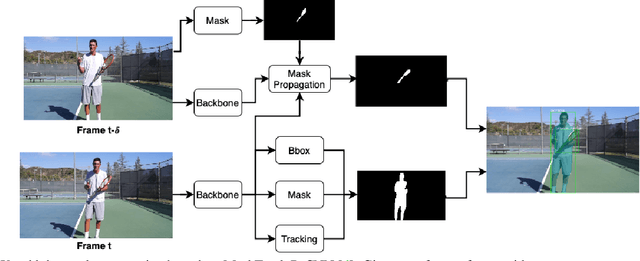
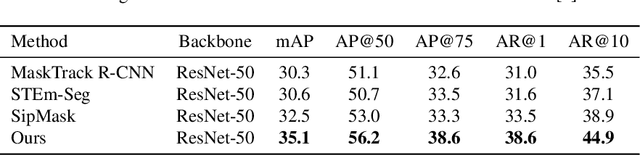
Abstract:Video instance segmentation aims to detect, segment, and track objects in a video. Current approaches extend image-level segmentation algorithms to the temporal domain. However, this results in temporally inconsistent masks. In this work, we identify the mask quality due to temporal stability as a performance bottleneck. Motivated by this, we propose a video instance segmentation method that alleviates the problem due to missing detections. Since this cannot be solved simply using spatial information, we leverage temporal context using inter-frame attentions. This allows our network to refocus on missing objects using box predictions from the neighbouring frame, thereby overcoming missing detections. Our method significantly outperforms previous state-of-the-art algorithms using the Mask R-CNN backbone, by achieving 35.1% mAP on the YouTube-VIS benchmark. Additionally, our method is completely online and requires no future frames. Our code is publicly available at https://github.com/anirudh-chakravarthy/ObjProp.
Three approaches to facilitate DNN generalization to objects in out-of-distribution orientations and illuminations: late-stopping, tuning batch normalization and invariance loss
Oct 30, 2021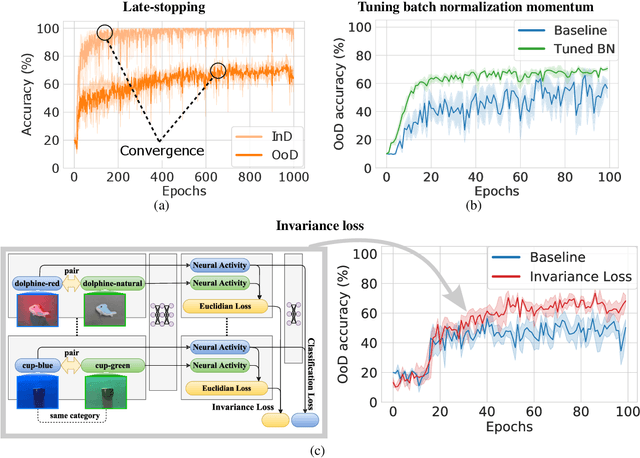
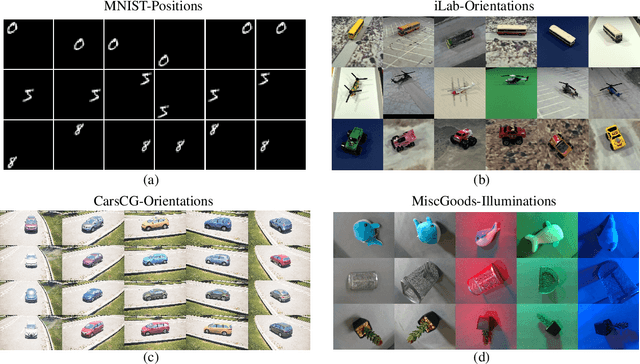
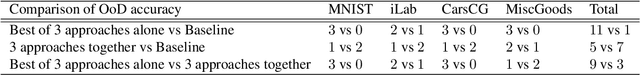

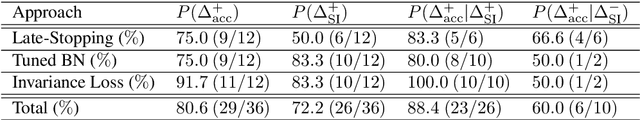
Abstract:The training data distribution is often biased towards objects in certain orientations and illumination conditions. While humans have a remarkable capability of recognizing objects in out-of-distribution (OoD) orientations and illuminations, Deep Neural Networks (DNNs) severely suffer in this case, even when large amounts of training examples are available. In this paper, we investigate three different approaches to improve DNNs in recognizing objects in OoD orientations and illuminations. Namely, these are (i) training much longer after convergence of the in-distribution (InD) validation accuracy, i.e., late-stopping, (ii) tuning the momentum parameter of the batch normalization layers, and (iii) enforcing invariance of the neural activity in an intermediate layer to orientation and illumination conditions. Each of these approaches substantially improves the DNN's OoD accuracy (more than 20% in some cases). We report results in four datasets: two datasets are modified from the MNIST and iLab datasets, and the other two are novel (one of 3D rendered cars and another of objects taken from various controlled orientations and illumination conditions). These datasets allow to study the effects of different amounts of bias and are challenging as DNNs perform poorly in OoD conditions. Finally, we demonstrate that even though the three approaches focus on different aspects of DNNs, they all tend to lead to the same underlying neural mechanism to enable OoD accuracy gains -- individual neurons in the intermediate layers become more selective to a category and also invariant to OoD orientations and illuminations.
MedMNIST v2: A Large-Scale Lightweight Benchmark for 2D and 3D Biomedical Image Classification
Oct 27, 2021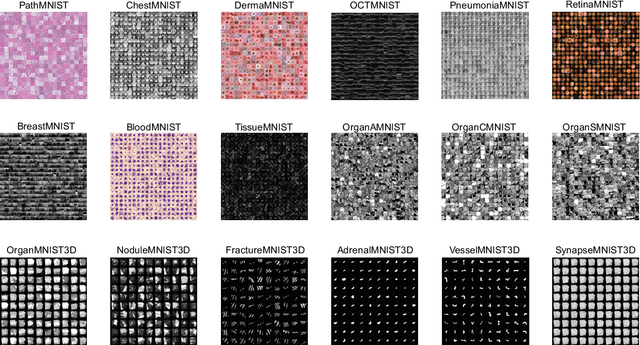
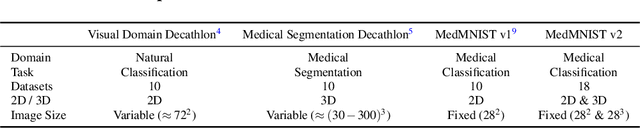

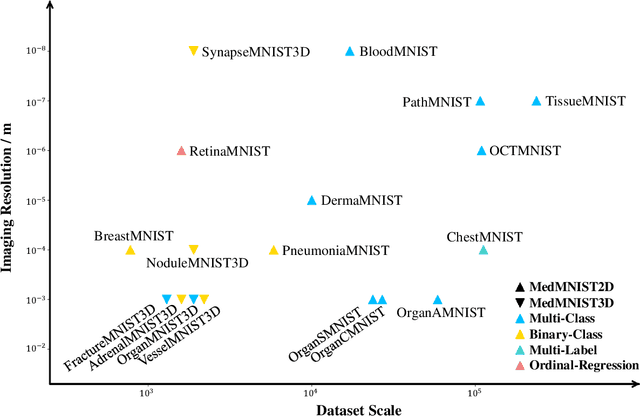
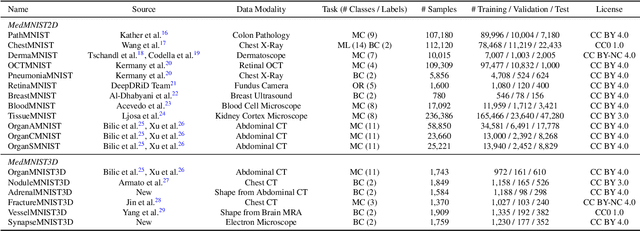
Abstract:We introduce MedMNIST v2, a large-scale MNIST-like dataset collection of standardized biomedical images, including 12 datasets for 2D and 6 datasets for 3D. All images are pre-processed into a small size of 28x28 (2D) or 28x28x28 (3D) with the corresponding classification labels so that no background knowledge is required for users. Covering primary data modalities in biomedical images, MedMNIST v2 is designed to perform classification on lightweight 2D and 3D images with various dataset scales (from 100 to 100,000) and diverse tasks (binary/multi-class, ordinal regression, and multi-label). The resulting dataset, consisting of 708,069 2D images and 10,214 3D images in total, could support numerous research / educational purposes in biomedical image analysis, computer vision, and machine learning. We benchmark several baseline methods on MedMNIST v2, including 2D / 3D neural networks and open-source / commercial AutoML tools. The data and code are publicly available at https://medmnist.com/.
GenNI: Human-AI Collaboration for Data-Backed Text Generation
Oct 19, 2021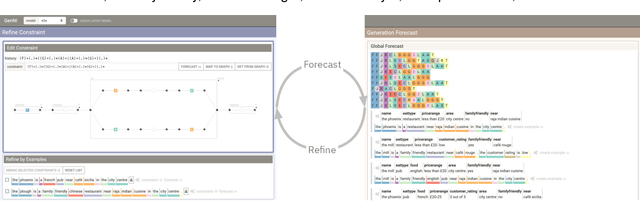
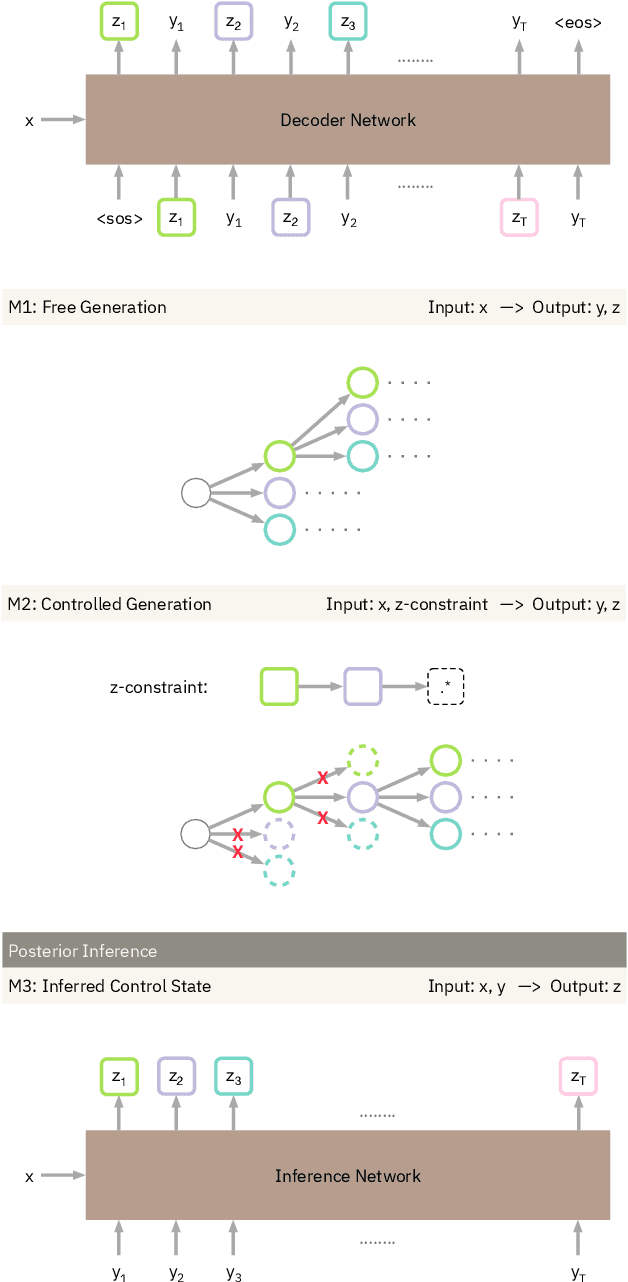
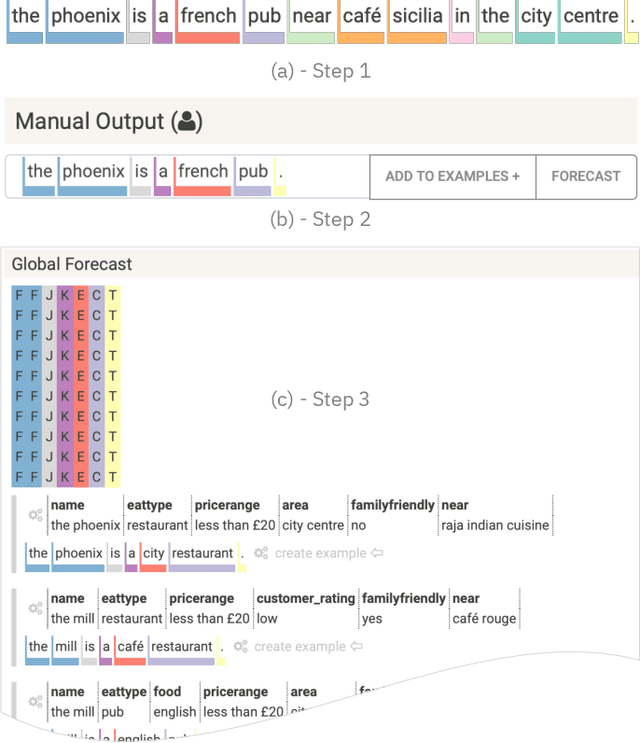
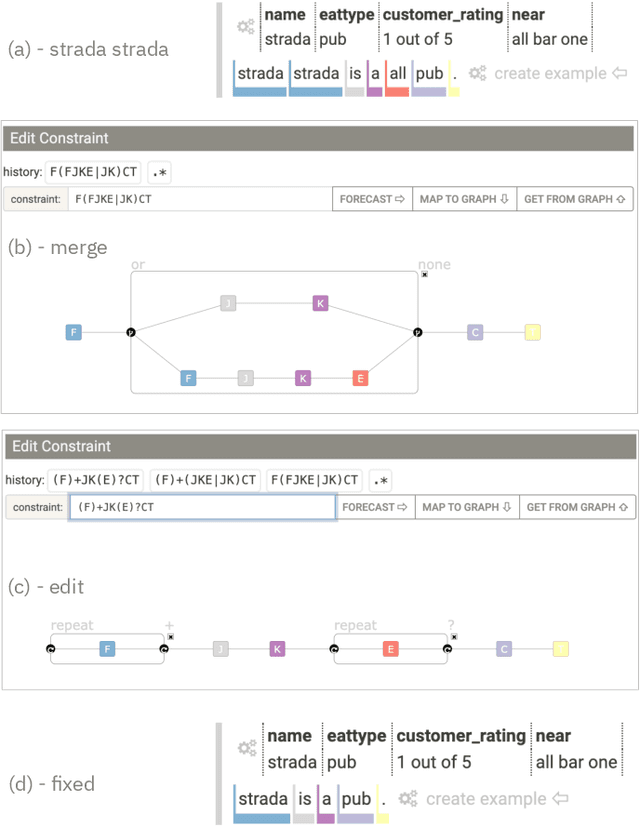
Abstract:Table2Text systems generate textual output based on structured data utilizing machine learning. These systems are essential for fluent natural language interfaces in tools such as virtual assistants; however, left to generate freely these ML systems often produce misleading or unexpected outputs. GenNI (Generation Negotiation Interface) is an interactive visual system for high-level human-AI collaboration in producing descriptive text. The tool utilizes a deep learning model designed with explicit control states. These controls allow users to globally constrain model generations, without sacrificing the representation power of the deep learning models. The visual interface makes it possible for users to interact with AI systems following a Refine-Forecast paradigm to ensure that the generation system acts in a manner human users find suitable. We report multiple use cases on two experiments that improve over uncontrolled generation approaches, while at the same time providing fine-grained control. A demo and source code are available at https://genni.vizhub.ai .
Scope2Screen: Focus+Context Techniques for Pathology Tumor Assessment in Multivariate Image Data
Oct 10, 2021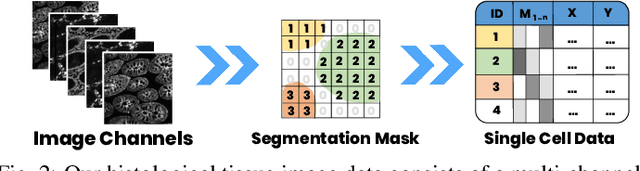

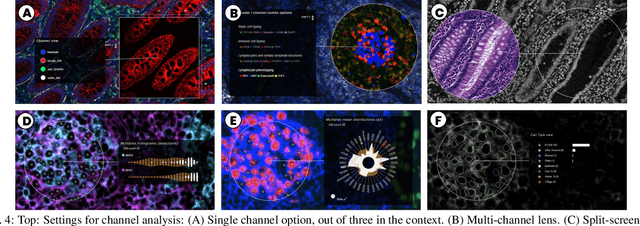
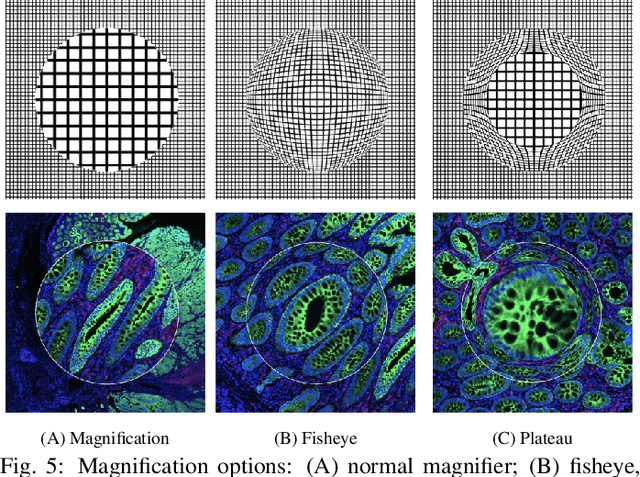
Abstract:Inspection of tissues using a light microscope is the primary method of diagnosing many diseases, notably cancer. Highly multiplexed tissue imaging builds on this foundation, enabling the collection of up to 60 channels of molecular information plus cell and tissue morphology using antibody staining. This provides unique insight into disease biology and promises to help with the design of patient-specific therapies. However, a substantial gap remains with respect to visualizing the resulting multivariate image data and effectively supporting pathology workflows in digital environments on screen. We, therefore, developed Scope2Screen, a scalable software system for focus+context exploration and annotation of whole-slide, high-plex, tissue images. Our approach scales to analyzing 100GB images of 10^9 or more pixels per channel, containing millions of cells. A multidisciplinary team of visualization experts, microscopists, and pathologists identified key image exploration and annotation tasks involving finding, magnifying, quantifying, and organizing ROIs in an intuitive and cohesive manner. Building on a scope2screen metaphor, we present interactive lensing techniques that operate at single-cell and tissue levels. Lenses are equipped with task-specific functionality and descriptive statistics, making it possible to analyze image features, cell types, and spatial arrangements (neighborhoods) across image channels and scales. A fast sliding-window search guides users to regions similar to those under the lens; these regions can be analyzed and considered either separately or as part of a larger image collection. A novel snapshot method enables linked lens configurations and image statistics to be saved, restored, and shared. We validate our designs with domain experts and apply Scope2Screen in two case studies involving lung and colorectal cancers to discover cancer-relevant image features.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge