Gangyong Jia
Spontaneous Spatial Cognition Emerges during Egocentric Video Viewing through Non-invasive BCI
Jul 16, 2025



Abstract:Humans possess a remarkable capacity for spatial cognition, allowing for self-localization even in novel or unfamiliar environments. While hippocampal neurons encoding position and orientation are well documented, the large-scale neural dynamics supporting spatial representation, particularly during naturalistic, passive experience, remain poorly understood. Here, we demonstrate for the first time that non-invasive brain-computer interfaces (BCIs) based on electroencephalography (EEG) can decode spontaneous, fine-grained egocentric 6D pose, comprising three-dimensional position and orientation, during passive viewing of egocentric video. Despite EEG's limited spatial resolution and high signal noise, we find that spatially coherent visual input (i.e., continuous and structured motion) reliably evokes decodable spatial representations, aligning with participants' subjective sense of spatial engagement. Decoding performance further improves when visual input is presented at a frame rate of 100 ms per image, suggesting alignment with intrinsic neural temporal dynamics. Using gradient-based backpropagation through a neural decoding model, we identify distinct EEG channels contributing to position -- and orientation specific -- components, revealing a distributed yet complementary neural encoding scheme. These findings indicate that the brain's spatial systems operate spontaneously and continuously, even under passive conditions, challenging traditional distinctions between active and passive spatial cognition. Our results offer a non-invasive window into the automatic construction of egocentric spatial maps and advance our understanding of how the human mind transforms everyday sensory experience into structured internal representations.
Spatial Temporal Attention based Target Vehicle Trajectory Prediction for Internet of Vehicles
Jan 01, 2025



Abstract:Forecasting vehicle behavior within complex traffic environments is pivotal within Intelligent Transportation Systems (ITS). Though this technology plays a significant role in alleviating the prevalent operational difficulties in logistics and transportation systems, the precise prediction of vehicle trajectories still poses a substantial challenge. To address this, our study introduces the Spatio Temporal Attention-based methodology for Target Vehicle Trajectory Prediction (STATVTPred). This approach integrates Global Positioning System(GPS) localization technology to track target movement and dynamically predict the vehicle's future path using comprehensive spatio-temporal trajectory data. We map the vehicle trajectory onto a directed graph, after which spatial attributes are extracted via a Graph Attention Networks(GATs). The Transformer technology is employed to yield temporal features from the sequence. These elements are then amalgamated with local road network structure maps to filter and deliver a smooth trajectory sequence, resulting in precise vehicle trajectory prediction.This study validates our proposed STATVTPred method on T-Drive and Chengdu taxi-trajectory datasets. The experimental results demonstrate that STATVTPred achieves 6.38% and 10.55% higher Average Match Rate (AMR) than the Transformer model on the Beijing and Chengdu datasets, respectively. Compared to the LSTM Encoder-Decoder model, STATVTPred boosts AMR by 37.45% and 36.06% on the same datasets. This is expected to establish STATVTPred as a new approach for handling trajectory prediction of targets in logistics and transportation scenarios, thereby enhancing prediction accuracy.
ICH-SCNet: Intracerebral Hemorrhage Segmentation and Prognosis Classification Network Using CLIP-guided SAM mechanism
Nov 07, 2024Abstract:Intracerebral hemorrhage (ICH) is the most fatal subtype of stroke and is characterized by a high incidence of disability. Accurate segmentation of the ICH region and prognosis prediction are critically important for developing and refining treatment plans for post-ICH patients. However, existing approaches address these two tasks independently and predominantly focus on imaging data alone, thereby neglecting the intrinsic correlation between the tasks and modalities. This paper introduces a multi-task network, ICH-SCNet, designed for both ICH segmentation and prognosis classification. Specifically, we integrate a SAM-CLIP cross-modal interaction mechanism that combines medical text and segmentation auxiliary information with neuroimaging data to enhance cross-modal feature recognition. Additionally, we develop an effective feature fusion module and a multi-task loss function to improve performance further. Extensive experiments on an ICH dataset reveal that our approach surpasses other state-of-the-art methods. It excels in the overall performance of classification tasks and outperforms competing models in all segmentation task metrics.
LPUWF-LDM: Enhanced Latent Diffusion Model for Precise Late-phase UWF-FA Generation on Limited Dataset
Sep 01, 2024



Abstract:Ultra-Wide-Field Fluorescein Angiography (UWF-FA) enables precise identification of ocular diseases using sodium fluorescein, which can be potentially harmful. Existing research has developed methods to generate UWF-FA from Ultra-Wide-Field Scanning Laser Ophthalmoscopy (UWF-SLO) to reduce the adverse reactions associated with injections. However, these methods have been less effective in producing high-quality late-phase UWF-FA, particularly in lesion areas and fine details. Two primary challenges hinder the generation of high-quality late-phase UWF-FA: the scarcity of paired UWF-SLO and early/late-phase UWF-FA datasets, and the need for realistic generation at lesion sites and potential blood leakage regions. This study introduces an improved latent diffusion model framework to generate high-quality late-phase UWF-FA from limited paired UWF images. To address the challenges as mentioned earlier, our approach employs a module utilizing Cross-temporal Regional Difference Loss, which encourages the model to focus on the differences between early and late phases. Additionally, we introduce a low-frequency enhanced noise strategy in the diffusion forward process to improve the realism of medical images. To further enhance the mapping capability of the variational autoencoder module, especially with limited datasets, we implement a Gated Convolutional Encoder to extract additional information from conditional images. Our Latent Diffusion Model for Ultra-Wide-Field Late-Phase Fluorescein Angiography (LPUWF-LDM) effectively reconstructs fine details in late-phase UWF-FA and achieves state-of-the-art results compared to other existing methods when working with limited datasets. Our source code is available at: https://github.com/Tinysqua/****.
Class-balanced Open-set Semi-supervised Object Detection for Medical Images
Aug 22, 2024



Abstract:Medical image datasets in the real world are often unlabeled and imbalanced, and Semi-Supervised Object Detection (SSOD) can utilize unlabeled data to improve an object detector. However, existing approaches predominantly assumed that the unlabeled data and test data do not contain out-of-distribution (OOD) classes. The few open-set semi-supervised object detection methods have two weaknesses: first, the class imbalance is not considered; second, the OOD instances are distinguished and simply discarded during pseudo-labeling. In this paper, we consider the open-set semi-supervised object detection problem which leverages unlabeled data that contain OOD classes to improve object detection for medical images. Our study incorporates two key innovations: Category Control Embed (CCE) and out-of-distribution Detection Fusion Classifier (OODFC). CCE is designed to tackle dataset imbalance by constructing a Foreground information Library, while OODFC tackles open-set challenges by integrating the ``unknown'' information into basic pseudo-labels. Our method outperforms the state-of-the-art SSOD performance, achieving a 4.25 mAP improvement on the public Parasite dataset.
ICHPro: Intracerebral Hemorrhage Prognosis Classification Via Joint-attention Fusion-based 3d Cross-modal Network
Feb 17, 2024



Abstract:Intracerebral Hemorrhage (ICH) is the deadliest subtype of stroke, necessitating timely and accurate prognostic evaluation to reduce mortality and disability. However, the multi-factorial nature and complexity of ICH make methods based solely on computed tomography (CT) image features inadequate. Despite the capacity of cross-modal networks to fuse additional information, the effective combination of different modal features remains a significant challenge. In this study, we propose a joint-attention fusion-based 3D cross-modal network termed ICHPro that simulates the ICH prognosis interpretation process utilized by neurosurgeons. ICHPro includes a joint-attention fusion module to fuse features from CT images with demographic and clinical textual data. To enhance the representation of cross-modal features, we introduce a joint loss function. ICHPro facilitates the extraction of richer cross-modal features, thereby improving classification performance. Upon testing our method using a five-fold cross-validation, we achieved an accuracy of 89.11%, an F1 score of 0.8767, and an AUC value of 0.9429. These results outperform those obtained from other advanced methods based on the test dataset, thereby demonstrating the superior efficacy of ICHPro. The code is available at our Github: https://github.com/YU-deep/ICH.
TTMFN: Two-stream Transformer-based Multimodal Fusion Network for Survival Prediction
Nov 13, 2023



Abstract:Survival prediction plays a crucial role in assisting clinicians with the development of cancer treatment protocols. Recent evidence shows that multimodal data can help in the diagnosis of cancer disease and improve survival prediction. Currently, deep learning-based approaches have experienced increasing success in survival prediction by integrating pathological images and gene expression data. However, most existing approaches overlook the intra-modality latent information and the complex inter-modality correlations. Furthermore, existing modalities do not fully exploit the immense representational capabilities of neural networks for feature aggregation and disregard the importance of relationships between features. Therefore, it is highly recommended to address these issues in order to enhance the prediction performance by proposing a novel deep learning-based method. We propose a novel framework named Two-stream Transformer-based Multimodal Fusion Network for survival prediction (TTMFN), which integrates pathological images and gene expression data. In TTMFN, we present a two-stream multimodal co-attention transformer module to take full advantage of the complex relationships between different modalities and the potential connections within the modalities. Additionally, we develop a multi-head attention pooling approach to effectively aggregate the feature representations of the two modalities. The experiment results on four datasets from The Cancer Genome Atlas demonstrate that TTMFN can achieve the best performance or competitive results compared to the state-of-the-art methods in predicting the overall survival of patients.
GCS-ICHNet: Assessment of Intracerebral Hemorrhage Prognosis using Self-Attention with Domain Knowledge Integration
Nov 08, 2023



Abstract:Intracerebral Hemorrhage (ICH) is a severe condition resulting from damaged brain blood vessel ruptures, often leading to complications and fatalities. Timely and accurate prognosis and management are essential due to its high mortality rate. However, conventional methods heavily rely on subjective clinician expertise, which can lead to inaccurate diagnoses and delays in treatment. Artificial intelligence (AI) models have been explored to assist clinicians, but many prior studies focused on model modification without considering domain knowledge. This paper introduces a novel deep learning algorithm, GCS-ICHNet, which integrates multimodal brain CT image data and the Glasgow Coma Scale (GCS) score to improve ICH prognosis. The algorithm utilizes a transformer-based fusion module for assessment. GCS-ICHNet demonstrates high sensitivity 81.03% and specificity 91.59%, outperforming average clinicians and other state-of-the-art methods.
UWAT-GAN: Fundus Fluorescein Angiography Synthesis via Ultra-wide-angle Transformation Multi-scale GAN
Jul 21, 2023



Abstract:Fundus photography is an essential examination for clinical and differential diagnosis of fundus diseases. Recently, Ultra-Wide-angle Fundus (UWF) techniques, UWF Fluorescein Angiography (UWF-FA) and UWF Scanning Laser Ophthalmoscopy (UWF-SLO) have been gradually put into use. However, Fluorescein Angiography (FA) and UWF-FA require injecting sodium fluorescein which may have detrimental influences. To avoid negative impacts, cross-modality medical image generation algorithms have been proposed. Nevertheless, current methods in fundus imaging could not produce high-resolution images and are unable to capture tiny vascular lesion areas. This paper proposes a novel conditional generative adversarial network (UWAT-GAN) to synthesize UWF-FA from UWF-SLO. Using multi-scale generators and a fusion module patch to better extract global and local information, our model can generate high-resolution images. Moreover, an attention transmit module is proposed to help the decoder learn effectively. Besides, a supervised approach is used to train the network using multiple new weighted losses on different scales of data. Experiments on an in-house UWF image dataset demonstrate the superiority of the UWAT-GAN over the state-of-the-art methods. The source code is available at: https://github.com/Tinysqua/UWAT-GAN.
DU-Net based Unsupervised Contrastive Learning for Cancer Segmentation in Histology Images
Jun 17, 2022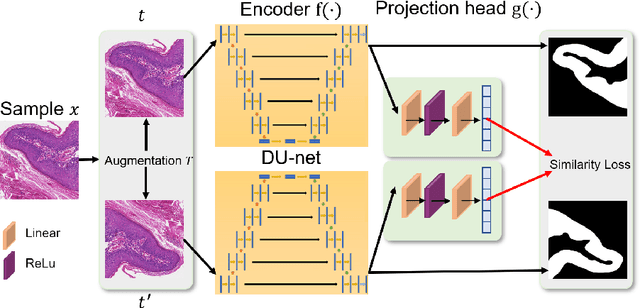
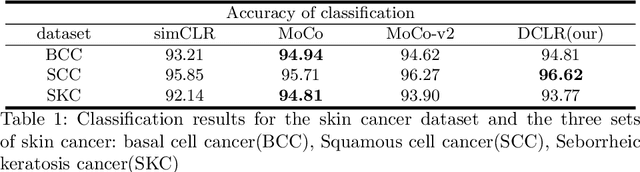
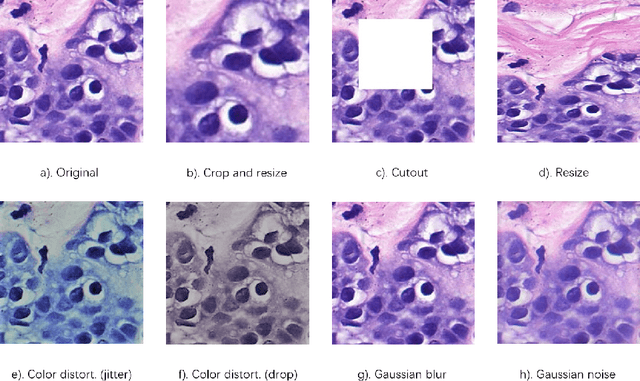
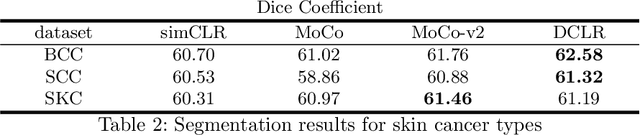
Abstract:In this paper, we introduce an unsupervised cancer segmentation framework for histology images. The framework involves an effective contrastive learning scheme for extracting distinctive visual representations for segmentation. The encoder is a Deep U-Net (DU-Net) structure that contains an extra fully convolution layer compared to the normal U-Net. A contrastive learning scheme is developed to solve the problem of lacking training sets with high-quality annotations on tumour boundaries. A specific set of data augmentation techniques are employed to improve the discriminability of the learned colour features from contrastive learning. Smoothing and noise elimination are conducted using convolutional Conditional Random Fields. The experiments demonstrate competitive performance in segmentation even better than some popular supervised networks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge