Fengyi Li
MTFM: A Scalable and Alignment-free Foundation Model for Industrial Recommendation in Meituan
Feb 13, 2026Abstract:Industrial recommendation systems typically involve multiple scenarios, yet existing cross-domain (CDR) and multi-scenario (MSR) methods often require prohibitive resources and strict input alignment, limiting their extensibility. We propose MTFM (Meituan Foundation Model for Recommendation), a transformer-based framework that addresses these challenges. Instead of pre-aligning inputs, MTFM transforms cross-domain data into heterogeneous tokens, capturing multi-scenario knowledge in an alignment-free manner. To enhance efficiency, we first introduce a multi-scenario user-level sample aggregation that significantly enhances training throughput by reducing the total number of instances. We further integrate Grouped-Query Attention and a customized Hybrid Target Attention to minimize memory usage and computational complexity. Furthermore, we implement various system-level optimizations, such as kernel fusion and the elimination of CPU-GPU blocking, to further enhance both training and inference throughput. Offline and online experiments validate the effectiveness of MTFM, demonstrating that significant performance gains are achieved by scaling both model capacity and multi-scenario training data.
Generative Modeling through Spectral Analysis of Koopman Operator
Dec 21, 2025



Abstract:We propose Koopman Spectral Wasserstein Gradient Descent (KSWGD), a generative modeling framework that combines operator-theoretic spectral analysis with optimal transport. The novel insight is that the spectral structure required for accelerated Wasserstein gradient descent can be directly estimated from trajectory data via Koopman operator approximation which can eliminate the need for explicit knowledge of the target potential or neural network training. We provide rigorous convergence analysis and establish connection to Feynman-Kac theory that clarifies the method's probabilistic foundation. Experiments across diverse settings, including compact manifold sampling, metastable multi-well systems, image generation, and high dimensional stochastic partial differential equation, demonstrate that KSWGD consistently achieves faster convergence than other existing methods while maintaining high sample quality.
BP-Seg: A graphical model approach to unsupervised and non-contiguous text segmentation using belief propagation
May 22, 2025

Abstract:Text segmentation based on the semantic meaning of sentences is a fundamental task with broad utility in many downstream applications. In this paper, we propose a graphical model-based unsupervised learning approach, named BP-Seg for efficient text segmentation. Our method not only considers local coherence, capturing the intuition that adjacent sentences are often more related, but also effectively groups sentences that are distant in the text yet semantically similar. This is achieved through belief propagation on the carefully constructed graphical models. Experimental results on both an illustrative example and a dataset with long-form documents demonstrate that our method performs favorably compared to competing approaches.
Expected Information Gain Estimation via Density Approximations: Sample Allocation and Dimension Reduction
Nov 13, 2024Abstract:Computing expected information gain (EIG) from prior to posterior (equivalently, mutual information between candidate observations and model parameters or other quantities of interest) is a fundamental challenge in Bayesian optimal experimental design. We formulate flexible transport-based schemes for EIG estimation in general nonlinear/non-Gaussian settings, compatible with both standard and implicit Bayesian models. These schemes are representative of two-stage methods for estimating or bounding EIG using marginal and conditional density estimates. In this setting, we analyze the optimal allocation of samples between training (density estimation) and approximation of the outer prior expectation. We show that with this optimal sample allocation, the MSE of the resulting EIG estimator converges more quickly than that of a standard nested Monte Carlo scheme. We then address the estimation of EIG in high dimensions, by deriving gradient-based upper bounds on the mutual information lost by projecting the parameters and/or observations to lower-dimensional subspaces. Minimizing these upper bounds yields projectors and hence low-dimensional EIG approximations that outperform approximations obtained via other linear dimension reduction schemes. Numerical experiments on a PDE-constrained Bayesian inverse problem also illustrate a favorable trade-off between dimension truncation and the modeling of non-Gaussianity, when estimating EIG from finite samples in high dimensions.
Sharp detection of low-dimensional structure in probability measures via dimensional logarithmic Sobolev inequalities
Jun 18, 2024


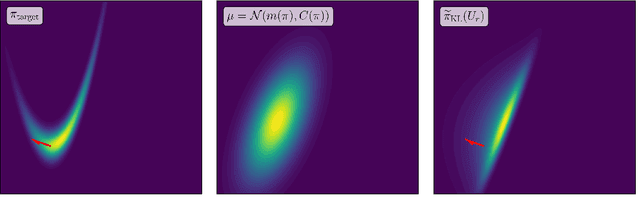
Abstract:Identifying low-dimensional structure in high-dimensional probability measures is an essential pre-processing step for efficient sampling. We introduce a method for identifying and approximating a target measure $\pi$ as a perturbation of a given reference measure $\mu$ along a few significant directions of $\mathbb{R}^{d}$. The reference measure can be a Gaussian or a nonlinear transformation of a Gaussian, as commonly arising in generative modeling. Our method extends prior work on minimizing majorizations of the Kullback--Leibler divergence to identify optimal approximations within this class of measures. Our main contribution unveils a connection between the \emph{dimensional} logarithmic Sobolev inequality (LSI) and approximations with this ansatz. Specifically, when the target and reference are both Gaussian, we show that minimizing the dimensional LSI is equivalent to minimizing the KL divergence restricted to this ansatz. For general non-Gaussian measures, the dimensional LSI produces majorants that uniformly improve on previous majorants for gradient-based dimension reduction. We further demonstrate the applicability of this analysis to the squared Hellinger distance, where analogous reasoning shows that the dimensional Poincar\'e inequality offers improved bounds.
Nonlinear Bayesian optimal experimental design using logarithmic Sobolev inequalities
Feb 23, 2024Abstract:We study the problem of selecting $k$ experiments from a larger candidate pool, where the goal is to maximize mutual information (MI) between the selected subset and the underlying parameters. Finding the exact solution is to this combinatorial optimization problem is computationally costly, not only due to the complexity of the combinatorial search but also the difficulty of evaluating MI in nonlinear/non-Gaussian settings. We propose greedy approaches based on new computationally inexpensive lower bounds for MI, constructed via log-Sobolev inequalities. We demonstrate that our method outperforms random selection strategies, Gaussian approximations, and nested Monte Carlo (NMC) estimators of MI in various settings, including optimal design for nonlinear models with non-additive noise.
Stable generative modeling using diffusion maps
Jan 09, 2024Abstract:We consider the problem of sampling from an unknown distribution for which only a sufficiently large number of training samples are available. Such settings have recently drawn considerable interest in the context of generative modelling. In this paper, we propose a generative model combining diffusion maps and Langevin dynamics. Diffusion maps are used to approximate the drift term from the available training samples, which is then implemented in a discrete-time Langevin sampler to generate new samples. By setting the kernel bandwidth to match the time step size used in the unadjusted Langevin algorithm, our method effectively circumvents any stability issues typically associated with time-stepping stiff stochastic differential equations. More precisely, we introduce a novel split-step scheme, ensuring that the generated samples remain within the convex hull of the training samples. Our framework can be naturally extended to generate conditional samples. We demonstrate the performance of our proposed scheme through experiments on synthetic datasets with increasing dimensions and on a stochastic subgrid-scale parametrization conditional sampling problem.
Diffusion map particle systems for generative modeling
Apr 01, 2023



Abstract:We propose a novel diffusion map particle system (DMPS) for generative modeling, based on diffusion maps and Laplacian-adjusted Wasserstein gradient descent (LAWGD). Diffusion maps are used to approximate the generator of the Langevin diffusion process from samples, and hence to learn the underlying data-generating manifold. On the other hand, LAWGD enables efficient sampling from the target distribution given a suitable choice of kernel, which we construct here via a spectral approximation of the generator, computed with diffusion maps. Numerical experiments show that our method outperforms others on synthetic datasets, including examples with manifold structure.
Bayesian model calibration for block copolymer self-assembly: Likelihood-free inference and expected information gain computation via measure transport
Jun 22, 2022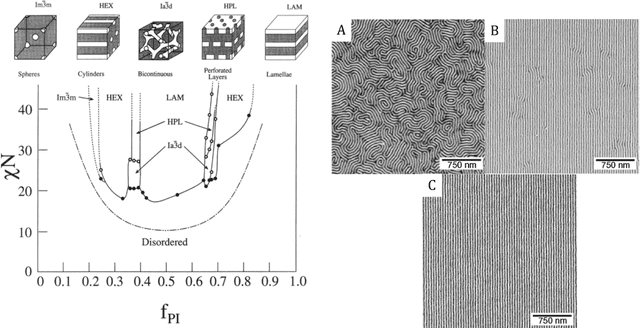
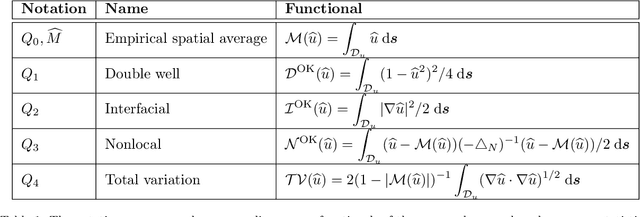
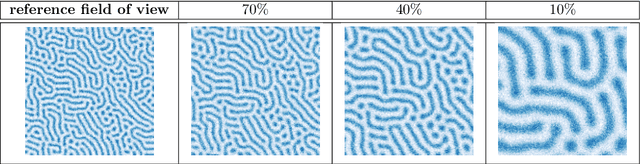
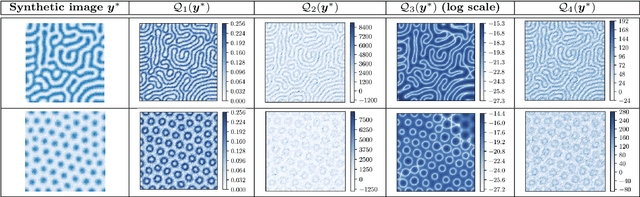
Abstract:We consider the Bayesian calibration of models describing the phenomenon of block copolymer (BCP) self-assembly using image data produced by microscopy or X-ray scattering techniques. To account for the random long-range disorder in BCP equilibrium structures, we introduce auxiliary variables to represent this aleatory uncertainty. These variables, however, result in an integrated likelihood for high-dimensional image data that is generally intractable to evaluate. We tackle this challenging Bayesian inference problem using a likelihood-free approach based on measure transport together with the construction of summary statistics for the image data. We also show that expected information gains (EIGs) from the observed data about the model parameters can be computed with no significant additional cost. Lastly, we present a numerical case study based on the Ohta--Kawasaki model for diblock copolymer thin film self-assembly and top-down microscopy characterization. For calibration, we introduce several domain-specific energy- and Fourier-based summary statistics, and quantify their informativeness using EIG. We demonstrate the power of the proposed approach to study the effect of data corruptions and experimental designs on the calibration results.
Automatic Fine-grained Glomerular Lesion Recognition in Kidney Pathology
Mar 11, 2022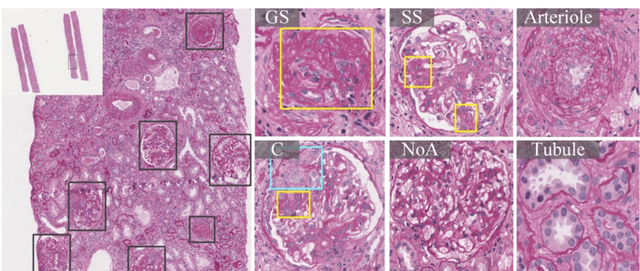
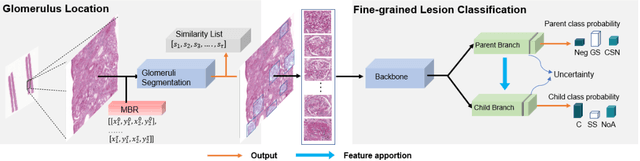
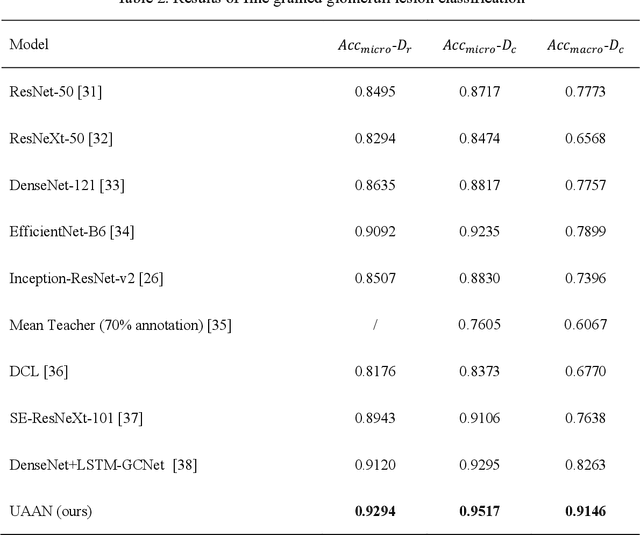
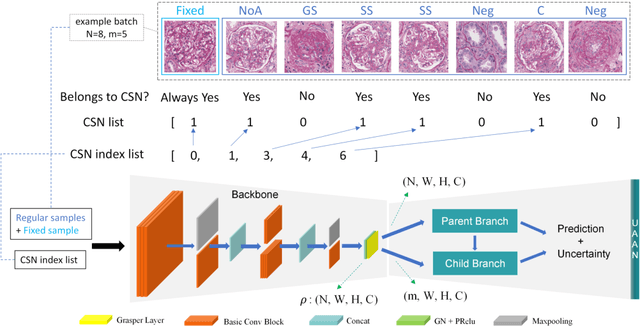
Abstract:Recognition of glomeruli lesions is the key for diagnosis and treatment planning in kidney pathology; however, the coexisting glomerular structures such as mesangial regions exacerbate the difficulties of this task. In this paper, we introduce a scheme to recognize fine-grained glomeruli lesions from whole slide images. First, a focal instance structural similarity loss is proposed to drive the model to locate all types of glomeruli precisely. Then an Uncertainty Aided Apportionment Network is designed to carry out the fine-grained visual classification without bounding-box annotations. This double branch-shaped structure extracts common features of the child class from the parent class and produces the uncertainty factor for reconstituting the training dataset. Results of slide-wise evaluation illustrate the effectiveness of the entire scheme, with an 8-22% improvement of the mean Average Precision compared with remarkable detection methods. The comprehensive results clearly demonstrate the effectiveness of the proposed method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge