Dong Ni
MUVF-YOLOX: A Multi-modal Ultrasound Video Fusion Network for Renal Tumor Diagnosis
Jul 15, 2023Abstract:Early diagnosis of renal cancer can greatly improve the survival rate of patients. Contrast-enhanced ultrasound (CEUS) is a cost-effective and non-invasive imaging technique and has become more and more frequently used for renal tumor diagnosis. However, the classification of benign and malignant renal tumors can still be very challenging due to the highly heterogeneous appearance of cancer and imaging artifacts. Our aim is to detect and classify renal tumors by integrating B-mode and CEUS-mode ultrasound videos. To this end, we propose a novel multi-modal ultrasound video fusion network that can effectively perform multi-modal feature fusion and video classification for renal tumor diagnosis. The attention-based multi-modal fusion module uses cross-attention and self-attention to extract modality-invariant features and modality-specific features in parallel. In addition, we design an object-level temporal aggregation (OTA) module that can automatically filter low-quality features and efficiently integrate temporal information from multiple frames to improve the accuracy of tumor diagnosis. Experimental results on a multicenter dataset show that the proposed framework outperforms the single-modal models and the competing methods. Furthermore, our OTA module achieves higher classification accuracy than the frame-level predictions. Our code is available at \url{https://github.com/JeunyuLi/MUAF}.
GSMorph: Gradient Surgery for cine-MRI Cardiac Deformable Registration
Jun 26, 2023Abstract:Deep learning-based deformable registration methods have been widely investigated in diverse medical applications. Learning-based deformable registration relies on weighted objective functions trading off registration accuracy and smoothness of the deformation field. Therefore, they inevitably require tuning the hyperparameter for optimal registration performance. Tuning the hyperparameters is highly computationally expensive and introduces undesired dependencies on domain knowledge. In this study, we construct a registration model based on the gradient surgery mechanism, named GSMorph, to achieve a hyperparameter-free balance on multiple losses. In GSMorph, we reformulate the optimization procedure by projecting the gradient of similarity loss orthogonally to the plane associated with the smoothness constraint, rather than additionally introducing a hyperparameter to balance these two competing terms. Furthermore, our method is model-agnostic and can be merged into any deep registration network without introducing extra parameters or slowing down inference. In this study, We compared our method with state-of-the-art (SOTA) deformable registration approaches over two publicly available cardiac MRI datasets. GSMorph proves superior to five SOTA learning-based registration models and two conventional registration techniques, SyN and Demons, on both registration accuracy and smoothness.
Inflated 3D Convolution-Transformer for Weakly-supervised Carotid Stenosis Grading with Ultrasound Videos
Jun 12, 2023



Abstract:Localization of the narrowest position of the vessel and corresponding vessel and remnant vessel delineation in carotid ultrasound (US) are essential for carotid stenosis grading (CSG) in clinical practice. However, the pipeline is time-consuming and tough due to the ambiguous boundaries of plaque and temporal variation. To automatize this procedure, a large number of manual delineations are usually required, which is not only laborious but also not reliable given the annotation difficulty. In this study, we present the first video classification framework for automatic CSG. Our contribution is three-fold. First, to avoid the requirement of laborious and unreliable annotation, we propose a novel and effective video classification network for weakly-supervised CSG. Second, to ease the model training, we adopt an inflation strategy for the network, where pre-trained 2D convolution weights can be adapted into the 3D counterpart in our network for an effective warm start. Third, to enhance the feature discrimination of the video, we propose a novel attention-guided multi-dimension fusion (AMDF) transformer encoder to model and integrate global dependencies within and across spatial and temporal dimensions, where two lightweight cross-dimensional attention mechanisms are designed. Our approach is extensively validated on a large clinically collected carotid US video dataset, demonstrating state-of-the-art performance compared with strong competitors.
ModeT: Learning Deformable Image Registration via Motion Decomposition Transformer
Jun 09, 2023Abstract:The Transformer structures have been widely used in computer vision and have recently made an impact in the area of medical image registration. However, the use of Transformer in most registration networks is straightforward. These networks often merely use the attention mechanism to boost the feature learning as the segmentation networks do, but do not sufficiently design to be adapted for the registration task. In this paper, we propose a novel motion decomposition Transformer (ModeT) to explicitly model multiple motion modalities by fully exploiting the intrinsic capability of the Transformer structure for deformation estimation. The proposed ModeT naturally transforms the multi-head neighborhood attention relationship into the multi-coordinate relationship to model multiple motion modes. Then the competitive weighting module (CWM) fuses multiple deformation sub-fields to generate the resulting deformation field. Extensive experiments on two public brain magnetic resonance imaging (MRI) datasets show that our method outperforms current state-of-the-art registration networks and Transformers, demonstrating the potential of our ModeT for the challenging non-rigid deformation estimation problem. The benchmarks and our code are publicly available at https://github.com/ZAX130/SmileCode.
Instructive Feature Enhancement for Dichotomous Medical Image Segmentation
Jun 06, 2023



Abstract:Deep neural networks have been widely applied in dichotomous medical image segmentation (DMIS) of many anatomical structures in several modalities, achieving promising performance. However, existing networks tend to struggle with task-specific, heavy and complex designs to improve accuracy. They made little instructions to which feature channels would be more beneficial for segmentation, and that may be why the performance and universality of these segmentation models are hindered. In this study, we propose an instructive feature enhancement approach, namely IFE, to adaptively select feature channels with rich texture cues and strong discriminability to enhance raw features based on local curvature or global information entropy criteria. Being plug-and-play and applicable for diverse DMIS tasks, IFE encourages the model to focus on texture-rich features which are especially important for the ambiguous and challenging boundary identification, simultaneously achieving simplicity, universality, and certain interpretability. To evaluate the proposed IFE, we constructed the first large-scale DMIS dataset Cosmos55k, which contains 55,023 images from 7 modalities and 26 anatomical structures. Extensive experiments show that IFE can improve the performance of classic segmentation networks across different anatomies and modalities with only slight modifications. Code is available at https://github.com/yezi-66/IFE
Fourier Test-time Adaptation with Multi-level Consistency for Robust Classification
Jun 05, 2023



Abstract:Deep classifiers may encounter significant performance degradation when processing unseen testing data from varying centers, vendors, and protocols. Ensuring the robustness of deep models against these domain shifts is crucial for their widespread clinical application. In this study, we propose a novel approach called Fourier Test-time Adaptation (FTTA), which employs a dual-adaptation design to integrate input and model tuning, thereby jointly improving the model robustness. The main idea of FTTA is to build a reliable multi-level consistency measurement of paired inputs for achieving self-correction of prediction. Our contribution is two-fold. First, we encourage consistency in global features and local attention maps between the two transformed images of the same input. Here, the transformation refers to Fourier-based input adaptation, which can transfer one unseen image into source style to reduce the domain gap. Furthermore, we leverage style-interpolated images to enhance the global and local features with learnable parameters, which can smooth the consistency measurement and accelerate convergence. Second, we introduce a regularization technique that utilizes style interpolation consistency in the frequency space to encourage self-consistency in the logit space of the model output. This regularization provides strong self-supervised signals for robustness enhancement. FTTA was extensively validated on three large classification datasets with different modalities and organs. Experimental results show that FTTA is general and outperforms other strong state-of-the-art methods.
Segment Anything Model for Medical Images?
May 01, 2023



Abstract:The Segment Anything Model (SAM) is the first foundation model for general image segmentation. It designed a novel promotable segmentation task, ensuring zero-shot image segmentation using the pre-trained model via two main modes including automatic everything and manual prompt. SAM has achieved impressive results on various natural image segmentation tasks. However, medical image segmentation (MIS) is more challenging due to the complex modalities, fine anatomical structures, uncertain and complex object boundaries, and wide-range object scales. Meanwhile, zero-shot and efficient MIS can well reduce the annotation time and boost the development of medical image analysis. Hence, SAM seems to be a potential tool and its performance on large medical datasets should be further validated. We collected and sorted 52 open-source datasets, and built a large medical segmentation dataset with 16 modalities, 68 objects, and 553K slices. We conducted a comprehensive analysis of different SAM testing strategies on the so-called COSMOS 553K dataset. Extensive experiments validate that SAM performs better with manual hints like points and boxes for object perception in medical images, leading to better performance in prompt mode compared to everything mode. Additionally, SAM shows remarkable performance in some specific objects and modalities, but is imperfect or even totally fails in other situations. Finally, we analyze the influence of different factors (e.g., the Fourier-based boundary complexity and size of the segmented objects) on SAM's segmentation performance. Extensive experiments validate that SAM's zero-shot segmentation capability is not sufficient to ensure its direct application to the MIS.
Hierarchical Agent-based Reinforcement Learning Framework for Automated Quality Assessment of Fetal Ultrasound Video
Apr 14, 2023


Abstract:Ultrasound is the primary modality to examine fetal growth during pregnancy, while the image quality could be affected by various factors. Quality assessment is essential for controlling the quality of ultrasound images to guarantee both the perceptual and diagnostic values. Existing automated approaches often require heavy structural annotations and the predictions may not necessarily be consistent with the assessment results by human experts. Furthermore, the overall quality of a scan and the correlation between the quality of frames should not be overlooked. In this work, we propose a reinforcement learning framework powered by two hierarchical agents that collaboratively learn to perform both frame-level and video-level quality assessments. It is equipped with a specially-designed reward mechanism that considers temporal dependency among frame quality and only requires sparse binary annotations to train. Experimental results on a challenging fetal brain dataset verify that the proposed framework could perform dual-level quality assessment and its predictions correlate well with the subjective assessment results.
RLIP: Relational Language-Image Pre-training for Human-Object Interaction Detection
Sep 05, 2022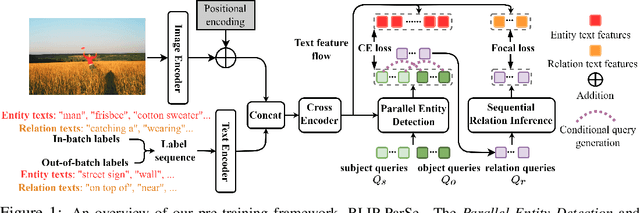
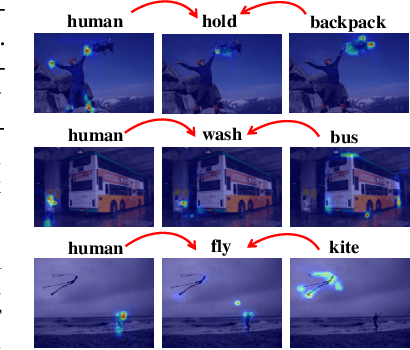
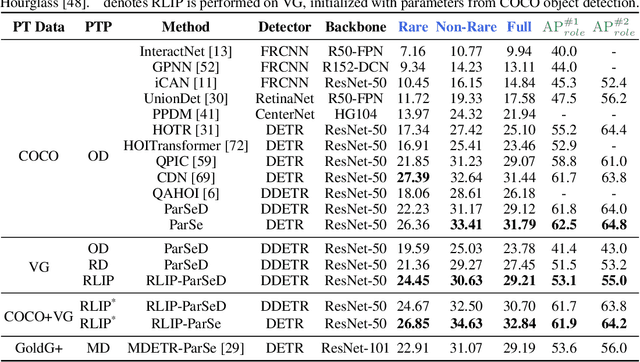
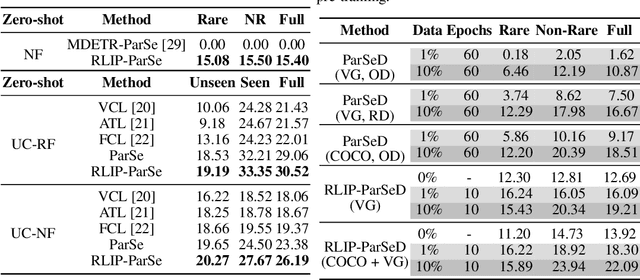

Abstract:The task of Human-Object Interaction (HOI) detection targets fine-grained visual parsing of humans interacting with their environment, enabling a broad range of applications. Prior work has demonstrated the benefits of effective architecture design and integration of relevant cues for more accurate HOI detection. However, the design of an appropriate pre-training strategy for this task remains underexplored by existing approaches. To address this gap, we propose Relational Language-Image Pre-training (RLIP), a strategy for contrastive pre-training that leverages both entity and relation descriptions. To make effective use of such pre-training, we make three technical contributions: (1) a new Parallel entity detection and Sequential relation inference (ParSe) architecture that enables the use of both entity and relation descriptions during holistically optimized pre-training; (2) a synthetic data generation framework, Label Sequence Extension, that expands the scale of language data available within each minibatch; (3) mechanisms to account for ambiguity, Relation Quality Labels and Relation Pseudo-Labels, to mitigate the influence of ambiguous/noisy samples in the pre-training data. Through extensive experiments, we demonstrate the benefits of these contributions, collectively termed RLIP-ParSe, for improved zero-shot, few-shot and fine-tuning HOI detection performance as well as increased robustness to learning from noisy annotations. Code will be available at \url{https://github.com/JacobYuan7/RLIP}.
Masked Video Modeling with Correlation-aware Contrastive Learning for Breast Cancer Diagnosis in Ultrasound
Aug 21, 2022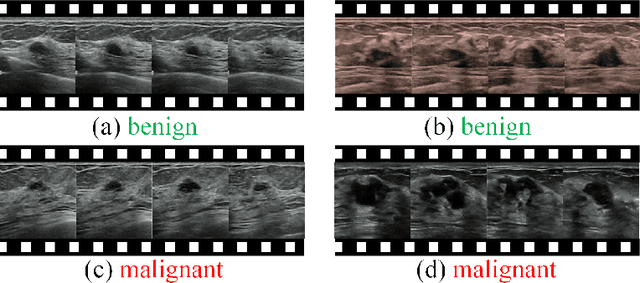
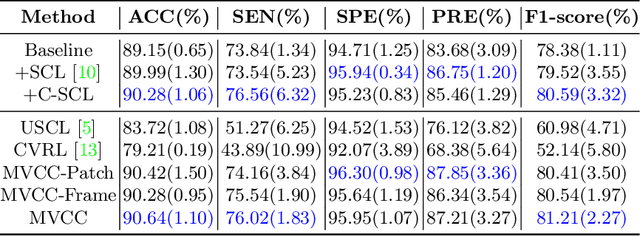
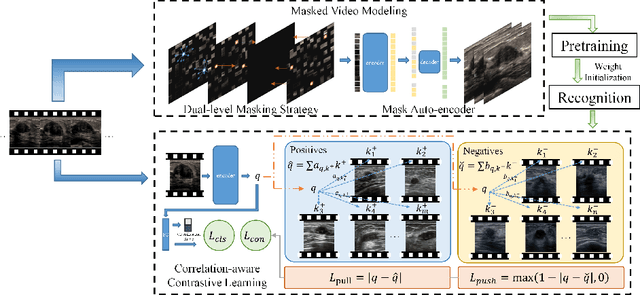
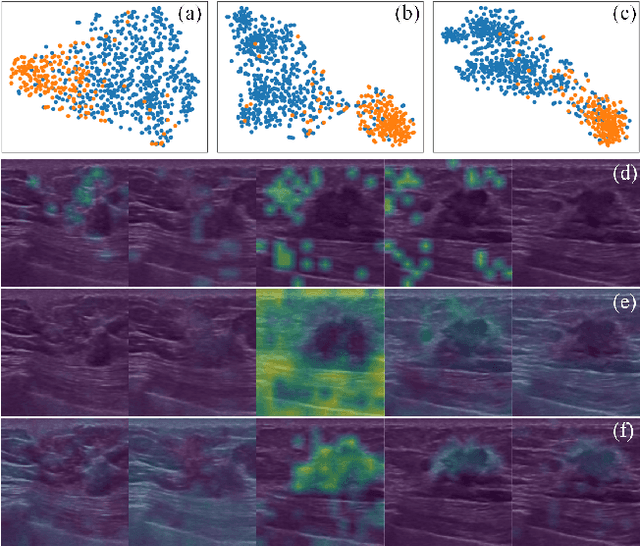
Abstract:Breast cancer is one of the leading causes of cancer deaths in women. As the primary output of breast screening, breast ultrasound (US) video contains exclusive dynamic information for cancer diagnosis. However, training models for video analysis is non-trivial as it requires a voluminous dataset which is also expensive to annotate. Furthermore, the diagnosis of breast lesion faces unique challenges such as inter-class similarity and intra-class variation. In this paper, we propose a pioneering approach that directly utilizes US videos in computer-aided breast cancer diagnosis. It leverages masked video modeling as pretraning to reduce reliance on dataset size and detailed annotations. Moreover, a correlation-aware contrastive loss is developed to facilitate the identifying of the internal and external relationship between benign and malignant lesions. Experimental results show that our proposed approach achieved promising classification performance and can outperform other state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge