Daniel Sonntag
The Speed-up Factor: A Quantitative Multi-Iteration Active Learning Performance Metric
Feb 13, 2026Abstract:Machine learning models excel with abundant annotated data, but annotation is often costly and time-intensive. Active learning (AL) aims to improve the performance-to-annotation ratio by using query methods (QMs) to iteratively select the most informative samples. While AL research focuses mainly on QM development, the evaluation of this iterative process lacks appropriate performance metrics. This work reviews eight years of AL evaluation literature and formally introduces the speed-up factor, a quantitative multi-iteration QM performance metric that indicates the fraction of samples needed to match random sampling performance. Using four datasets from diverse domains and seven QMs of various types, we empirically evaluate the speed-up factor and compare it with state-of-the-art AL performance metrics. The results confirm the assumptions underlying the speed-up factor, demonstrate its accuracy in capturing the described fraction, and reveal its superior stability across iterations.
How Many Tokens Do 3D Point Cloud Transformer Architectures Really Need?
Nov 07, 2025Abstract:Recent advances in 3D point cloud transformers have led to state-of-the-art results in tasks such as semantic segmentation and reconstruction. However, these models typically rely on dense token representations, incurring high computational and memory costs during training and inference. In this work, we present the finding that tokens are remarkably redundant, leading to substantial inefficiency. We introduce gitmerge3D, a globally informed graph token merging method that can reduce the token count by up to 90-95% while maintaining competitive performance. This finding challenges the prevailing assumption that more tokens inherently yield better performance and highlights that many current models are over-tokenized and under-optimized for scalability. We validate our method across multiple 3D vision tasks and show consistent improvements in computational efficiency. This work is the first to assess redundancy in large-scale 3D transformer models, providing insights into the development of more efficient 3D foundation architectures. Our code and checkpoints are publicly available at https://gitmerge3d.github.io
S-Chain: Structured Visual Chain-of-Thought For Medicine
Oct 26, 2025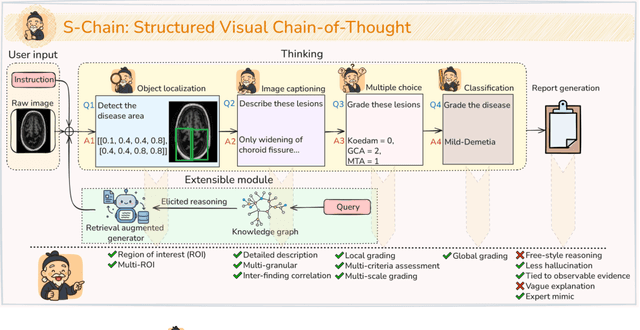
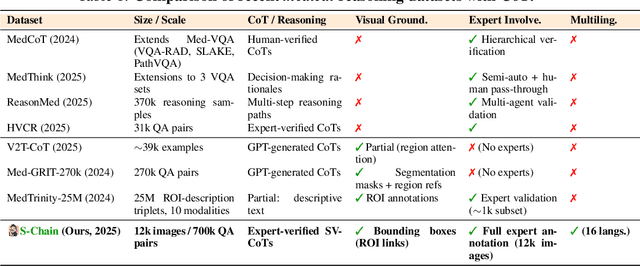
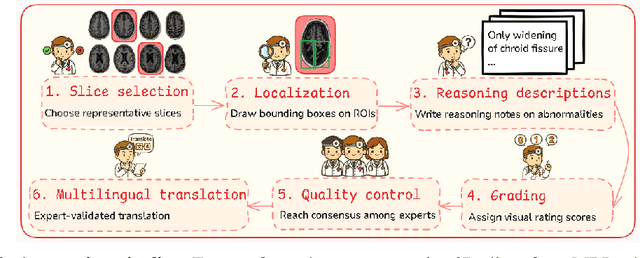
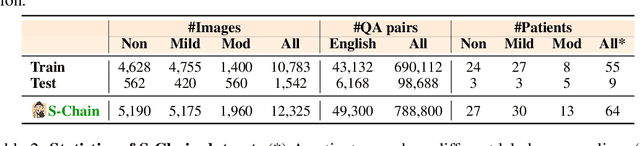
Abstract:Faithful reasoning in medical vision-language models (VLMs) requires not only accurate predictions but also transparent alignment between textual rationales and visual evidence. While Chain-of-Thought (CoT) prompting has shown promise in medical visual question answering (VQA), no large-scale expert-level dataset has captured stepwise reasoning with precise visual grounding. We introduce S-Chain, the first large-scale dataset of 12,000 expert-annotated medical images with bounding boxes and structured visual CoT (SV-CoT), explicitly linking visual regions to reasoning steps. The dataset further supports 16 languages, totaling over 700k VQA pairs for broad multilingual applicability. Using S-Chain, we benchmark state-of-the-art medical VLMs (ExGra-Med, LLaVA-Med) and general-purpose VLMs (Qwen2.5-VL, InternVL2.5), showing that SV-CoT supervision significantly improves interpretability, grounding fidelity, and robustness. Beyond benchmarking, we study its synergy with retrieval-augmented generation, revealing how domain knowledge and visual grounding interact during autoregressive reasoning. Finally, we propose a new mechanism that strengthens the alignment between visual evidence and reasoning, improving both reliability and efficiency. S-Chain establishes a new benchmark for grounded medical reasoning and paves the way toward more trustworthy and explainable medical VLMs.
Mitigating Reward Over-optimization in Direct Alignment Algorithms with Importance Sampling
Jun 11, 2025Abstract:Direct Alignment Algorithms (DAAs) such as Direct Preference Optimization (DPO) have emerged as alternatives to the standard Reinforcement Learning from Human Feedback (RLHF) for aligning large language models (LLMs) with human values. However, these methods are more susceptible to over-optimization, in which the model drifts away from the reference policy, leading to degraded performance as training progresses. This paper proposes a novel importance-sampling approach to mitigate the over-optimization problem of offline DAAs. This approach, called (IS-DAAs), multiplies the DAA objective with an importance ratio that accounts for the reference policy distribution. IS-DAAs additionally avoid the high variance issue associated with importance sampling by clipping the importance ratio to a maximum value. Our extensive experiments demonstrate that IS-DAAs can effectively mitigate over-optimization, especially under low regularization strength, and achieve better performance than other methods designed to address this problem. Our implementations are provided publicly at this link.
CBM-RAG: Demonstrating Enhanced Interpretability in Radiology Report Generation with Multi-Agent RAG and Concept Bottleneck Models
Apr 29, 2025
Abstract:Advancements in generative Artificial Intelligence (AI) hold great promise for automating radiology workflows, yet challenges in interpretability and reliability hinder clinical adoption. This paper presents an automated radiology report generation framework that combines Concept Bottleneck Models (CBMs) with a Multi-Agent Retrieval-Augmented Generation (RAG) system to bridge AI performance with clinical explainability. CBMs map chest X-ray features to human-understandable clinical concepts, enabling transparent disease classification. Meanwhile, the RAG system integrates multi-agent collaboration and external knowledge to produce contextually rich, evidence-based reports. Our demonstration showcases the system's ability to deliver interpretable predictions, mitigate hallucinations, and generate high-quality, tailored reports with an interactive interface addressing accuracy, trust, and usability challenges. This framework provides a pathway to improving diagnostic consistency and empowering radiologists with actionable insights.
InFL-UX: A Toolkit for Web-Based Interactive Federated Learning
Mar 06, 2025

Abstract:This paper presents InFL-UX, an interactive, proof-of-concept browser-based Federated Learning (FL) toolkit designed to integrate user contributions seamlessly into the machine learning (ML) workflow. InFL-UX enables users across multiple devices to upload datasets, define classes, and collaboratively train classification models directly in the browser using modern web technologies. Unlike traditional FL toolkits, which often focus on backend simulations, InFL-UX provides a simple user interface for researchers to explore how users interact with and contribute to FL systems in real-world, interactive settings. By prioritising usability and decentralised model training, InFL-UX bridges the gap between FL and Interactive Machine Learning (IML), empowering non-technical users to actively participate in ML classification tasks.
MGPATH: Vision-Language Model with Multi-Granular Prompt Learning for Few-Shot WSI Classification
Feb 11, 2025



Abstract:Whole slide pathology image classification presents challenges due to gigapixel image sizes and limited annotation labels, hindering model generalization. This paper introduces a prompt learning method to adapt large vision-language models for few-shot pathology classification. We first extend the Prov-GigaPath vision foundation model, pre-trained on 1.3 billion pathology image tiles, into a vision-language model by adding adaptors and aligning it with medical text encoders via contrastive learning on 923K image-text pairs. The model is then used to extract visual features and text embeddings from few-shot annotations and fine-tunes with learnable prompt embeddings. Unlike prior methods that combine prompts with frozen features using prefix embeddings or self-attention, we propose multi-granular attention that compares interactions between learnable prompts with individual image patches and groups of them. This approach improves the model's ability to capture both fine-grained details and broader context, enhancing its recognition of complex patterns across sub-regions. To further improve accuracy, we leverage (unbalanced) optimal transport-based visual-text distance to secure model robustness by mitigating perturbations that might occur during the data augmentation process. Empirical experiments on lung, kidney, and breast pathology modalities validate the effectiveness of our approach; thereby, we surpass several of the latest competitors and consistently improve performance across diverse architectures, including CLIP, PLIP, and Prov-GigaPath integrated PLIP. We release our implementations and pre-trained models at this MGPATH.
Enhancing Online Learning Efficiency Through Heterogeneous Resource Integration with a Multi-Agent RAG System
Feb 06, 2025


Abstract:Efficient online learning requires seamless access to diverse resources such as videos, code repositories, documentation, and general web content. This poster paper introduces early-stage work on a Multi-Agent Retrieval-Augmented Generation (RAG) System designed to enhance learning efficiency by integrating these heterogeneous resources. Using specialized agents tailored for specific resource types (e.g., YouTube tutorials, GitHub repositories, documentation websites, and search engines), the system automates the retrieval and synthesis of relevant information. By streamlining the process of finding and combining knowledge, this approach reduces manual effort and enhances the learning experience. A preliminary user study confirmed the system's strong usability and moderate-high utility, demonstrating its potential to improve the efficiency of knowledge acquisition.
On Zero-Initialized Attention: Optimal Prompt and Gating Factor Estimation
Feb 05, 2025



Abstract:The LLaMA-Adapter has recently emerged as an efficient fine-tuning technique for LLaMA models, leveraging zero-initialized attention to stabilize training and enhance performance. However, despite its empirical success, the theoretical foundations of zero-initialized attention remain largely unexplored. In this paper, we provide a rigorous theoretical analysis, establishing a connection between zero-initialized attention and mixture-of-expert models. We prove that both linear and non-linear prompts, along with gating functions, can be optimally estimated, with non-linear prompts offering greater flexibility for future applications. Empirically, we validate our findings on the open LLM benchmarks, demonstrating that non-linear prompts outperform linear ones. Notably, even with limited training data, both prompt types consistently surpass vanilla attention, highlighting the robustness and adaptability of zero-initialized attention.
Deep Learning for Ophthalmology: The State-of-the-Art and Future Trends
Jan 07, 2025



Abstract:The emergence of artificial intelligence (AI), particularly deep learning (DL), has marked a new era in the realm of ophthalmology, offering transformative potential for the diagnosis and treatment of posterior segment eye diseases. This review explores the cutting-edge applications of DL across a range of ocular conditions, including diabetic retinopathy, glaucoma, age-related macular degeneration, and retinal vessel segmentation. We provide a comprehensive overview of foundational ML techniques and advanced DL architectures, such as CNNs, attention mechanisms, and transformer-based models, highlighting the evolving role of AI in enhancing diagnostic accuracy, optimizing treatment strategies, and improving overall patient care. Additionally, we present key challenges in integrating AI solutions into clinical practice, including ensuring data diversity, improving algorithm transparency, and effectively leveraging multimodal data. This review emphasizes AI's potential to improve disease diagnosis and enhance patient care while stressing the importance of collaborative efforts to overcome these barriers and fully harness AI's impact in advancing eye care.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge