Mathias Niepert
SymDrift: One-Shot Generative Modeling under Symmetries
May 07, 2026Abstract:Generative modeling of physical systems, such as molecules, requires learning distributions that are invariant under global symmetries, such as rotations in three-dimensional space. Equivariant diffusion and flow matching models can incorporate such invariances effectively, even when trained on a non-invariant empirical distribution, but they typically rely on costly multi-step sampling. Recently, drifting models have emerged as an efficient alternative, enabling single-step generation and achieving state-of-the-art performance in generative modeling tasks. However, we show that drifting models face a symmetry-specific challenge, since an equivariant generator does not generally produce the same drifting field as the one obtained from the symmetrized target distribution. Addressing this issue would require expensive symmetrization of the empirical distribution. To avoid this cost, we propose SymDrift, a framework that makes the drifting field itself symmetry-aware. We introduce two complementary strategies: (i) a symmetrized drift in coordinate space based on optimal alignment, and (ii) a $G$-invariant embedding that removes symmetry ambiguity by construction. Empirically, SymDrift outperforms existing one-shot methods on standard benchmarks for conformer and transition state generation, while remaining competitive with significantly more expensive multi-step approaches. By enabling one-shot inference, SymDrift reduces computational overhead by up to 40$\times$ compared to existing baselines, making it promising for high-throughput applications such as virtual drug screening and large-scale reaction network exploration.
Logical Guidance for the Exact Composition of Diffusion Models
Feb 05, 2026Abstract:We propose LOGDIFF (Logical Guidance for the Exact Composition of Diffusion Models), a guidance framework for diffusion models that enables principled constrained generation with complex logical expressions at inference time. We study when exact score-based guidance for complex logical formulas can be obtained from guidance signals associated with atomic properties. First, we derive an exact Boolean calculus that provides a sufficient condition for exact logical guidance. Specifically, if a formula admits a circuit representation in which conjunctions combine conditionally independent subformulas and disjunctions combine subformulas that are either conditionally independent or mutually exclusive, exact logical guidance is achievable. In this case, the guidance signal can be computed exactly from atomic scores and posterior probabilities using an efficient recursive algorithm. Moreover, we show that, for commonly encountered classes of distributions, any desired Boolean formula is compilable into such a circuit representation. Second, by combining atomic guidance scores with posterior probability estimates, we introduce a hybrid guidance approach that bridges classifierguidance and classifier-free guidance, applicable to both compositional logical guidance and standard conditional generation. We demonstrate the effectiveness of our framework on multiple image and protein structure generation tasks.
GraIP: A Benchmarking Framework For Neural Graph Inverse Problems
Jan 26, 2026Abstract:A wide range of graph learning tasks, such as structure discovery, temporal graph analysis, and combinatorial optimization, focus on inferring graph structures from data, rather than making predictions on given graphs. However, the respective methods to solve such problems are often developed in an isolated, task-specific manner and thus lack a unifying theoretical foundation. Here, we provide a stepping stone towards the formation of such a foundation and further development by introducing the Neural Graph Inverse Problem (GraIP) conceptual framework, which formalizes and reframes a broad class of graph learning tasks as inverse problems. Unlike discriminative approaches that directly predict target variables from given graph inputs, the GraIP paradigm addresses inverse problems, i.e., it relies on observational data and aims to recover the underlying graph structure by reversing the forward process, such as message passing or network dynamics, that produced the observed outputs. We demonstrate the versatility of GraIP across various graph learning tasks, including rewiring, causal discovery, and neural relational inference. We also propose benchmark datasets and metrics for each GraIP domain considered, and characterize and empirically evaluate existing baseline methods used to solve them. Overall, our unifying perspective bridges seemingly disparate applications and provides a principled approach to structural learning in constrained and combinatorial settings while encouraging cross-pollination of existing methods across graph inverse problems.
SMART: Scalable Mesh-free Aerodynamic Simulations from Raw Geometries using a Transformer-based Surrogate Model
Jan 26, 2026Abstract:Machine learning-based surrogate models have emerged as more efficient alternatives to numerical solvers for physical simulations over complex geometries, such as car bodies. Many existing models incorporate the simulation mesh as an additional input, thereby reducing prediction errors. However, generating a simulation mesh for new geometries is computationally costly. In contrast, mesh-free methods, which do not rely on the simulation mesh, typically incur higher errors. Motivated by these considerations, we introduce SMART, a neural surrogate model that predicts physical quantities at arbitrary query locations using only a point-cloud representation of the geometry, without requiring access to the simulation mesh. The geometry and simulation parameters are encoded into a shared latent space that captures both structural and parametric characteristics of the physical field. A physics decoder then attends to the encoder's intermediate latent representations to map spatial queries to physical quantities. Through this cross-layer interaction, the model jointly updates latent geometric features and the evolving physical field. Extensive experiments show that SMART is competitive with and often outperforms existing methods that rely on the simulation mesh as input, demonstrating its capabilities for industry-level simulations.
How Many Tokens Do 3D Point Cloud Transformer Architectures Really Need?
Nov 07, 2025Abstract:Recent advances in 3D point cloud transformers have led to state-of-the-art results in tasks such as semantic segmentation and reconstruction. However, these models typically rely on dense token representations, incurring high computational and memory costs during training and inference. In this work, we present the finding that tokens are remarkably redundant, leading to substantial inefficiency. We introduce gitmerge3D, a globally informed graph token merging method that can reduce the token count by up to 90-95% while maintaining competitive performance. This finding challenges the prevailing assumption that more tokens inherently yield better performance and highlights that many current models are over-tokenized and under-optimized for scalability. We validate our method across multiple 3D vision tasks and show consistent improvements in computational efficiency. This work is the first to assess redundancy in large-scale 3D transformer models, providing insights into the development of more efficient 3D foundation architectures. Our code and checkpoints are publicly available at https://gitmerge3d.github.io
S-Chain: Structured Visual Chain-of-Thought For Medicine
Oct 26, 2025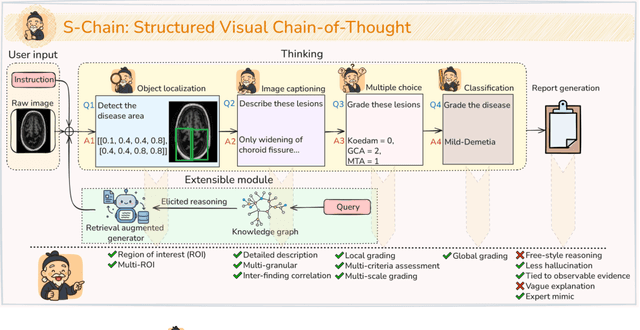
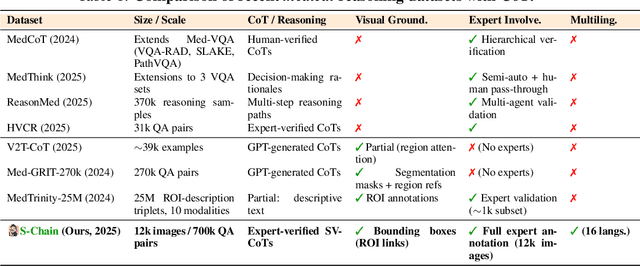
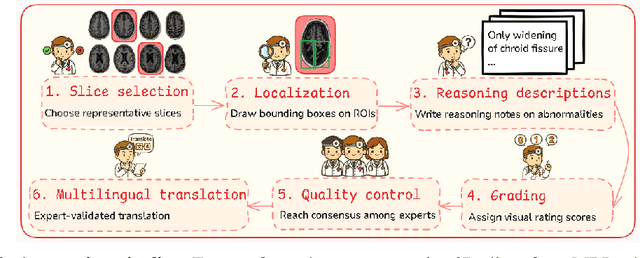
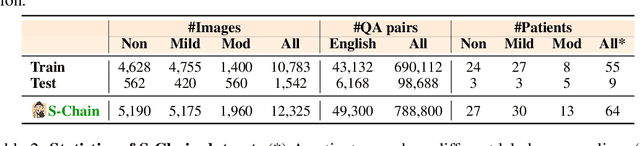
Abstract:Faithful reasoning in medical vision-language models (VLMs) requires not only accurate predictions but also transparent alignment between textual rationales and visual evidence. While Chain-of-Thought (CoT) prompting has shown promise in medical visual question answering (VQA), no large-scale expert-level dataset has captured stepwise reasoning with precise visual grounding. We introduce S-Chain, the first large-scale dataset of 12,000 expert-annotated medical images with bounding boxes and structured visual CoT (SV-CoT), explicitly linking visual regions to reasoning steps. The dataset further supports 16 languages, totaling over 700k VQA pairs for broad multilingual applicability. Using S-Chain, we benchmark state-of-the-art medical VLMs (ExGra-Med, LLaVA-Med) and general-purpose VLMs (Qwen2.5-VL, InternVL2.5), showing that SV-CoT supervision significantly improves interpretability, grounding fidelity, and robustness. Beyond benchmarking, we study its synergy with retrieval-augmented generation, revealing how domain knowledge and visual grounding interact during autoregressive reasoning. Finally, we propose a new mechanism that strengthens the alignment between visual evidence and reasoning, improving both reliability and efficiency. S-Chain establishes a new benchmark for grounded medical reasoning and paves the way toward more trustworthy and explainable medical VLMs.
Learning the Neighborhood: Contrast-Free Multimodal Self-Supervised Molecular Graph Pretraining
Sep 26, 2025



Abstract:High-quality molecular representations are essential for property prediction and molecular design, yet large labeled datasets remain scarce. While self-supervised pretraining on molecular graphs has shown promise, many existing approaches either depend on hand-crafted augmentations or complex generative objectives, and often rely solely on 2D topology, leaving valuable 3D structural information underutilized. To address this gap, we introduce C-FREE (Contrast-Free Representation learning on Ego-nets), a simple framework that integrates 2D graphs with ensembles of 3D conformers. C-FREE learns molecular representations by predicting subgraph embeddings from their complementary neighborhoods in the latent space, using fixed-radius ego-nets as modeling units across different conformers. This design allows us to integrate both geometric and topological information within a hybrid Graph Neural Network (GNN)-Transformer backbone, without negatives, positional encodings, or expensive pre-processing. Pretraining on the GEOM dataset, which provides rich 3D conformational diversity, C-FREE achieves state-of-the-art results on MoleculeNet, surpassing contrastive, generative, and other multimodal self-supervised methods. Fine-tuning across datasets with diverse sizes and molecule types further demonstrates that pretraining transfers effectively to new chemical domains, highlighting the importance of 3D-informed molecular representations.
CALM-PDE: Continuous and Adaptive Convolutions for Latent Space Modeling of Time-dependent PDEs
May 19, 2025Abstract:Solving time-dependent Partial Differential Equations (PDEs) using a densely discretized spatial domain is a fundamental problem in various scientific and engineering disciplines, including modeling climate phenomena and fluid dynamics. However, performing these computations directly in the physical space often incurs significant computational costs. To address this issue, several neural surrogate models have been developed that operate in a compressed latent space to solve the PDE. While these approaches reduce computational complexity, they often use Transformer-based attention mechanisms to handle irregularly sampled domains, resulting in increased memory consumption. In contrast, convolutional neural networks allow memory-efficient encoding and decoding but are limited to regular discretizations. Motivated by these considerations, we propose CALM-PDE, a model class that efficiently solves arbitrarily discretized PDEs in a compressed latent space. We introduce a novel continuous convolution-based encoder-decoder architecture that uses an epsilon-neighborhood-constrained kernel and learns to apply the convolution operator to adaptive and optimized query points. We demonstrate the effectiveness of CALM-PDE on a diverse set of PDEs with both regularly and irregularly sampled spatial domains. CALM-PDE is competitive with or outperforms existing baseline methods while offering significant improvements in memory and inference time efficiency compared to Transformer-based methods.
Learning (Approximately) Equivariant Networks via Constrained Optimization
May 19, 2025Abstract:Equivariant neural networks are designed to respect symmetries through their architecture, boosting generalization and sample efficiency when those symmetries are present in the data distribution. Real-world data, however, often departs from perfect symmetry because of noise, structural variation, measurement bias, or other symmetry-breaking effects. Strictly equivariant models may struggle to fit the data, while unconstrained models lack a principled way to leverage partial symmetries. Even when the data is fully symmetric, enforcing equivariance can hurt training by limiting the model to a restricted region of the parameter space. Guided by homotopy principles, where an optimization problem is solved by gradually transforming a simpler problem into a complex one, we introduce Adaptive Constrained Equivariance (ACE), a constrained optimization approach that starts with a flexible, non-equivariant model and gradually reduces its deviation from equivariance. This gradual tightening smooths training early on and settles the model at a data-driven equilibrium, balancing between equivariance and non-equivariance. Across multiple architectures and tasks, our method consistently improves performance metrics, sample efficiency, and robustness to input perturbations compared with strictly equivariant models and heuristic equivariance relaxations.
LOGLO-FNO: Efficient Learning of Local and Global Features in Fourier Neural Operators
Apr 05, 2025



Abstract:Modeling high-frequency information is a critical challenge in scientific machine learning. For instance, fully turbulent flow simulations of Navier-Stokes equations at Reynolds numbers 3500 and above can generate high-frequency signals due to swirling fluid motions caused by eddies and vortices. Faithfully modeling such signals using neural networks depends on accurately reconstructing moderate to high frequencies. However, it has been well known that deep neural nets exhibit the so-called spectral bias toward learning low-frequency components. Meanwhile, Fourier Neural Operators (FNOs) have emerged as a popular class of data-driven models in recent years for solving Partial Differential Equations (PDEs) and for surrogate modeling in general. Although impressive results have been achieved on several PDE benchmark problems, FNOs often perform poorly in learning non-dominant frequencies characterized by local features. This limitation stems from the spectral bias inherent in neural networks and the explicit exclusion of high-frequency modes in FNOs and their variants. Therefore, to mitigate these issues and improve FNO's spectral learning capabilities to represent a broad range of frequency components, we propose two key architectural enhancements: (i) a parallel branch performing local spectral convolutions (ii) a high-frequency propagation module. Moreover, we propose a novel frequency-sensitive loss term based on radially binned spectral errors. This introduction of a parallel branch for local convolutions reduces number of trainable parameters by up to 50% while achieving the accuracy of baseline FNO that relies solely on global convolutions. Experiments on three challenging PDE problems in fluid mechanics and biological pattern formation, and the qualitative and spectral analysis of predictions show the effectiveness of our method over the state-of-the-art neural operator baselines.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge