Duy M. H. Nguyen
How Many Tokens Do 3D Point Cloud Transformer Architectures Really Need?
Nov 07, 2025Abstract:Recent advances in 3D point cloud transformers have led to state-of-the-art results in tasks such as semantic segmentation and reconstruction. However, these models typically rely on dense token representations, incurring high computational and memory costs during training and inference. In this work, we present the finding that tokens are remarkably redundant, leading to substantial inefficiency. We introduce gitmerge3D, a globally informed graph token merging method that can reduce the token count by up to 90-95% while maintaining competitive performance. This finding challenges the prevailing assumption that more tokens inherently yield better performance and highlights that many current models are over-tokenized and under-optimized for scalability. We validate our method across multiple 3D vision tasks and show consistent improvements in computational efficiency. This work is the first to assess redundancy in large-scale 3D transformer models, providing insights into the development of more efficient 3D foundation architectures. Our code and checkpoints are publicly available at https://gitmerge3d.github.io
S-Chain: Structured Visual Chain-of-Thought For Medicine
Oct 26, 2025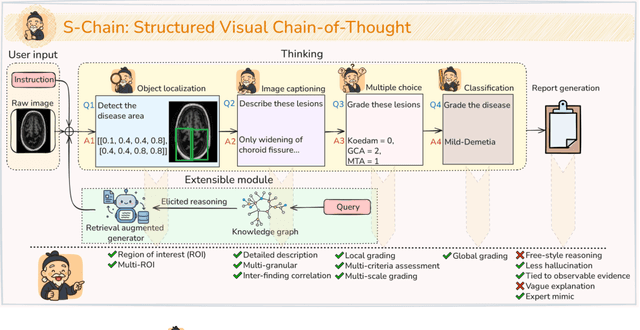
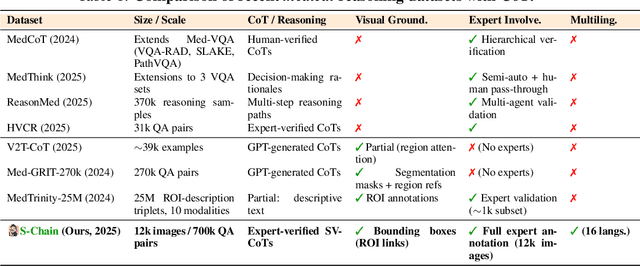
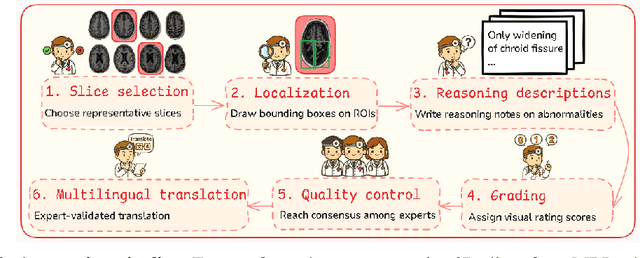
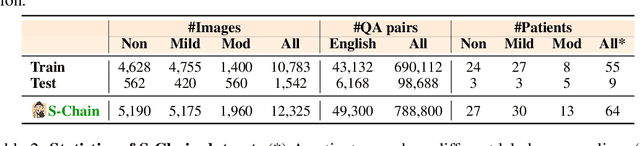
Abstract:Faithful reasoning in medical vision-language models (VLMs) requires not only accurate predictions but also transparent alignment between textual rationales and visual evidence. While Chain-of-Thought (CoT) prompting has shown promise in medical visual question answering (VQA), no large-scale expert-level dataset has captured stepwise reasoning with precise visual grounding. We introduce S-Chain, the first large-scale dataset of 12,000 expert-annotated medical images with bounding boxes and structured visual CoT (SV-CoT), explicitly linking visual regions to reasoning steps. The dataset further supports 16 languages, totaling over 700k VQA pairs for broad multilingual applicability. Using S-Chain, we benchmark state-of-the-art medical VLMs (ExGra-Med, LLaVA-Med) and general-purpose VLMs (Qwen2.5-VL, InternVL2.5), showing that SV-CoT supervision significantly improves interpretability, grounding fidelity, and robustness. Beyond benchmarking, we study its synergy with retrieval-augmented generation, revealing how domain knowledge and visual grounding interact during autoregressive reasoning. Finally, we propose a new mechanism that strengthens the alignment between visual evidence and reasoning, improving both reliability and efficiency. S-Chain establishes a new benchmark for grounded medical reasoning and paves the way toward more trustworthy and explainable medical VLMs.
The Reasoning Boundary Paradox: How Reinforcement Learning Constrains Language Models
Oct 02, 2025Abstract:Reinforcement Learning with Verifiable Rewards (RLVR) has emerged as a key method for improving Large Language Models' reasoning capabilities, yet recent evidence suggests it may paradoxically shrink the reasoning boundary rather than expand it. This paper investigates the shrinkage issue of RLVR by analyzing its learning dynamics and reveals two critical phenomena that explain this failure. First, we expose negative interference in RLVR, where learning to solve certain training problems actively reduces the likelihood of correct solutions for others, leading to the decline of Pass@$k$ performance, or the probability of generating a correct solution within $k$ attempts. Second, we uncover the winner-take-all phenomenon: RLVR disproportionately reinforces problems with high likelihood, correct solutions, under the base model, while suppressing other initially low-likelihood ones. Through extensive theoretical and empirical analysis on multiple mathematical reasoning benchmarks, we show that this effect arises from the inherent on-policy sampling in standard RL objectives, causing the model to converge toward narrow solution strategies. Based on these insights, we propose a simple yet effective data curation algorithm that focuses RLVR learning on low-likelihood problems, achieving notable improvement in Pass@$k$ performance. Our code is available at https://github.com/mail-research/SELF-llm-interference.
On Zero-Initialized Attention: Optimal Prompt and Gating Factor Estimation
Feb 05, 2025



Abstract:The LLaMA-Adapter has recently emerged as an efficient fine-tuning technique for LLaMA models, leveraging zero-initialized attention to stabilize training and enhance performance. However, despite its empirical success, the theoretical foundations of zero-initialized attention remain largely unexplored. In this paper, we provide a rigorous theoretical analysis, establishing a connection between zero-initialized attention and mixture-of-expert models. We prove that both linear and non-linear prompts, along with gating functions, can be optimally estimated, with non-linear prompts offering greater flexibility for future applications. Empirically, we validate our findings on the open LLM benchmarks, demonstrating that non-linear prompts outperform linear ones. Notably, even with limited training data, both prompt types consistently surpass vanilla attention, highlighting the robustness and adaptability of zero-initialized attention.
Deep Learning for Ophthalmology: The State-of-the-Art and Future Trends
Jan 07, 2025



Abstract:The emergence of artificial intelligence (AI), particularly deep learning (DL), has marked a new era in the realm of ophthalmology, offering transformative potential for the diagnosis and treatment of posterior segment eye diseases. This review explores the cutting-edge applications of DL across a range of ocular conditions, including diabetic retinopathy, glaucoma, age-related macular degeneration, and retinal vessel segmentation. We provide a comprehensive overview of foundational ML techniques and advanced DL architectures, such as CNNs, attention mechanisms, and transformer-based models, highlighting the evolving role of AI in enhancing diagnostic accuracy, optimizing treatment strategies, and improving overall patient care. Additionally, we present key challenges in integrating AI solutions into clinical practice, including ensuring data diversity, improving algorithm transparency, and effectively leveraging multimodal data. This review emphasizes AI's potential to improve disease diagnosis and enhance patient care while stressing the importance of collaborative efforts to overcome these barriers and fully harness AI's impact in advancing eye care.
LoGra-Med: Long Context Multi-Graph Alignment for Medical Vision-Language Model
Oct 03, 2024



Abstract:State-of-the-art medical multi-modal large language models (med-MLLM), like LLaVA-Med or BioMedGPT, leverage instruction-following data in pre-training. However, those models primarily focus on scaling the model size and data volume to boost performance while mainly relying on the autoregressive learning objectives. Surprisingly, we reveal that such learning schemes might result in a weak alignment between vision and language modalities, making these models highly reliant on extensive pre-training datasets - a significant challenge in medical domains due to the expensive and time-consuming nature of curating high-quality instruction-following instances. We address this with LoGra-Med, a new multi-graph alignment algorithm that enforces triplet correlations across image modalities, conversation-based descriptions, and extended captions. This helps the model capture contextual meaning, handle linguistic variability, and build cross-modal associations between visuals and text. To scale our approach, we designed an efficient end-to-end learning scheme using black-box gradient estimation, enabling faster LLaMa 7B training. Our results show LoGra-Med matches LLAVA-Med performance on 600K image-text pairs for Medical VQA and significantly outperforms it when trained on 10% of the data. For example, on VQA-RAD, we exceed LLAVA-Med by 20.13% and nearly match the 100% pre-training score (72.52% vs. 72.64%). We also surpass SOTA methods like BiomedGPT on visual chatbots and RadFM on zero-shot image classification with VQA, highlighting the effectiveness of multi-graph alignment.
Dude: Dual Distribution-Aware Context Prompt Learning For Large Vision-Language Model
Jul 05, 2024



Abstract:Prompt learning methods are gaining increasing attention due to their ability to customize large vision-language models to new domains using pre-trained contextual knowledge and minimal training data. However, existing works typically rely on optimizing unified prompt inputs, often struggling with fine-grained classification tasks due to insufficient discriminative attributes. To tackle this, we consider a new framework based on a dual context of both domain-shared and class-specific contexts, where the latter is generated by Large Language Models (LLMs) such as GPTs. Such dual prompt methods enhance the model's feature representation by joining implicit and explicit factors encoded in LLM knowledge. Moreover, we formulate the Unbalanced Optimal Transport (UOT) theory to quantify the relationships between constructed prompts and visual tokens. Through partial matching, UOT can properly align discrete sets of visual tokens and prompt embeddings under different mass distributions, which is particularly valuable for handling irrelevant or noisy elements, ensuring that the preservation of mass does not restrict transport solutions. Furthermore, UOT's characteristics integrate seamlessly with image augmentation, expanding the training sample pool while maintaining a reasonable distance between perturbed images and prompt inputs. Extensive experiments across few-shot classification and adapter settings substantiate the superiority of our model over current state-of-the-art baselines.
I-MPN: Inductive Message Passing Network for Effective and Efficient Human-in-the-Loop Annotation of Mobile Eye Tracking Data
Jun 10, 2024



Abstract:Understanding human visual processing in dynamic environments is essential for psychology and human-centered interaction design. Mobile eye-tracking systems, combining egocentric video and gaze signals, offer valuable insights. However, manual analysis of these recordings is time-intensive. In this work, we present a novel human-centered learning algorithm designed for automated object recognition within mobile eye-tracking settings. Our approach seamlessly integrates an object detector with an inductive message-passing network technique (I-MPN), harnessing node features such as node profile information and positions. This integration enables our algorithm to learn embedding functions capable of generalizing to new object angle views, thereby facilitating rapid adaptation and efficient reasoning in dynamic contexts as users navigate through their environment. Through experiments conducted on three distinct video sequences, our \textit{interactive-based method} showcases significant performance improvements over fixed training/testing algorithms, even when trained on considerably smaller annotated samples collected through user feedback. Furthermore, we showcase exceptional efficiency in data annotation processes, surpassing approaches that use complete object detectors, combine detectors with convolutional networks, or employ interactive video segmentation.
Accelerating Transformers with Spectrum-Preserving Token Merging
May 25, 2024



Abstract:Increasing the throughput of the Transformer architecture, a foundational component used in numerous state-of-the-art models for vision and language tasks (e.g., GPT, LLaVa), is an important problem in machine learning. One recent and effective strategy is to merge token representations within Transformer models, aiming to reduce computational and memory requirements while maintaining accuracy. Prior works have proposed algorithms based on Bipartite Soft Matching (BSM), which divides tokens into distinct sets and merges the top k similar tokens. However, these methods have significant drawbacks, such as sensitivity to token-splitting strategies and damage to informative tokens in later layers. This paper presents a novel paradigm called PiToMe, which prioritizes the preservation of informative tokens using an additional metric termed the energy score. This score identifies large clusters of similar tokens as high-energy, indicating potential candidates for merging, while smaller (unique and isolated) clusters are considered as low-energy and preserved. Experimental findings demonstrate that PiToMe saved from 40-60\% FLOPs of the base models while exhibiting superior off-the-shelf performance on image classification (0.5\% average performance drop of ViT-MAE-H compared to 2.6\% as baselines), image-text retrieval (0.3\% average performance drop of CLIP on Flickr30k compared to 4.5\% as others), and analogously in visual questions answering with LLaVa-7B. Furthermore, PiToMe is theoretically shown to preserve intrinsic spectral properties of the original token space under mild conditions
Structure-Aware E(3)-Invariant Molecular Conformer Aggregation Networks
Feb 03, 2024



Abstract:A molecule's 2D representation consists of its atoms, their attributes, and the molecule's covalent bonds. A 3D (geometric) representation of a molecule is called a conformer and consists of its atom types and Cartesian coordinates. Every conformer has a potential energy, and the lower this energy, the more likely it occurs in nature. Most existing machine learning methods for molecular property prediction consider either 2D molecular graphs or 3D conformer structure representations in isolation. Inspired by recent work on using ensembles of conformers in conjunction with 2D graph representations, we propose E(3)-invariant molecular conformer aggregation networks. The method integrates a molecule's 2D representation with that of multiple of its conformers. Contrary to prior work, we propose a novel 2D--3D aggregation mechanism based on a differentiable solver for the \emph{Fused Gromov-Wasserstein Barycenter} problem and the use of an efficient online conformer generation method based on distance geometry. We show that the proposed aggregation mechanism is E(3) invariant and provides an efficient GPU implementation. Moreover, we demonstrate that the aggregation mechanism helps to outperform state-of-the-art property prediction methods on established datasets significantly.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge