Daguang Xu
Automated ischemic stroke lesion segmentation from 3D MRI
Sep 21, 2022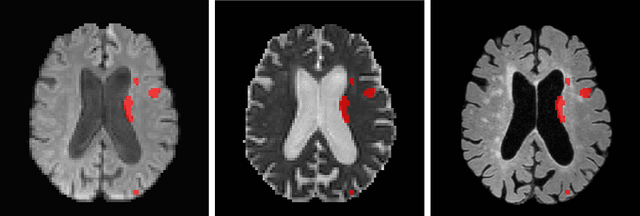
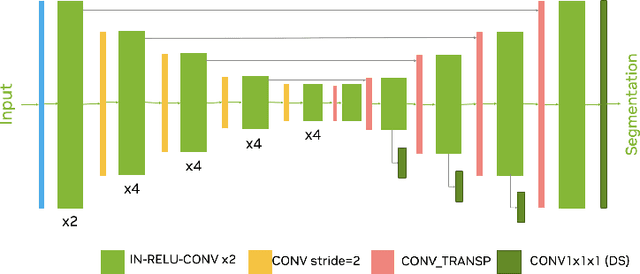
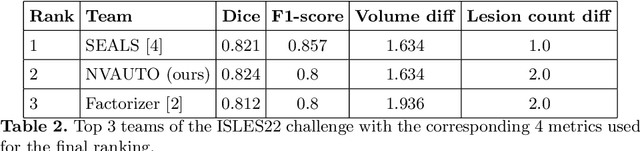
Abstract:Ischemic Stroke Lesion Segmentation challenge (ISLES 2022) offers a platform for researchers to compare their solutions to 3D segmentation of ischemic stroke regions from 3D MRIs. In this work, we describe our solution to ISLES 2022 segmentation task. We re-sample all images to a common resolution, use two input MRI modalities (DWI and ADC) and train SegResNet semantic segmentation network from MONAI. The final submission is an ensemble of 15 models (from 3 runs of 5-fold cross validation). Our solution (team name NVAUTO) achieves the top place in terms of Dice metric (0.824), and overall rank 2 (based on the combined metric ranking).
Warm Start Active Learning with Proxy Labels \& Selection via Semi-Supervised Fine-Tuning
Sep 13, 2022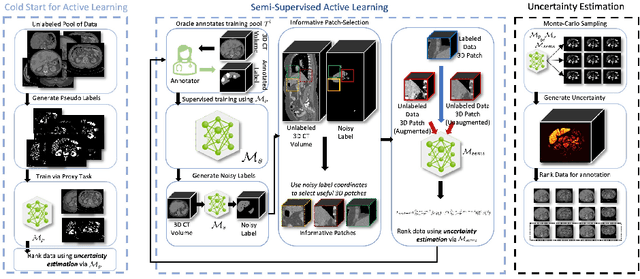
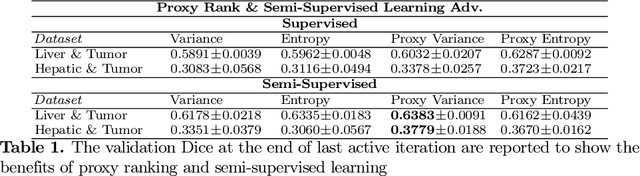
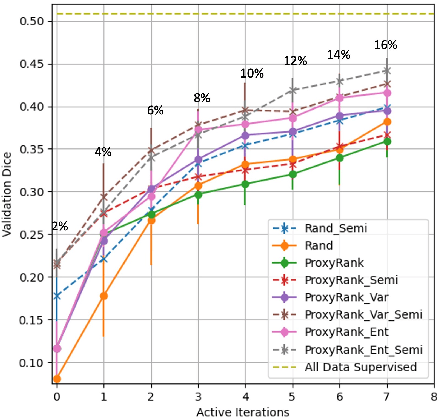
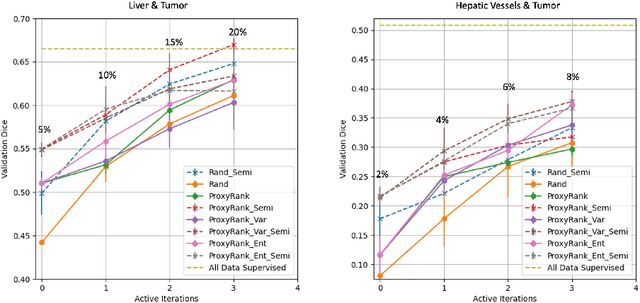
Abstract:Which volume to annotate next is a challenging problem in building medical imaging datasets for deep learning. One of the promising methods to approach this question is active learning (AL). However, AL has been a hard nut to crack in terms of which AL algorithm and acquisition functions are most useful for which datasets. Also, the problem is exacerbated with which volumes to label first when there is zero labeled data to start with. This is known as the cold start problem in AL. We propose two novel strategies for AL specifically for 3D image segmentation. First, we tackle the cold start problem by proposing a proxy task and then utilizing uncertainty generated from the proxy task to rank the unlabeled data to be annotated. Second, we craft a two-stage learning framework for each active iteration where the unlabeled data is also used in the second stage as a semi-supervised fine-tuning strategy. We show the promise of our approach on two well-known large public datasets from medical segmentation decathlon. The results indicate that the initial selection of data and semi-supervised framework both showed significant improvement for several AL strategies.
Split-U-Net: Preventing Data Leakage in Split Learning for Collaborative Multi-Modal Brain Tumor Segmentation
Aug 22, 2022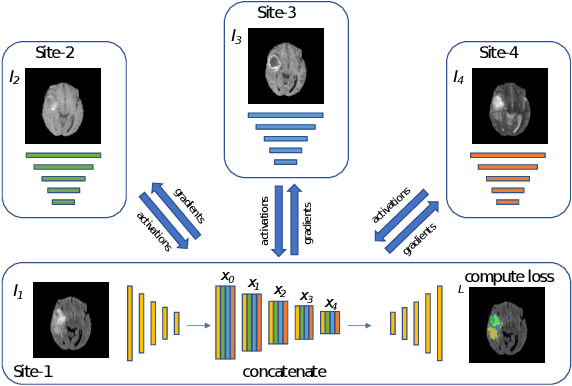
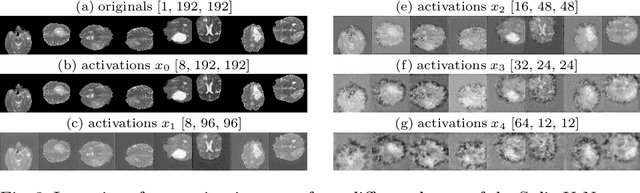

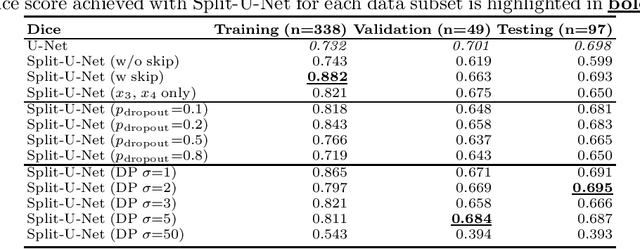
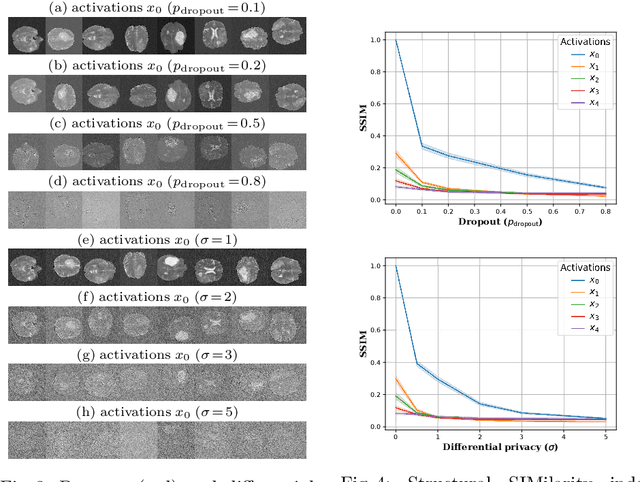
Abstract:Split learning (SL) has been proposed to train deep learning models in a decentralized manner. For decentralized healthcare applications with vertical data partitioning, SL can be beneficial as it allows institutes with complementary features or images for a shared set of patients to jointly develop more robust and generalizable models. In this work, we propose "Split-U-Net" and successfully apply SL for collaborative biomedical image segmentation. Nonetheless, SL requires the exchanging of intermediate activation maps and gradients to allow training models across different feature spaces, which might leak data and raise privacy concerns. Therefore, we also quantify the amount of data leakage in common SL scenarios for biomedical image segmentation and provide ways to counteract such leakage by applying appropriate defense strategies.
Fetal Brain Tissue Annotation and Segmentation Challenge Results
Apr 20, 2022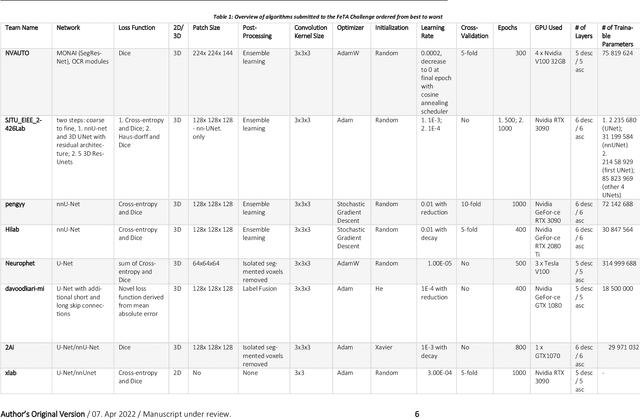
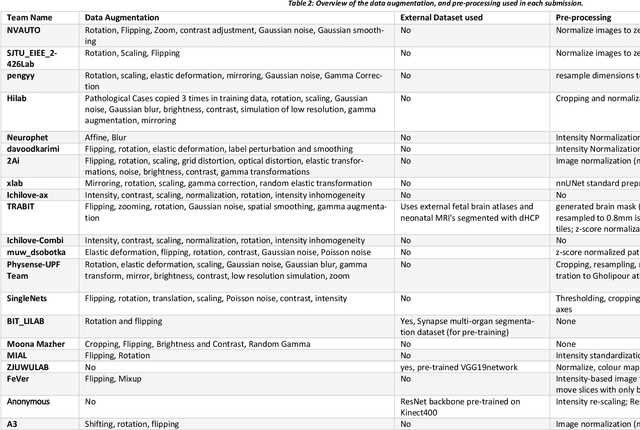
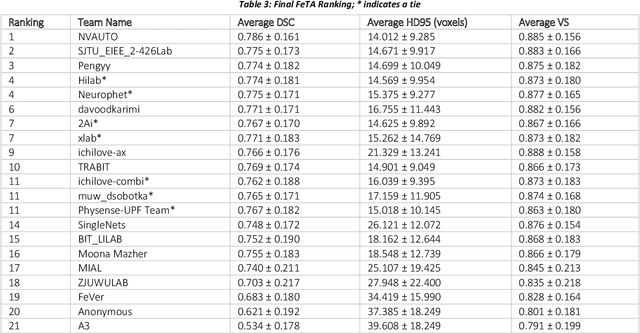
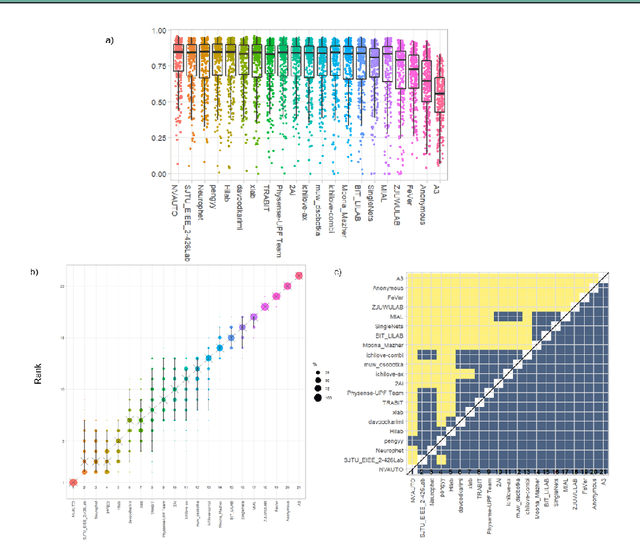
Abstract:In-utero fetal MRI is emerging as an important tool in the diagnosis and analysis of the developing human brain. Automatic segmentation of the developing fetal brain is a vital step in the quantitative analysis of prenatal neurodevelopment both in the research and clinical context. However, manual segmentation of cerebral structures is time-consuming and prone to error and inter-observer variability. Therefore, we organized the Fetal Tissue Annotation (FeTA) Challenge in 2021 in order to encourage the development of automatic segmentation algorithms on an international level. The challenge utilized FeTA Dataset, an open dataset of fetal brain MRI reconstructions segmented into seven different tissues (external cerebrospinal fluid, grey matter, white matter, ventricles, cerebellum, brainstem, deep grey matter). 20 international teams participated in this challenge, submitting a total of 21 algorithms for evaluation. In this paper, we provide a detailed analysis of the results from both a technical and clinical perspective. All participants relied on deep learning methods, mainly U-Nets, with some variability present in the network architecture, optimization, and image pre- and post-processing. The majority of teams used existing medical imaging deep learning frameworks. The main differences between the submissions were the fine tuning done during training, and the specific pre- and post-processing steps performed. The challenge results showed that almost all submissions performed similarly. Four of the top five teams used ensemble learning methods. However, one team's algorithm performed significantly superior to the other submissions, and consisted of an asymmetrical U-Net network architecture. This paper provides a first of its kind benchmark for future automatic multi-tissue segmentation algorithms for the developing human brain in utero.
UNetFormer: A Unified Vision Transformer Model and Pre-Training Framework for 3D Medical Image Segmentation
Apr 05, 2022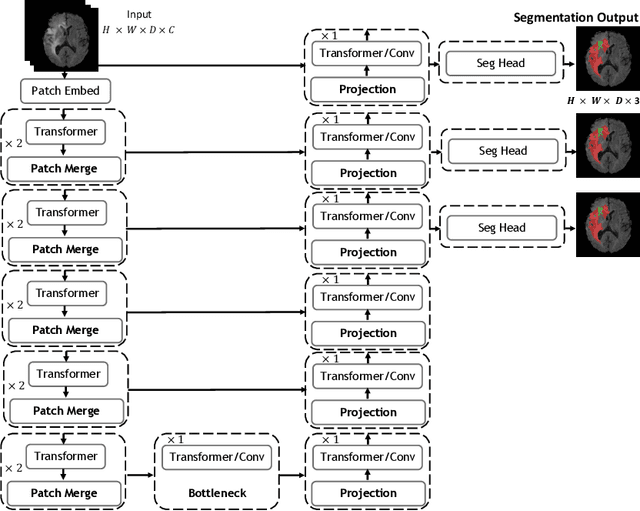
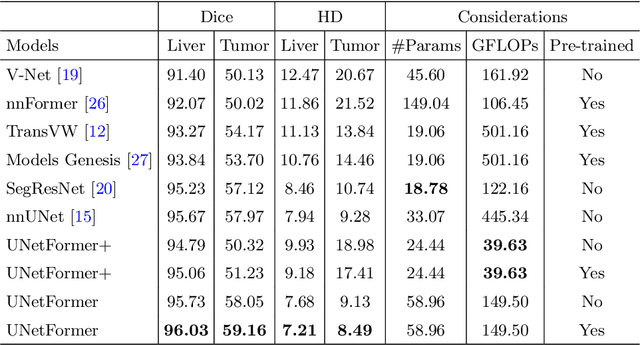
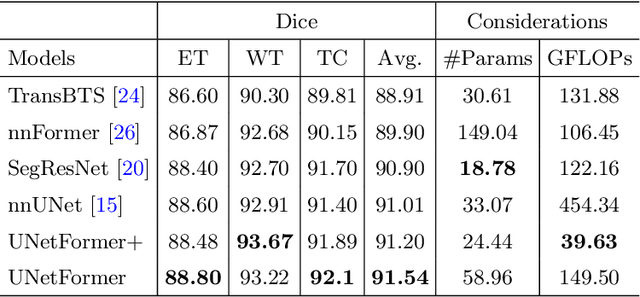
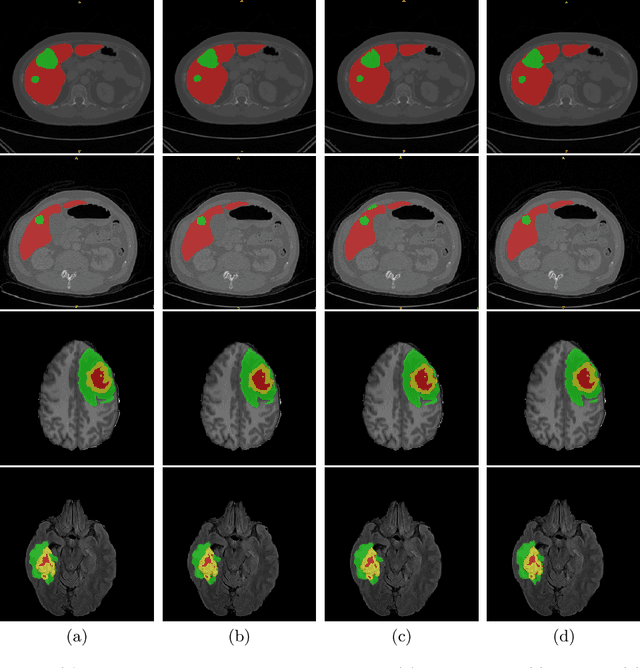
Abstract:Vision Transformers (ViT)s have recently become popular due to their outstanding modeling capabilities, in particular for capturing long-range information, and scalability to dataset and model sizes which has led to state-of-the-art performance in various computer vision and medical image analysis tasks. In this work, we introduce a unified framework consisting of two architectures, dubbed UNetFormer, with a 3D Swin Transformer-based encoder and Convolutional Neural Network (CNN) and transformer-based decoders. In the proposed model, the encoder is linked to the decoder via skip connections at five different resolutions with deep supervision. The design of proposed architecture allows for meeting a wide range of trade-off requirements between accuracy and computational cost. In addition, we present a methodology for self-supervised pre-training of the encoder backbone via learning to predict randomly masked volumetric tokens using contextual information of visible tokens. We pre-train our framework on a cohort of $5050$ CT images, gathered from publicly available CT datasets, and present a systematic investigation of various components such as masking ratio and patch size that affect the representation learning capability and performance of downstream tasks. We validate the effectiveness of our pre-training approach by fine-tuning and testing our model on liver and liver tumor segmentation task using the Medical Segmentation Decathlon (MSD) dataset and achieve state-of-the-art performance in terms of various segmentation metrics. To demonstrate its generalizability, we train and test the model on BraTS 21 dataset for brain tumor segmentation using MRI images and outperform other methods in terms of Dice score. Code: https://github.com/Project-MONAI/research-contributions
GradViT: Gradient Inversion of Vision Transformers
Mar 28, 2022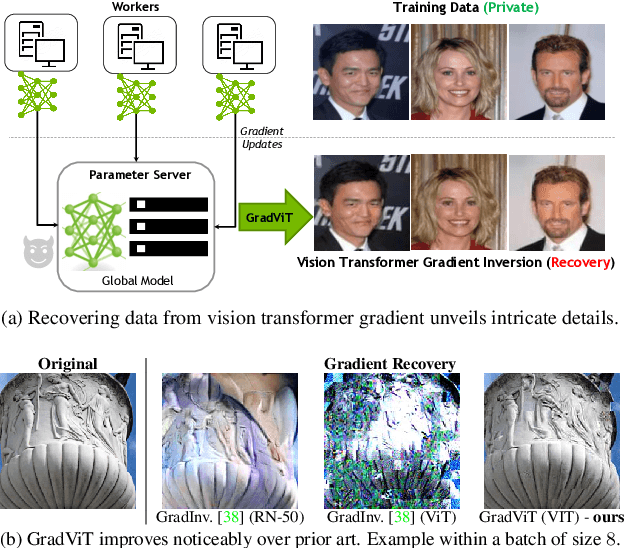
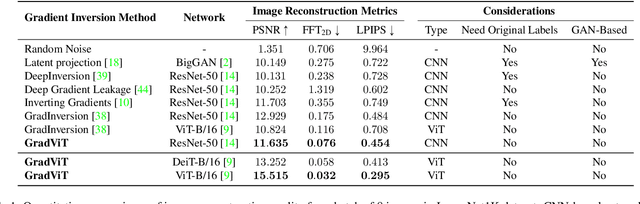
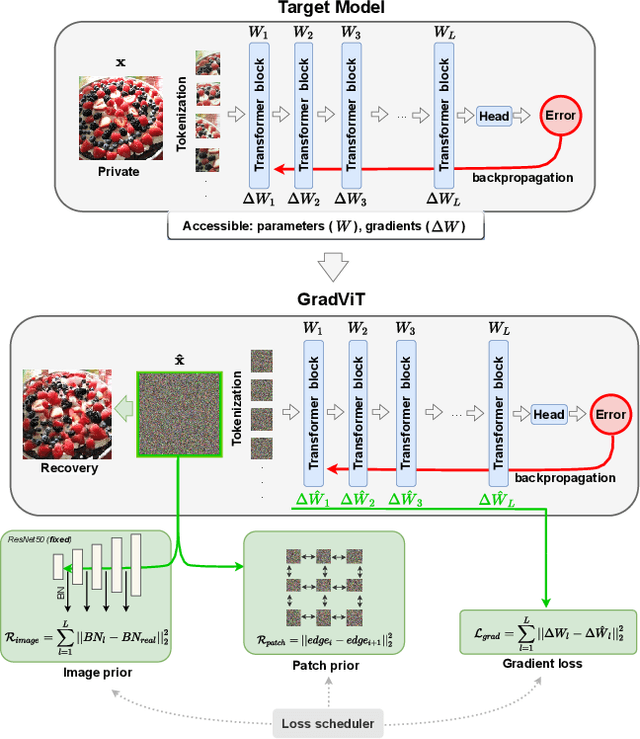
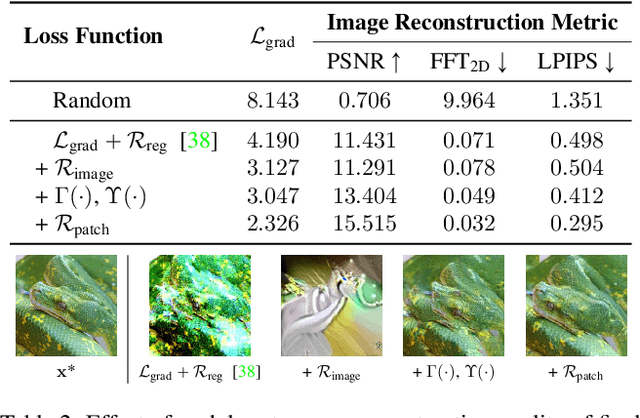
Abstract:In this work we demonstrate the vulnerability of vision transformers (ViTs) to gradient-based inversion attacks. During this attack, the original data batch is reconstructed given model weights and the corresponding gradients. We introduce a method, named GradViT, that optimizes random noise into naturally looking images via an iterative process. The optimization objective consists of (i) a loss on matching the gradients, (ii) image prior in the form of distance to batch-normalization statistics of a pretrained CNN model, and (iii) a total variation regularization on patches to guide correct recovery locations. We propose a unique loss scheduling function to overcome local minima during optimization. We evaluate GadViT on ImageNet1K and MS-Celeb-1M datasets, and observe unprecedentedly high fidelity and closeness to the original (hidden) data. During the analysis we find that vision transformers are significantly more vulnerable than previously studied CNNs due to the presence of the attention mechanism. Our method demonstrates new state-of-the-art results for gradient inversion in both qualitative and quantitative metrics. Project page at https://gradvit.github.io/.
MONAI Label: A framework for AI-assisted Interactive Labeling of 3D Medical Images
Mar 23, 2022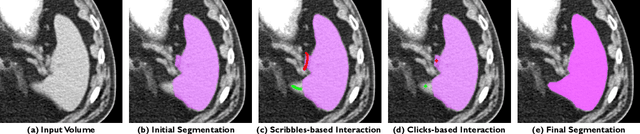
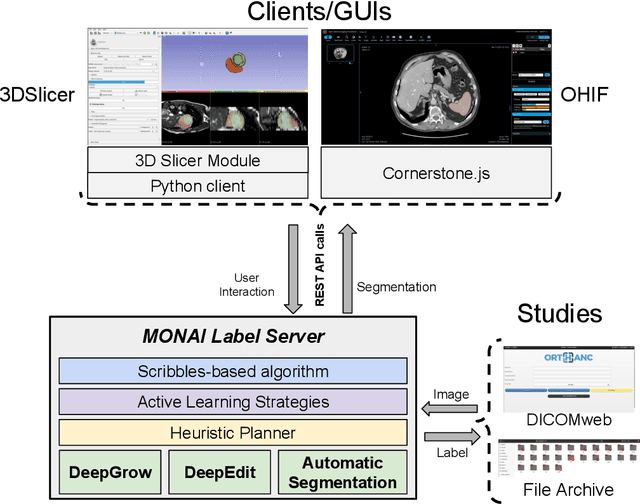
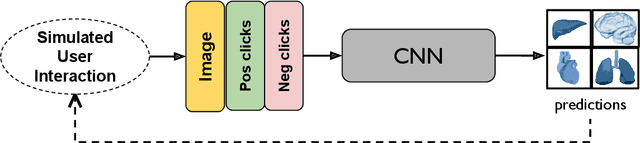
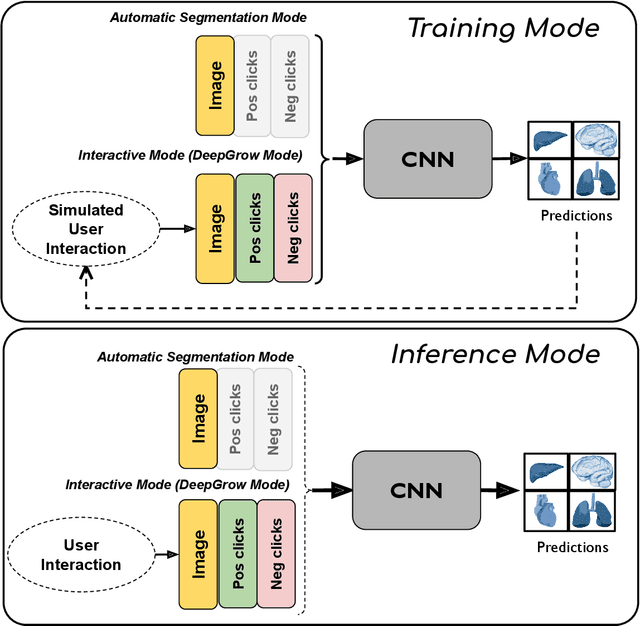
Abstract:The lack of annotated datasets is a major challenge in training new task-specific supervised AI algorithms as manual annotation is expensive and time-consuming. To address this problem, we present MONAI Label, a free and open-source platform that facilitates the development of AI-based applications that aim at reducing the time required to annotate 3D medical image datasets. Through MONAI Label researchers can develop annotation applications focusing on their domain of expertise. It allows researchers to readily deploy their apps as services, which can be made available to clinicians via their preferred user-interface. Currently, MONAI Label readily supports locally installed (3DSlicer) and web-based (OHIF) frontends, and offers two Active learning strategies to facilitate and speed up the training of segmentation algorithms. MONAI Label allows researchers to make incremental improvements to their labeling apps by making them available to other researchers and clinicians alike. Lastly, MONAI Label provides sample labeling apps, namely DeepEdit and DeepGrow, demonstrating dramatically reduced annotation times.
Closing the Generalization Gap of Cross-silo Federated Medical Image Segmentation
Mar 18, 2022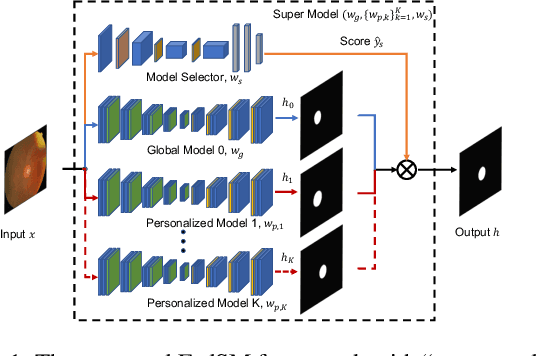
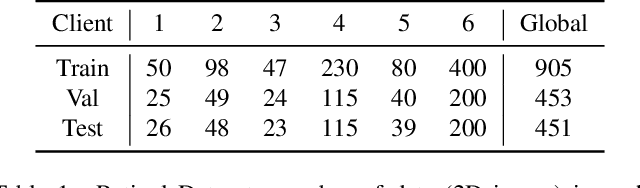
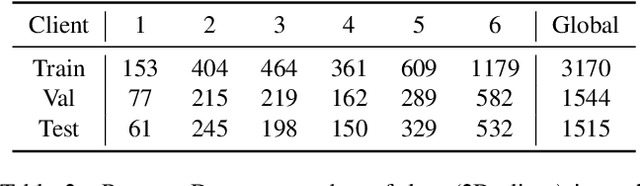
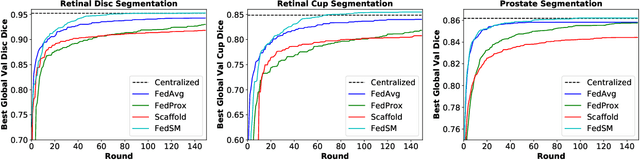
Abstract:Cross-silo federated learning (FL) has attracted much attention in medical imaging analysis with deep learning in recent years as it can resolve the critical issues of insufficient data, data privacy, and training efficiency. However, there can be a generalization gap between the model trained from FL and the one from centralized training. This important issue comes from the non-iid data distribution of the local data in the participating clients and is well-known as client drift. In this work, we propose a novel training framework FedSM to avoid the client drift issue and successfully close the generalization gap compared with the centralized training for medical image segmentation tasks for the first time. We also propose a novel personalized FL objective formulation and a new method SoftPull to solve it in our proposed framework FedSM. We conduct rigorous theoretical analysis to guarantee its convergence for optimizing the non-convex smooth objective function. Real-world medical image segmentation experiments using deep FL validate the motivations and effectiveness of our proposed method.
Auto-FedRL: Federated Hyperparameter Optimization for Multi-institutional Medical Image Segmentation
Mar 12, 2022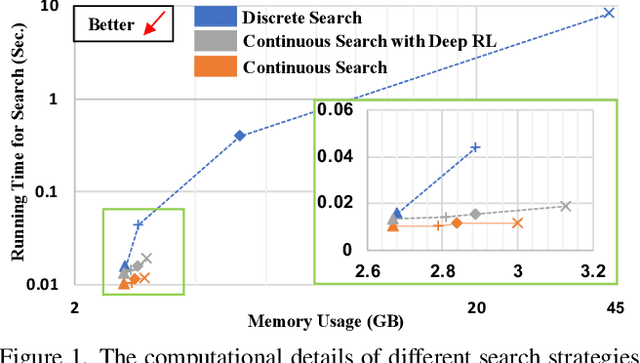
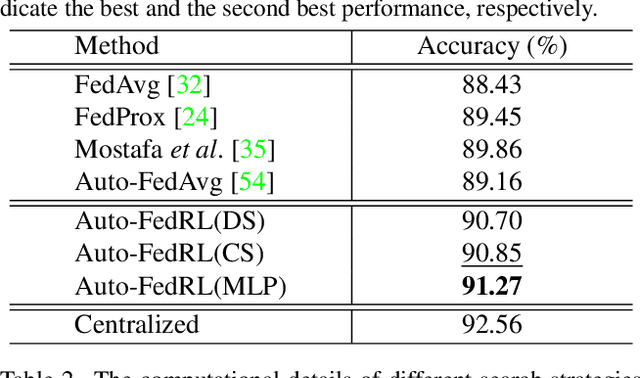

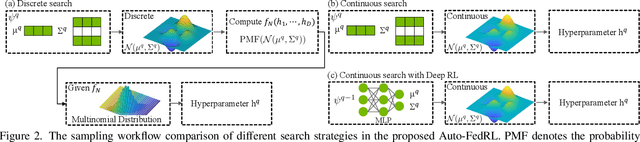

Abstract:Federated learning (FL) is a distributed machine learning technique that enables collaborative model training while avoiding explicit data sharing. The inherent privacy-preserving property of FL algorithms makes them especially attractive to the medical field. However, in case of heterogeneous client data distributions, standard FL methods are unstable and require intensive hyperparameter tuning to achieve optimal performance. Conventional hyperparameter optimization algorithms are impractical in real-world FL applications as they involve numerous training trials, which are often not affordable with limited compute budgets. In this work, we propose an efficient reinforcement learning~(RL)-based federated hyperparameter optimization algorithm, termed Auto-FedRL, in which an online RL agent can dynamically adjust hyperparameters of each client based on the current training progress. Extensive experiments are conducted to investigate different search strategies and RL agents. The effectiveness of the proposed method is validated on a heterogeneous data split of the CIFAR-10 dataset as well as two real-world medical image segmentation datasets for COVID-19 lesion segmentation in chest CT and pancreas segmentation in abdominal CT.
Do Gradient Inversion Attacks Make Federated Learning Unsafe?
Feb 14, 2022



Abstract:Federated learning (FL) allows the collaborative training of AI models without needing to share raw data. This capability makes it especially interesting for healthcare applications where patient and data privacy is of utmost concern. However, recent works on the inversion of deep neural networks from model gradients raised concerns about the security of FL in preventing the leakage of training data. In this work, we show that these attacks presented in the literature are impractical in real FL use-cases and provide a new baseline attack that works for more realistic scenarios where the clients' training involves updating the Batch Normalization (BN) statistics. Furthermore, we present new ways to measure and visualize potential data leakage in FL. Our work is a step towards establishing reproducible methods of measuring data leakage in FL and could help determine the optimal tradeoffs between privacy-preserving techniques, such as differential privacy, and model accuracy based on quantifiable metrics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge