Interpretable Multi-Step Reasoning with Knowledge Extraction on Complex Healthcare Question Answering
Aug 06, 2020Ye Liu, Shaika Chowdhury, Chenwei Zhang, Cornelia Caragea, Philip S. Yu
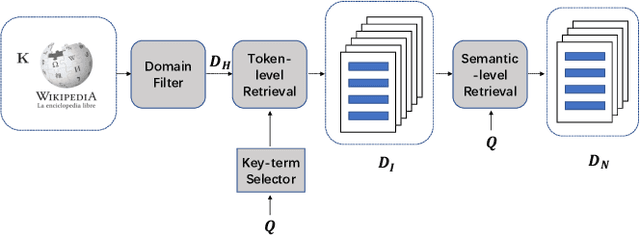
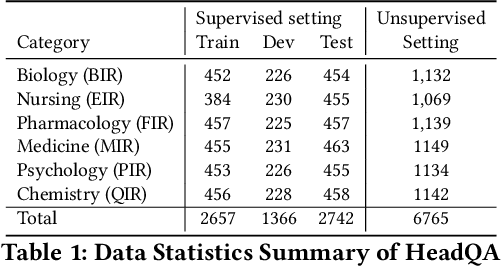
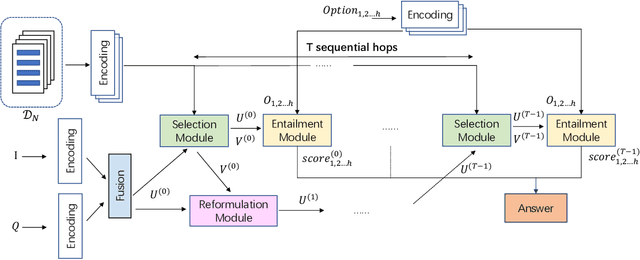
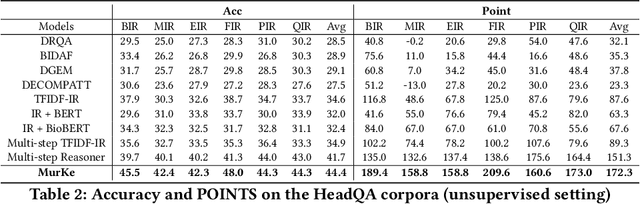
Healthcare question answering assistance aims to provide customer healthcare information, which widely appears in both Web and mobile Internet. The questions usually require the assistance to have proficient healthcare background knowledge as well as the reasoning ability on the knowledge. Recently a challenge involving complex healthcare reasoning, HeadQA dataset, has been proposed, which contains multiple-choice questions authorized for the public healthcare specialization exam. Unlike most other QA tasks that focus on linguistic understanding, HeadQA requires deeper reasoning involving not only knowledge extraction, but also complex reasoning with healthcare knowledge. These questions are the most challenging for current QA systems, and the current performance of the state-of-the-art method is slightly better than a random guess. In order to solve this challenging task, we present a Multi-step reasoning with Knowledge extraction framework (MurKe). The proposed framework first extracts the healthcare knowledge as supporting documents from the large corpus. In order to find the reasoning chain and choose the correct answer, MurKe iterates between selecting the supporting documents, reformulating the query representation using the supporting documents and getting entailment score for each choice using the entailment model. The reformulation module leverages selected documents for missing evidence, which maintains interpretability. Moreover, we are striving to make full use of off-the-shelf pre-trained models. With less trainable weight, the pre-trained model can easily adapt to healthcare tasks with limited training samples. From the experimental results and ablation study, our system is able to outperform several strong baselines on the HeadQA dataset.
AutoKnow: Self-Driving Knowledge Collection for Products of Thousands of Types
Jun 24, 2020Xin Luna Dong, Xiang He, Andrey Kan, Xian Li, Yan Liang, Jun Ma, Yifan Ethan Xu, Chenwei Zhang, Tong Zhao, Gabriel Blanco Saldana, Saurabh Deshpande, Alexandre Michetti Manduca, Jay Ren, Surender Pal Singh, Fan Xiao, Haw-Shiuan Chang, Giannis Karamanolakis, Yuning Mao, Yaqing Wang, Christos Faloutsos, Andrew McCallum, Jiawei Han
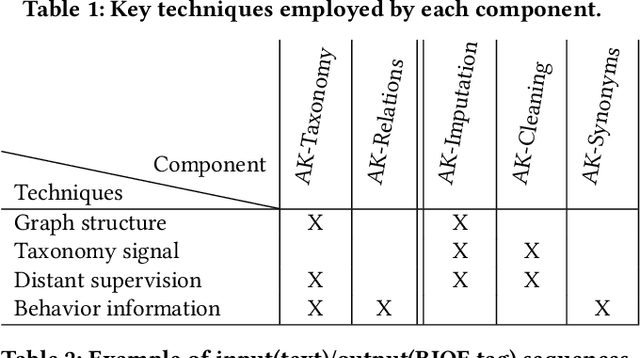
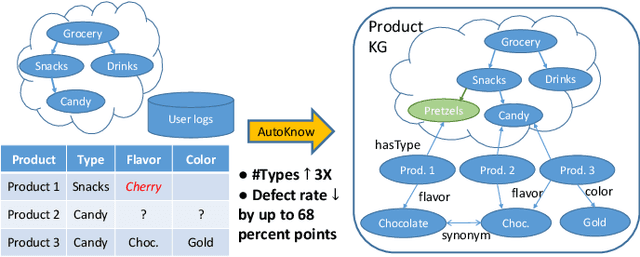

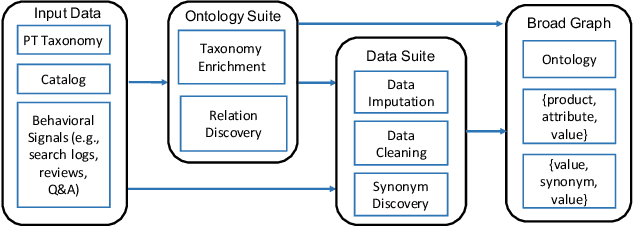
Can one build a knowledge graph (KG) for all products in the world? Knowledge graphs have firmly established themselves as valuable sources of information for search and question answering, and it is natural to wonder if a KG can contain information about products offered at online retail sites. There have been several successful examples of generic KGs, but organizing information about products poses many additional challenges, including sparsity and noise of structured data for products, complexity of the domain with millions of product types and thousands of attributes, heterogeneity across large number of categories, as well as large and constantly growing number of products. We describe AutoKnow, our automatic (self-driving) system that addresses these challenges. The system includes a suite of novel techniques for taxonomy construction, product property identification, knowledge extraction, anomaly detection, and synonym discovery. AutoKnow is (a) automatic, requiring little human intervention, (b) multi-scalable, scalable in multiple dimensions (many domains, many products, and many attributes), and (c) integrative, exploiting rich customer behavior logs. AutoKnow has been operational in collecting product knowledge for over 11K product types.
Octet: Online Catalog Taxonomy Enrichment with Self-Supervision
Jun 18, 2020Yuning Mao, Tong Zhao, Andrey Kan, Chenwei Zhang, Xin Luna Dong, Christos Faloutsos, Jiawei Han
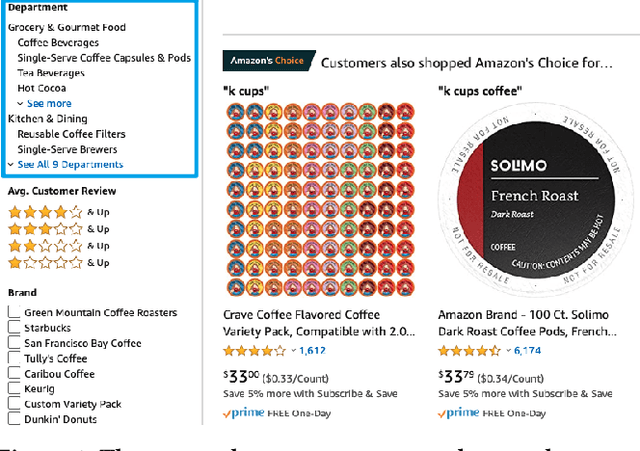
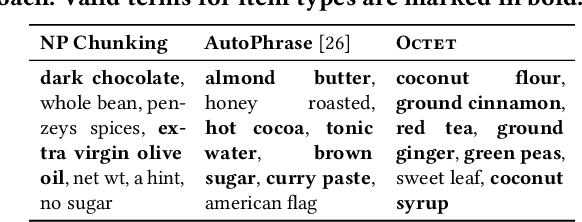
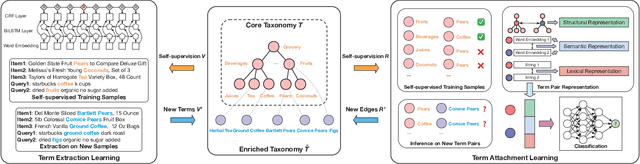
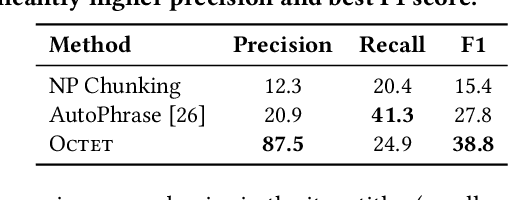
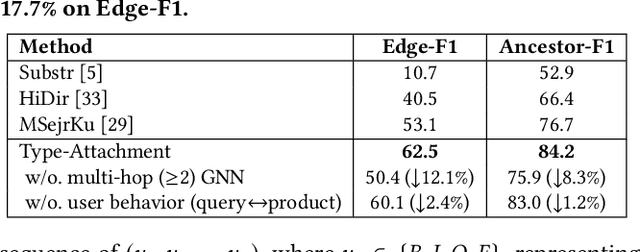
Taxonomies have found wide applications in various domains, especially online for item categorization, browsing, and search. Despite the prevalent use of online catalog taxonomies, most of them in practice are maintained by humans, which is labor-intensive and difficult to scale. While taxonomy construction from scratch is considerably studied in the literature, how to effectively enrich existing incomplete taxonomies remains an open yet important research question. Taxonomy enrichment not only requires the robustness to deal with emerging terms but also the consistency between existing taxonomy structure and new term attachment. In this paper, we present a self-supervised end-to-end framework, Octet, for Online Catalog Taxonomy EnrichmenT. Octet leverages heterogeneous information unique to online catalog taxonomies such as user queries, items, and their relations to the taxonomy nodes while requiring no other supervision than the existing taxonomies. We propose to distantly train a sequence labeling model for term extraction and employ graph neural networks (GNNs) to capture the taxonomy structure as well as the query-item-taxonomy interactions for term attachment. Extensive experiments in different online domains demonstrate the superiority of Octet over state-of-the-art methods via both automatic and human evaluations. Notably, Octet enriches an online catalog taxonomy in production to 2 times larger in the open-world evaluation.
SelfORE: Self-supervised Relational Feature Learning for Open Relation Extraction
Apr 06, 2020Xuming Hu, Lijie Wen, Yusong Xu, Chenwei Zhang, Philip S. Yu
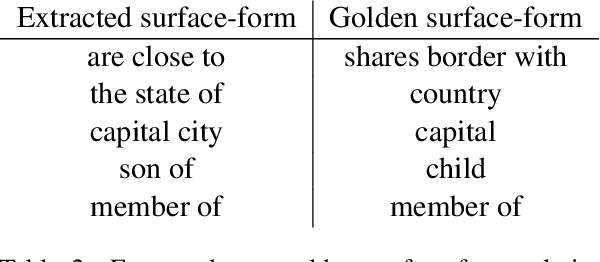
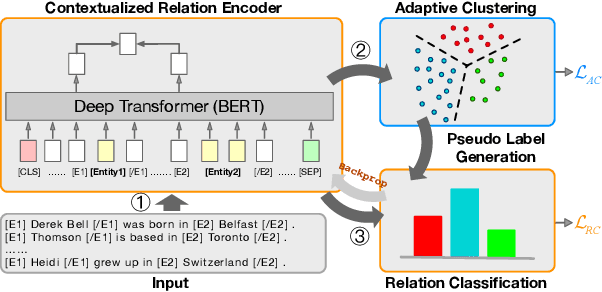
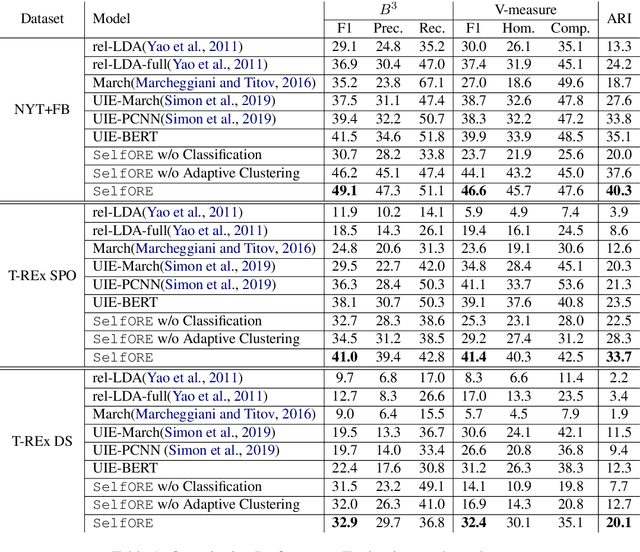
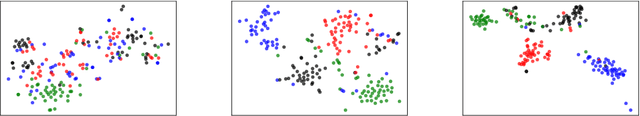

Open relation extraction is the task of extracting open-domain relation facts from natural language sentences. Existing works either utilize heuristics or distant-supervised annotations to train a supervised classifier over pre-defined relations, or adopt unsupervised methods with additional assumptions that have less discriminative power. In this work, we proposed a self-supervised framework named SelfORE, which exploits weak, self-supervised signals by leveraging large pretrained language model for adaptive clustering on contextualized relational features, and bootstraps the self-supervised signals by improving contextualized features in relation classification. Experimental results on three datasets show the effectiveness and robustness of SelfORE on open-domain Relation Extraction when comparing with competitive baselines. Source code is available at https://github.com/THU-BPM/SelfORE.
CG-BERT: Conditional Text Generation with BERT for Generalized Few-shot Intent Detection
Apr 04, 2020Congying Xia, Chenwei Zhang, Hoang Nguyen, Jiawei Zhang, Philip Yu
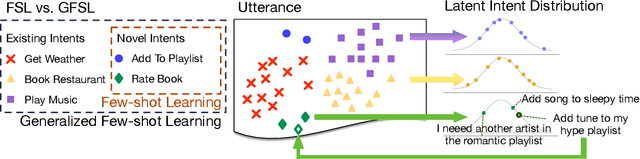
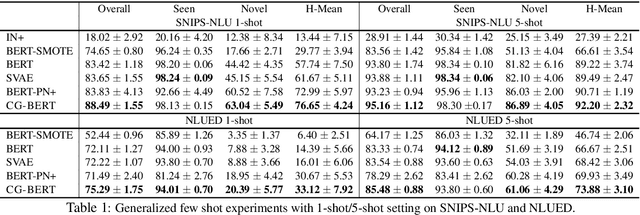
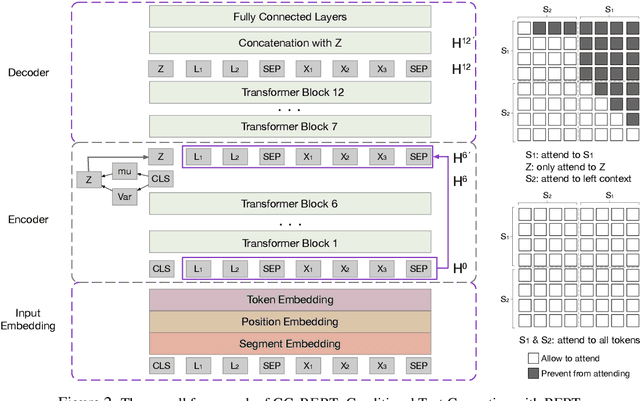
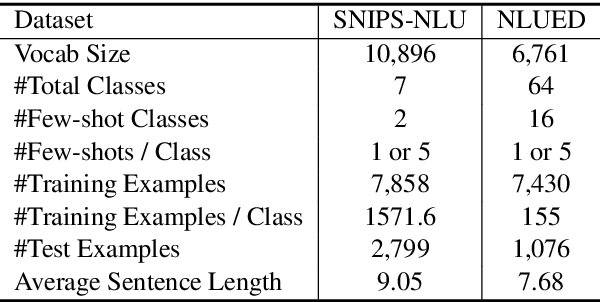
In this paper, we formulate a more realistic and difficult problem setup for the intent detection task in natural language understanding, namely Generalized Few-Shot Intent Detection (GFSID). GFSID aims to discriminate a joint label space consisting of both existing intents which have enough labeled data and novel intents which only have a few examples for each class. To approach this problem, we propose a novel model, Conditional Text Generation with BERT (CG-BERT). CG-BERT effectively leverages a large pre-trained language model to generate text conditioned on the intent label. By modeling the utterance distribution with variational inference, CG-BERT can generate diverse utterances for the novel intents even with only a few utterances available. Experimental results show that CG-BERT achieves state-of-the-art performance on the GFSID task with 1-shot and 5-shot settings on two real-world datasets.
Attribute2vec: Deep Network Embedding Through Multi-Filtering GCN
Apr 03, 2020Tingyi Wanyan, Chenwei Zhang, Ariful Azad, Xiaomin Liang, Daifeng Li, Ying Ding
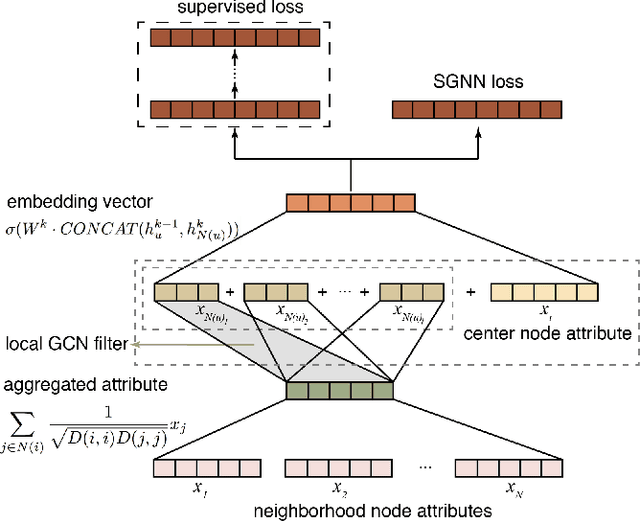
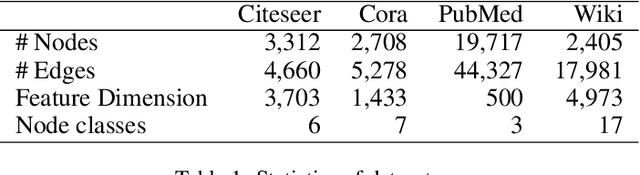
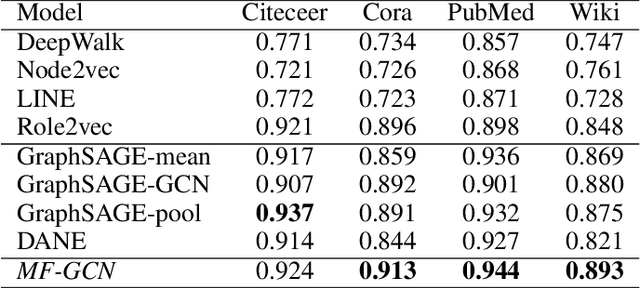
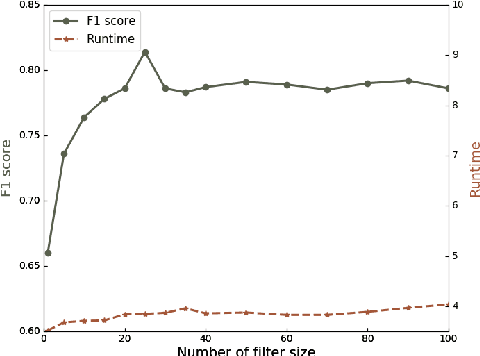
We present a multi-filtering Graph Convolution Neural Network (GCN) framework for network embedding task. It uses multiple local GCN filters to do feature extraction in every propagation layer. We show this approach could capture different important aspects of node features against the existing attribute embedding based method. We also show that with multi-filtering GCN approach, we can achieve significant improvement against baseline methods when training data is limited. We also perform many empirical experiments and demonstrate the benefit of using multiple filters against single filter as well as most current existing network embedding methods for both the link prediction and node classification tasks.
Med2Meta: Learning Representations of Medical Concepts with Meta-Embeddings
Jan 04, 2020Shaika Chowdhury, Chenwei Zhang, Philip S. Yu, Yuan Luo
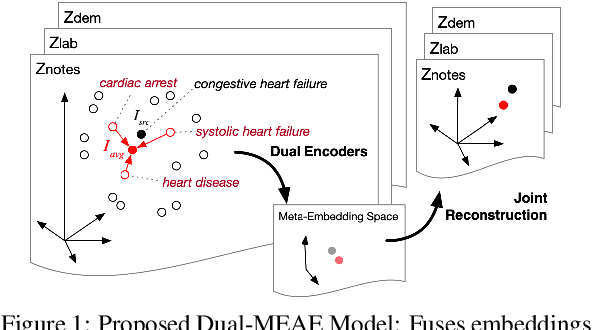
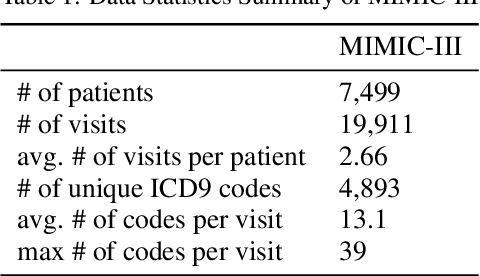
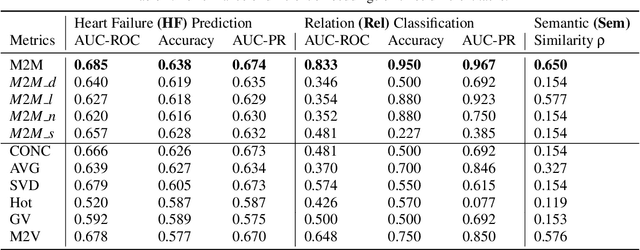

Distributed representations of medical concepts have been used to support downstream clinical tasks recently. Electronic Health Records (EHR) capture different aspects of patients' hospital encounters and serve as a rich source for augmenting clinical decision making by learning robust medical concept embeddings. However, the same medical concept can be recorded in different modalities (e.g., clinical notes, lab results)-with each capturing salient information unique to that modality-and a holistic representation calls for relevant feature ensemble from all information sources. We hypothesize that representations learned from heterogeneous data types would lead to performance enhancement on various clinical informatics and predictive modeling tasks. To this end, our proposed approach makes use of meta-embeddings, embeddings aggregated from learned embeddings. Firstly, modality-specific embeddings for each medical concept is learned with graph autoencoders. The ensemble of all the embeddings is then modeled as a meta-embedding learning problem to incorporate their correlating and complementary information through a joint reconstruction. Empirical results of our model on both quantitative and qualitative clinical evaluations have shown improvements over state-of-the-art embedding models, thus validating our hypothesis.
* 9 pages
Generative Temporal Link Prediction via Self-tokenized Sequence Modeling
Nov 26, 2019Yue Wang, Chenwei Zhang, Shen Wang, Philip S. Yu, Lu Bai, Lixin Cui, Guandong Xu
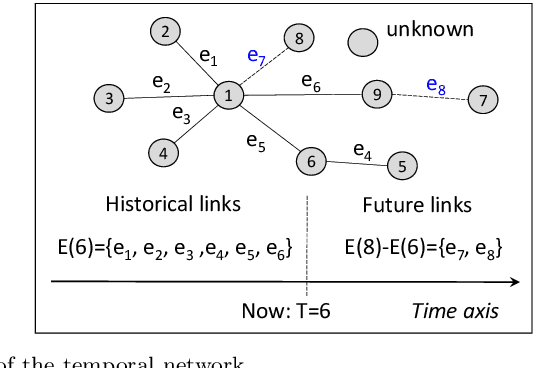
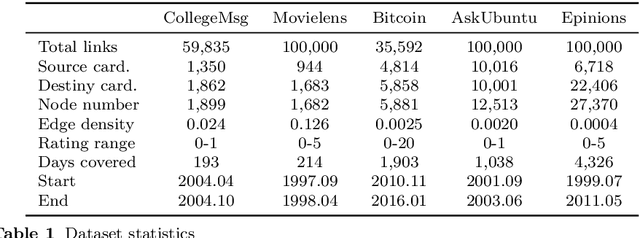
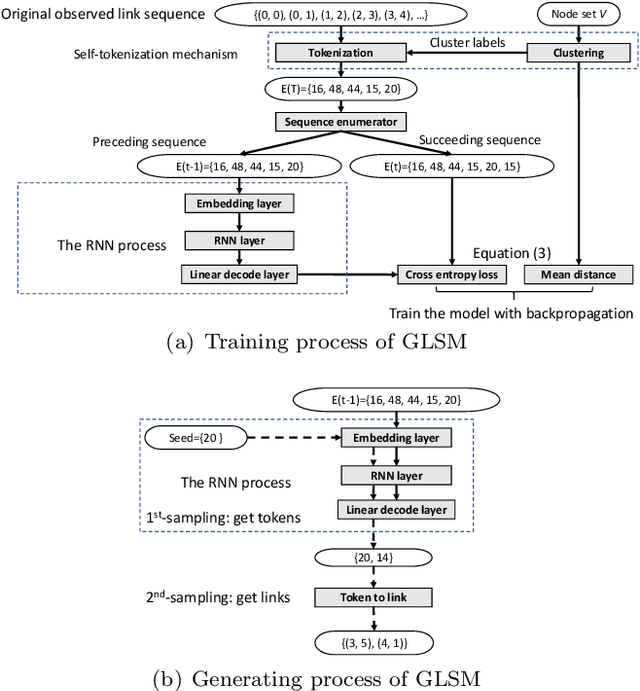
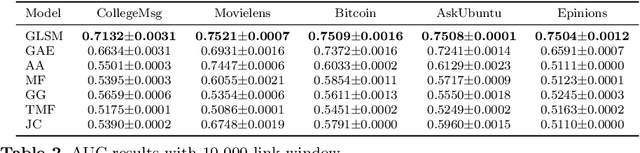
We formalize networks with evolving structures as temporal networks and propose a generative link prediction model, Generative Link Sequence Modeling (GLSM), to predict future links for temporal networks. GLSM captures the temporal link formation patterns from the observed links with a sequence modeling framework and has the ability to generate the emerging links by inferring from the probability distribution on the potential future links. To avoid overfitting caused by treating each link as a unique token, we propose a self-tokenization mechanism to transform each raw link in the network to an abstract aggregation token automatically. The self-tokenization is seamlessly integrated into the sequence modeling framework, which allows the proposed GLSM model to have the generalization capability to discover link formation patterns beyond raw link sequences. We compare GLSM with the existing state-of-art methods on five real-world datasets. The experimental results demonstrate that GLSM obtains future positive links effectively in a generative fashion while achieving the best performance (2-10\% improvements on AUC) among other alternatives.
Hierarchical Semantic Correspondence Learning for Post-Discharge Patient Mortality Prediction
Oct 15, 2019Shaika Chowdhury, Chenwei Zhang, Philip S. Yu, Yuan Luo
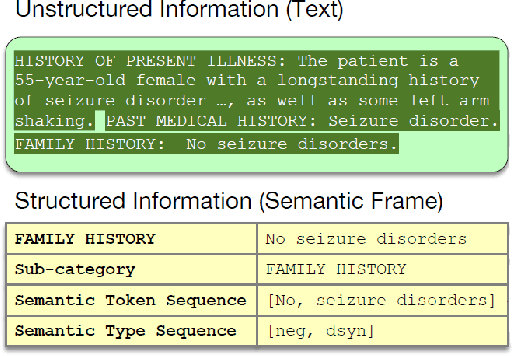

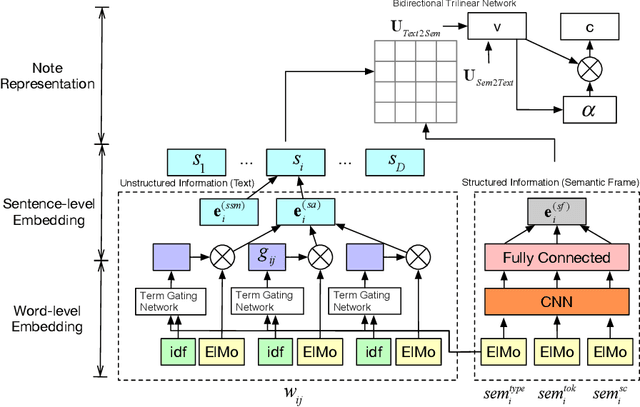
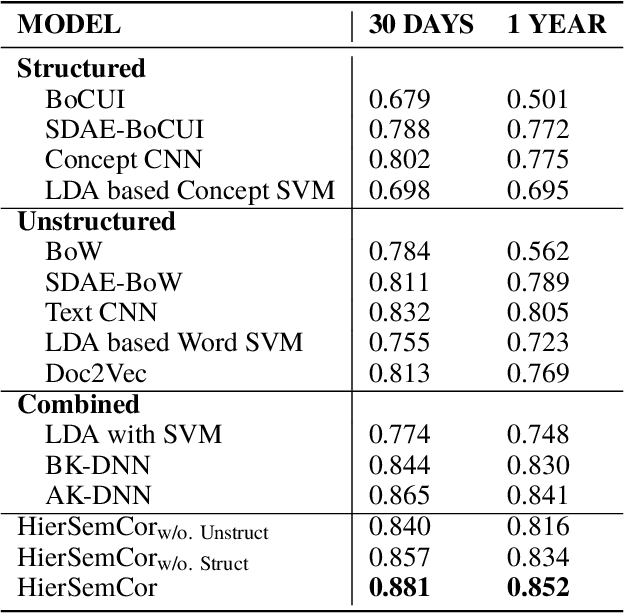
Predicting patient mortality is an important and challenging problem in the healthcare domain, especially for intensive care unit (ICU) patients. Electronic health notes serve as a rich source for learning patient representations, that can facilitate effective risk assessment. However, a large portion of clinical notes are unstructured and also contain domain specific terminologies, from which we need to extract structured information. In this paper, we introduce an embedding framework to learn semantically-plausible distributed representations of clinical notes that exploits the semantic correspondence between the unstructured texts and their corresponding structured knowledge, known as semantic frame, in a hierarchical fashion. Our approach integrates text modeling and semantic correspondence learning into a single model that comprises 1) an unstructured embedding module that makes use of self-similarity matrix representations in order to inject structural regularities of different segments inherent in clinical texts to promote local coherence, 2) a structured embedding module to embed the semantic frames (e.g., UMLS semantic types) with deep ConvNet and 3) a hierarchical semantic correspondence module that embeds by enhancing the interactions between text-semantic frame embedding pairs at multiple levels (i.e., words, sentence, note). Evaluations on multiple embedding benchmarks on post discharge intensive care patient mortality prediction tasks demonstrate its effectiveness compared to approaches that do not exploit the semantic interactions between structured and unstructured information present in clinical notes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge