Carola-Bibiane Schönlieb
on behalf of the AIX-COVNET collaboration
Can Physics-Informed Neural Networks beat the Finite Element Method?
Feb 08, 2023



Abstract:Partial differential equations play a fundamental role in the mathematical modelling of many processes and systems in physical, biological and other sciences. To simulate such processes and systems, the solutions of PDEs often need to be approximated numerically. The finite element method, for instance, is a usual standard methodology to do so. The recent success of deep neural networks at various approximation tasks has motivated their use in the numerical solution of PDEs. These so-called physics-informed neural networks and their variants have shown to be able to successfully approximate a large range of partial differential equations. So far, physics-informed neural networks and the finite element method have mainly been studied in isolation of each other. In this work, we compare the methodologies in a systematic computational study. Indeed, we employ both methods to numerically solve various linear and nonlinear partial differential equations: Poisson in 1D, 2D, and 3D, Allen-Cahn in 1D, semilinear Schr\"odinger in 1D and 2D. We then compare computational costs and approximation accuracies. In terms of solution time and accuracy, physics-informed neural networks have not been able to outperform the finite element method in our study. In some experiments, they were faster at evaluating the solved PDE.
Continuous U-Net: Faster, Greater and Noiseless
Feb 01, 2023



Abstract:Image segmentation is a fundamental task in image analysis and clinical practice. The current state-of-the-art techniques are based on U-shape type encoder-decoder networks with skip connections, called U-Net. Despite the powerful performance reported by existing U-Net type networks, they suffer from several major limitations. Issues include the hard coding of the receptive field size, compromising the performance and computational cost, as well as the fact that they do not account for inherent noise in the data. They have problems associated with discrete layers, and do not offer any theoretical underpinning. In this work we introduce continuous U-Net, a novel family of networks for image segmentation. Firstly, continuous U-Net is a continuous deep neural network that introduces new dynamic blocks modelled by second order ordinary differential equations. Secondly, we provide theoretical guarantees for our network demonstrating faster convergence, higher robustness and less sensitivity to noise. Thirdly, we derive qualitative measures to tailor-made segmentation tasks. We demonstrate, through extensive numerical and visual results, that our model outperforms existing U-Net blocks for several medical image segmentation benchmarking datasets.
Your diffusion model secretly knows the dimension of the data manifold
Dec 23, 2022



Abstract:In this work, we propose a novel framework for estimating the dimension of the data manifold using a trained diffusion model. A trained diffusion model approximates the gradient of the log density of a noise-corrupted version of the target distribution for varying levels of corruption. If the data concentrates around a manifold embedded in the high-dimensional ambient space, then as the level of corruption decreases, the score function points towards the manifold, as this direction becomes the direction of maximum likelihood increase. Therefore, for small levels of corruption, the diffusion model provides us with access to an approximation of the normal bundle of the data manifold. This allows us to estimate the dimension of the tangent space, thus, the intrinsic dimension of the data manifold. Our method outperforms linear methods for dimensionality detection such as PPCA in controlled experiments.
SCOTCH and SODA: A Transformer Video Shadow Detection Framework
Nov 13, 2022



Abstract:Shadows in videos are difficult to detect because of the large shadow deformation between frames. In this work, we argue that accounting for the shadow deformation is essential when designing a video shadow detection method. To this end, we introduce the shadow deformation attention trajectory (SODA), a new type of video self-attention module, specially designed to handle the large shadow deformations in videos. Moreover, we present a shadow contrastive learning mechanism (SCOTCH) which aims at guiding the network to learn a high-level representation of shadows, unified across different videos. We demonstrate empirically the effectiveness of our two contributions in an ablation study. Furthermore, we show that SCOTCH and SODA significantly outperforms existing techniques for video shadow detection. Code will be available upon the acceptance of this work.
Dynamical systems' based neural networks
Oct 05, 2022
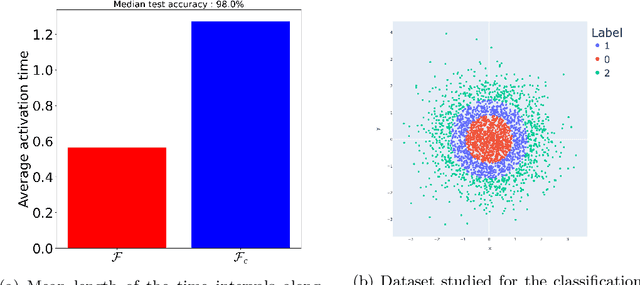
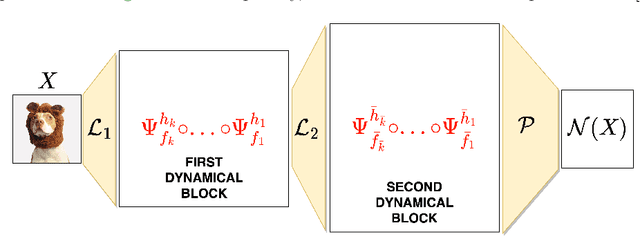
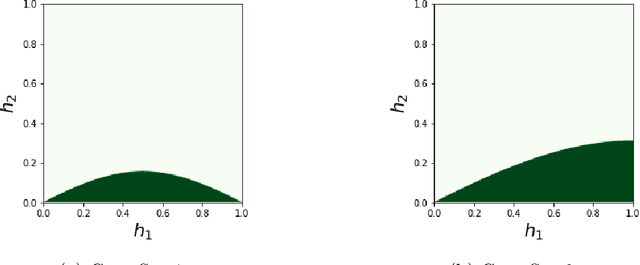
Abstract:Neural networks have gained much interest because of their effectiveness in many applications. However, their mathematical properties are generally not well understood. If there is some underlying geometric structure inherent to the data or to the function to approximate, it is often desirable to take this into account in the design of the neural network. In this work, we start with a non-autonomous ODE and build neural networks using a suitable, structure-preserving, numerical time-discretisation. The structure of the neural network is then inferred from the properties of the ODE vector field. Besides injecting more structure into the network architectures, this modelling procedure allows a better theoretical understanding of their behaviour. We present two universal approximation results and demonstrate how to impose some particular properties on the neural networks. A particular focus is on 1-Lipschitz architectures including layers that are not 1-Lipschitz. These networks are expressive and robust against adversarial attacks, as shown for the CIFAR-10 dataset.
Why Deep Surgical Models Fail?: Revisiting Surgical Action Triplet Recognition through the Lens of Robustness
Sep 18, 2022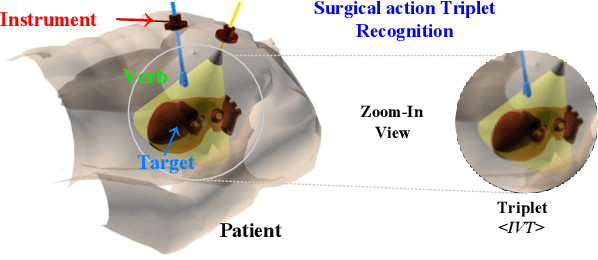
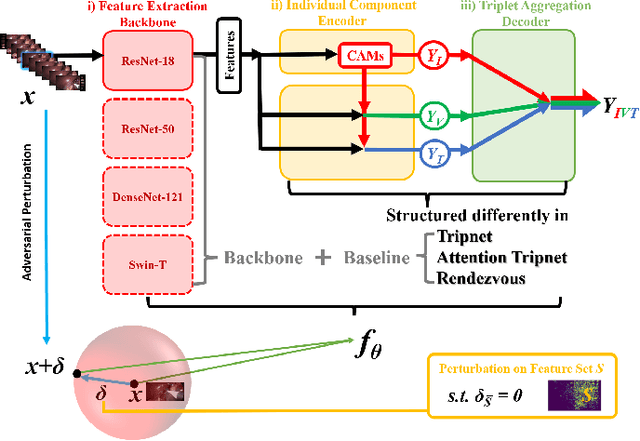
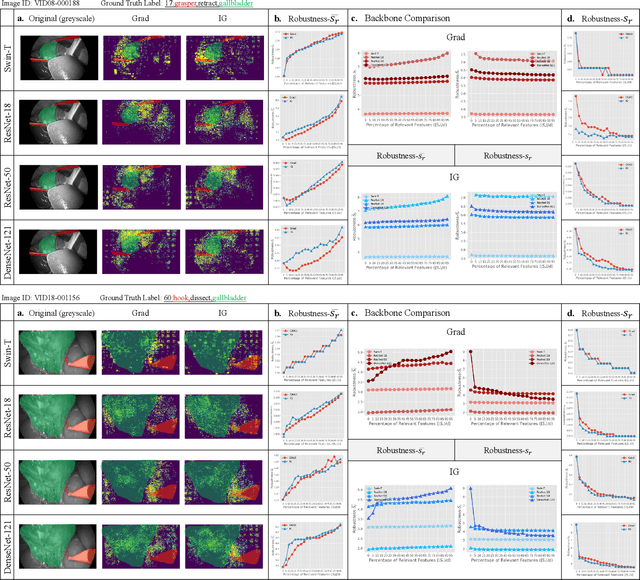
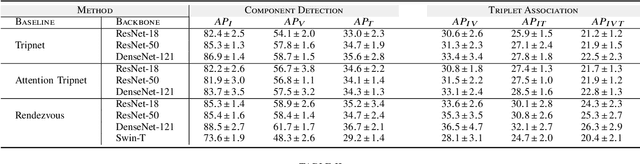
Abstract:Surgical action triplet recognition provides a better understanding of the surgical scene. This task is of high relevance as it provides to the surgeon with context-aware support and safety. The current go-to strategy for improving performance is the development of new network mechanisms. However, the performance of current state-of-the-art techniques is substantially lower than other surgical tasks. Why is this happening? This is the question that we address in this work. We present the first study to understand the failure of existing deep learning models through the lens of robustness and explainabilty. Firstly, we study current existing models under weak and strong $\delta-$perturbations via adversarial optimisation scheme. We then provide the failure modes via feature based explanations. Our study revels that the key for improving performance and increasing reliability is in the core and spurious attributes. Our work opens the door to more trustworthiness and reliability deep learning models in surgical science.
Self-Supervised Learning of Phenotypic Representations from Cell Images with Weak Labels
Sep 16, 2022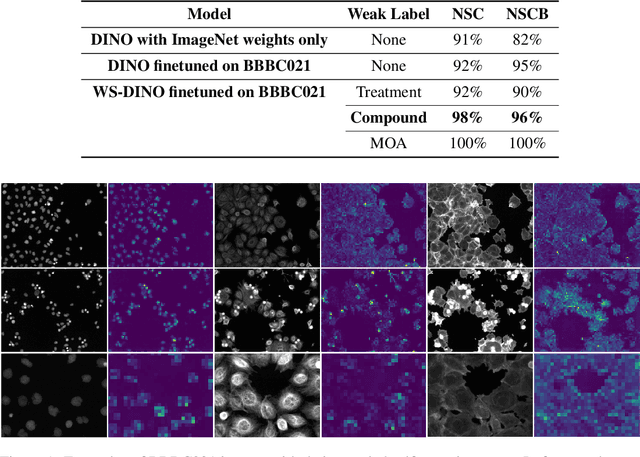
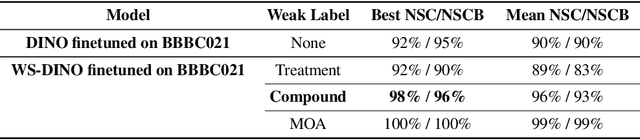
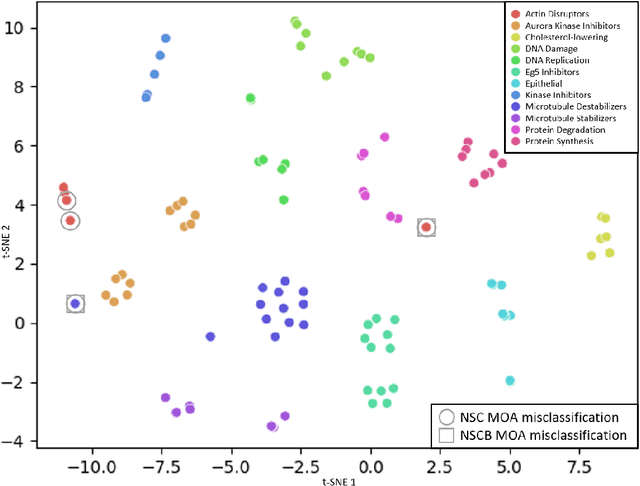
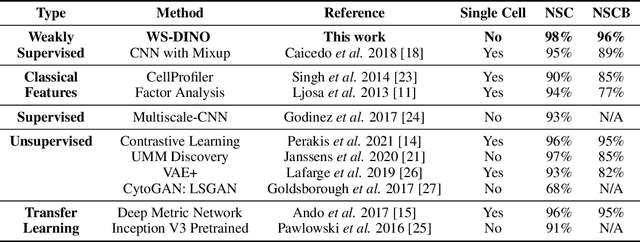
Abstract:We propose WS-DINO as a novel framework to use weak label information in learning phenotypic representations from high-content fluorescent images of cells. Our model is based on a knowledge distillation approach with a vision transformer backbone (DINO), and we use this as a benchmark model for our study. Using WS-DINO, we fine-tuned with weak label information available in high-content microscopy screens (treatment and compound), and achieve state-of-the-art performance in not-same-compound mechanism of action prediction on the BBBC021 dataset (98%), and not-same-compound-and-batch performance (96%) using the compound as the weak label. Our method bypasses single cell cropping as a pre-processing step, and using self-attention maps we show that the model learns structurally meaningful phenotypic profiles.
Spectral decomposition of atomic structures in heterogeneous cryo-EM
Sep 12, 2022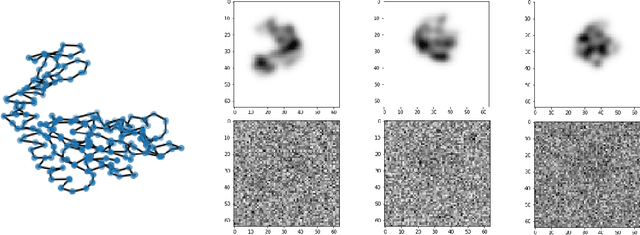
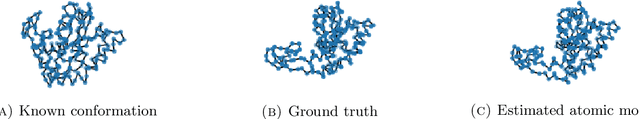
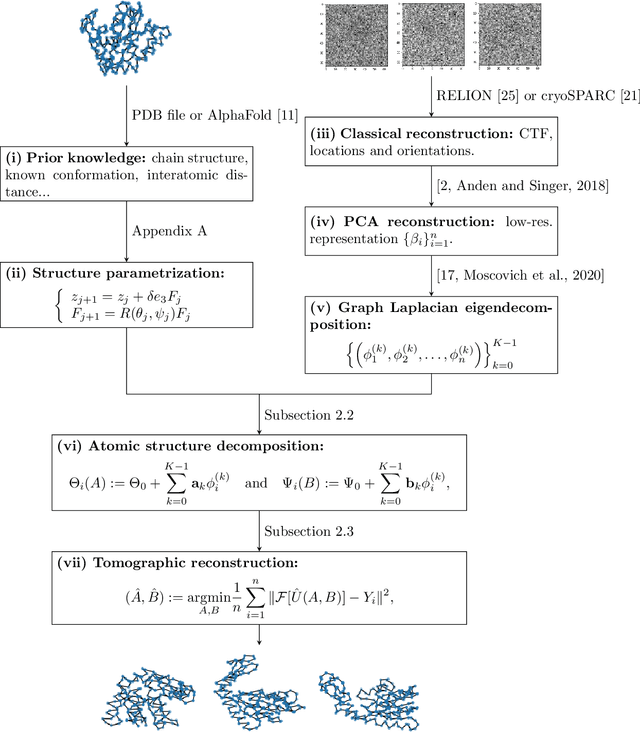


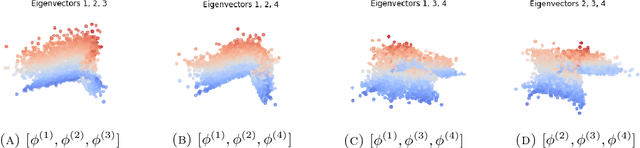
Abstract:We consider the problem of recovering the three-dimensional atomic structure of a flexible macromolecule from a heterogeneous cryo-EM dataset. The dataset contains noisy tomographic projections of the electrostatic potential of the macromolecule, taken from different viewing directions, and in the heterogeneous case, each image corresponds to a different conformation of the macromolecule. Under the assumption that the macromolecule can be modelled as a chain, or discrete curve (as it is for instance the case for a protein backbone with a single chain of amino-acids), we introduce a method to estimate the deformation of the atomic model with respect to a given conformation, which is assumed to be known a priori. Our method consists on estimating the torsion and bond angles of the atomic model in each conformation as a linear combination of the eigenfunctions of the Laplace operator in the manifold of conformations. These eigenfunctions can be approximated by means of a well-known technique in manifold learning, based on the construction of a graph Laplacian using the cryo-EM dataset. Finally, we test our approach with synthetic datasets, for which we recover the atomic model of two-dimensional and three-dimensional flexible structures from noisy tomographic projections.
Imaging with Equivariant Deep Learning
Sep 05, 2022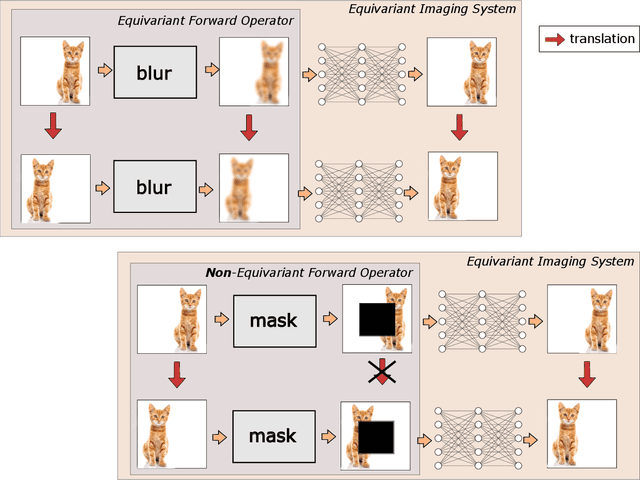
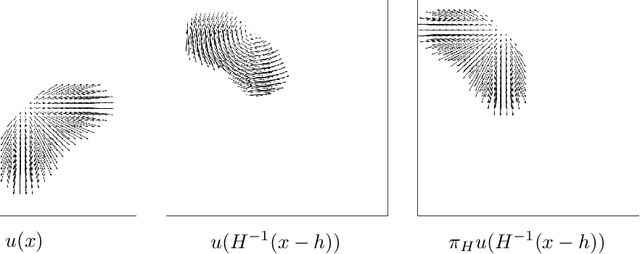


Abstract:From early image processing to modern computational imaging, successful models and algorithms have relied on a fundamental property of natural signals: symmetry. Here symmetry refers to the invariance property of signal sets to transformations such as translation, rotation or scaling. Symmetry can also be incorporated into deep neural networks in the form of equivariance, allowing for more data-efficient learning. While there has been important advances in the design of end-to-end equivariant networks for image classification in recent years, computational imaging introduces unique challenges for equivariant network solutions since we typically only observe the image through some noisy ill-conditioned forward operator that itself may not be equivariant. We review the emerging field of equivariant imaging and show how it can provide improved generalization and new imaging opportunities. Along the way we show the interplay between the acquisition physics and group actions and links to iterative reconstruction, blind compressed sensing and self-supervised learning.
Accelerating Deep Unrolling Networks via Dimensionality Reduction
Aug 31, 2022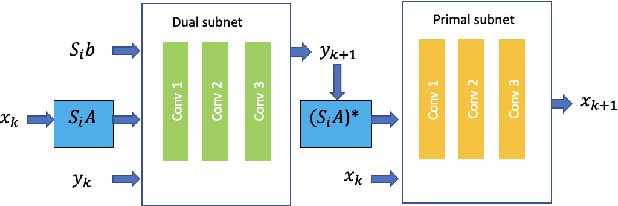
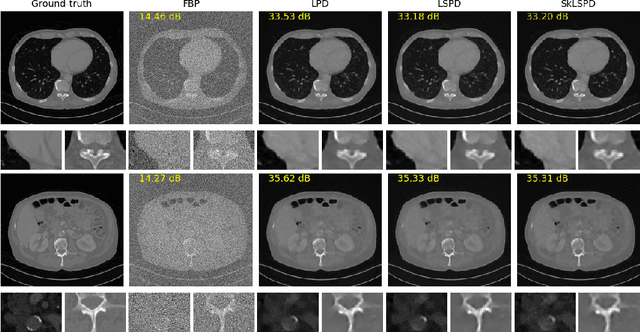
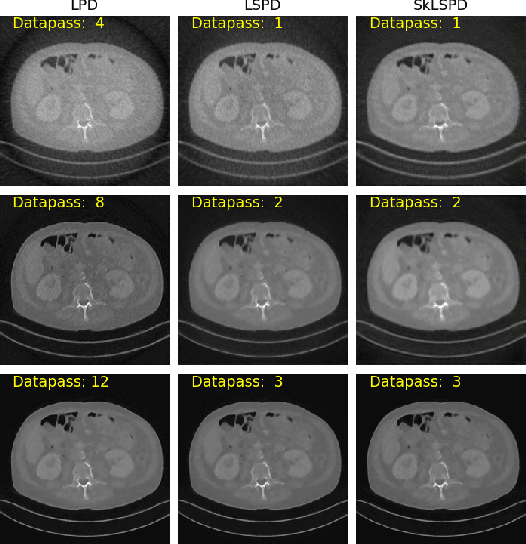
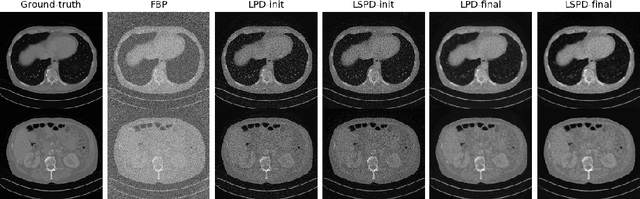
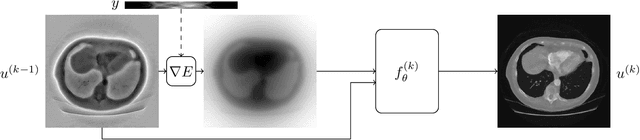
Abstract:In this work we propose a new paradigm for designing efficient deep unrolling networks using dimensionality reduction schemes, including minibatch gradient approximation and operator sketching. The deep unrolling networks are currently the state-of-the-art solutions for imaging inverse problems. However, for high-dimensional imaging tasks, especially X-ray CT and MRI imaging, the deep unrolling schemes typically become inefficient both in terms of memory and computation, due to the need of computing multiple times the high-dimensional forward and adjoint operators. Recently researchers have found that such limitations can be partially addressed by unrolling the stochastic gradient descent (SGD), inspired by the success of stochastic first-order optimization. In this work, we explore further this direction and propose first a more expressive and practical stochastic primal-dual unrolling, based on the state-of-the-art Learned Primal-Dual (LPD) network, and also a further acceleration upon stochastic primal-dual unrolling, using sketching techniques to approximate products in the high-dimensional image space. The operator sketching can be jointly applied with stochastic unrolling for the best acceleration and compression performance. Our numerical experiments on X-ray CT image reconstruction demonstrate the remarkable effectiveness of our accelerated unrolling schemes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge