Bingbing Ni
Learning to Combine: Knowledge Aggregation for Multi-Source Domain Adaptation
Jul 28, 2020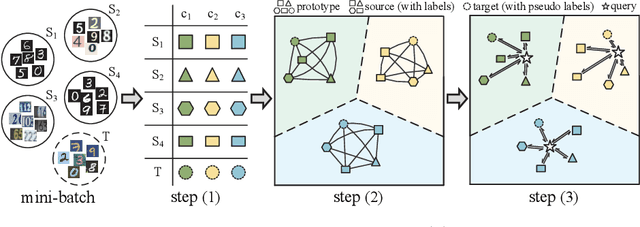
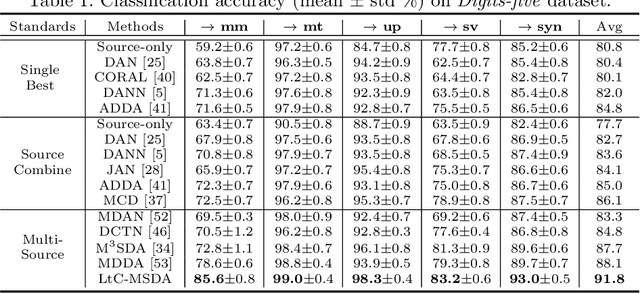
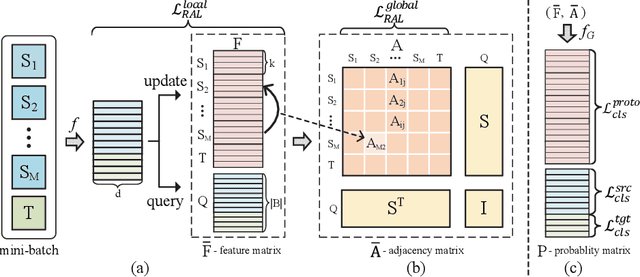

Abstract:Transferring knowledges learned from multiple source domains to target domain is a more practical and challenging task than conventional single-source domain adaptation. Furthermore, the increase of modalities brings more difficulty in aligning feature distributions among multiple domains. To mitigate these problems, we propose a Learning to Combine for Multi-Source Domain Adaptation (LtC-MSDA) framework via exploring interactions among domains. In the nutshell, a knowledge graph is constructed on the prototypes of various domains to realize the information propagation among semantically adjacent representations. On such basis, a graph model is learned to predict query samples under the guidance of correlated prototypes. In addition, we design a Relation Alignment Loss (RAL) to facilitate the consistency of categories' relational interdependency and the compactness of features, which boosts features' intra-class invariance and inter-class separability. Comprehensive results on public benchmark datasets demonstrate that our approach outperforms existing methods with a remarkable margin. Our code is available at \url{https://github.com/ChrisAllenMing/LtC-MSDA}
Self-Prediction for Joint Instance and Semantic Segmentation of Point Clouds
Jul 27, 2020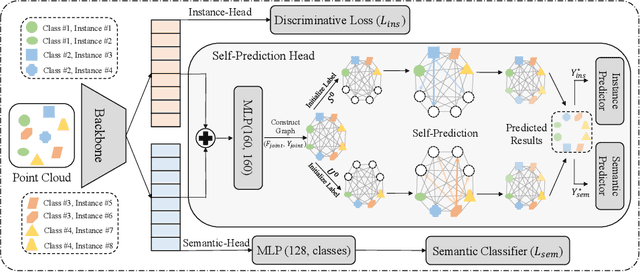
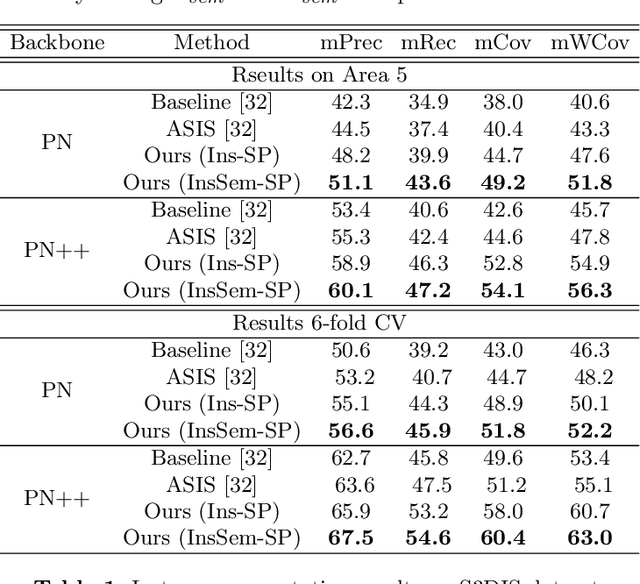
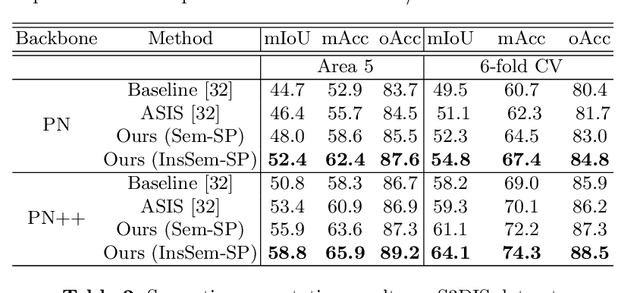
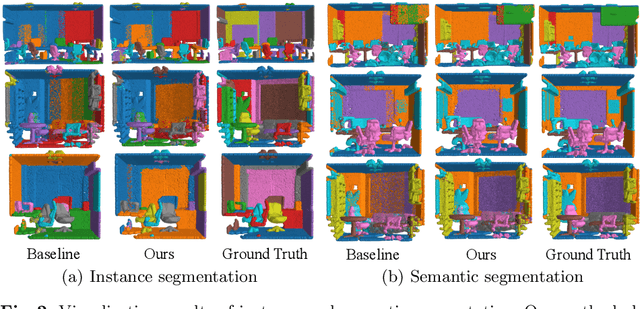
Abstract:We develop a novel learning scheme named Self-Prediction for 3D instance and semantic segmentation of point clouds. Distinct from most existing methods that focus on designing convolutional operators, our method designs a new learning scheme to enhance point relation exploring for better segmentation. More specifically, we divide a point cloud sample into two subsets and construct a complete graph based on their representations. Then we use label propagation algorithm to predict labels of one subset when given labels of the other subset. By training with this Self-Prediction task, the backbone network is constrained to fully explore relational context/geometric/shape information and learn more discriminative features for segmentation. Moreover, a general associated framework equipped with our Self-Prediction scheme is designed for enhancing instance and semantic segmentation simultaneously, where instance and semantic representations are combined to perform Self-Prediction. Through this way, instance and semantic segmentation are collaborated and mutually reinforced. Significant performance improvements on instance and semantic segmentation compared with baseline are achieved on S3DIS and ShapeNet. Our method achieves state-of-the-art instance segmentation results on S3DIS and comparable semantic segmentation results compared with state-of-the-arts on S3DIS and ShapeNet when we only take PointNet++ as the backbone network.
Collaborative Learning for Faster StyleGAN Embedding
Jul 03, 2020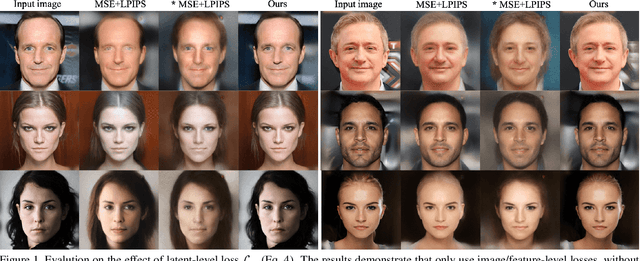
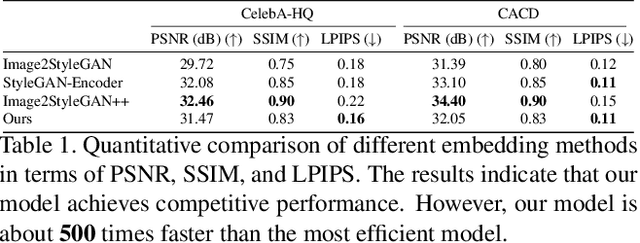
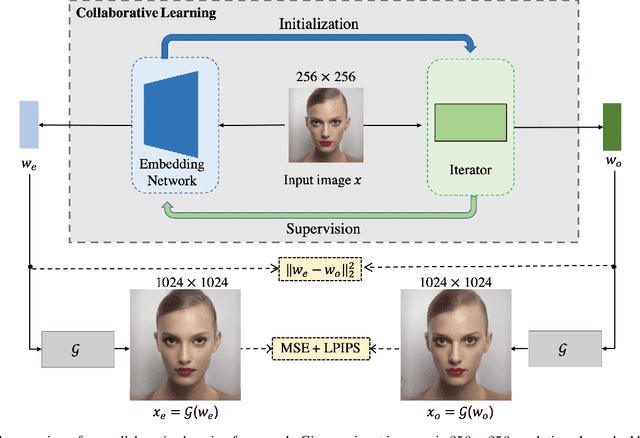
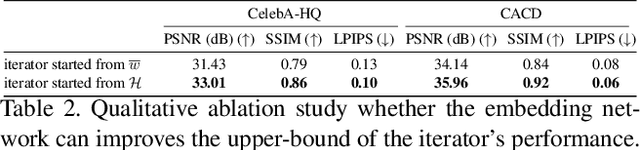
Abstract:The latent code of the recent popular model StyleGAN has learned disentangled representations thanks to the multi-layer style-based generator. Embedding a given image back to the latent space of StyleGAN enables wide interesting semantic image editing applications. Although previous works are able to yield impressive inversion results based on an optimization framework, which however suffers from the efficiency issue. In this work, we propose a novel collaborative learning framework that consists of an efficient embedding network and an optimization-based iterator. On one hand, with the progress of training, the embedding network gives a reasonable latent code initialization for the iterator. On the other hand, the updated latent code from the iterator in turn supervises the embedding network. In the end, high-quality latent code can be obtained efficiently with a single forward pass through our embedding network. Extensive experiments demonstrate the effectiveness and efficiency of our work.
Video Prediction via Example Guidance
Jul 03, 2020
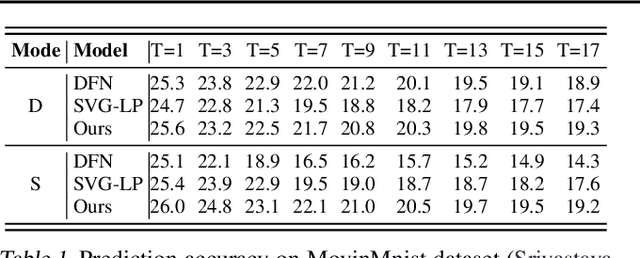
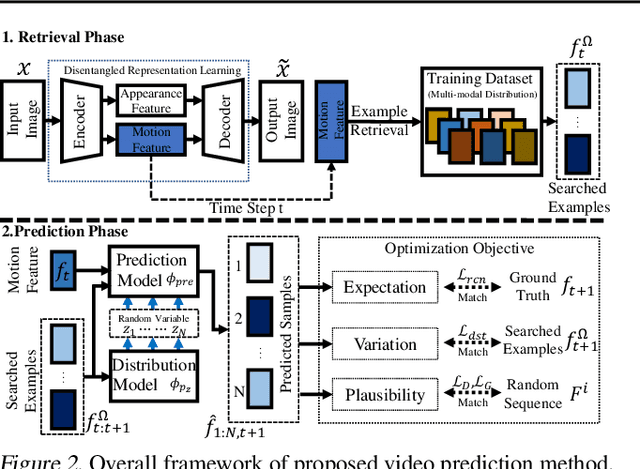
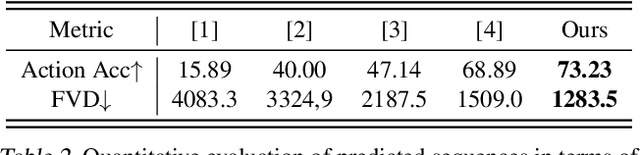
Abstract:In video prediction tasks, one major challenge is to capture the multi-modal nature of future contents and dynamics. In this work, we propose a simple yet effective framework that can efficiently predict plausible future states. The key insight is that the potential distribution of a sequence could be approximated with analogous ones in a repertoire of training pool, namely, expert examples. By further incorporating a novel optimization scheme into the training procedure, plausible predictions can be sampled efficiently from distribution constructed from the retrieved examples. Meanwhile, our method could be seamlessly integrated with existing stochastic predictive models; significant enhancement is observed with comprehensive experiments in both quantitative and qualitative aspects. We also demonstrate the generalization ability to predict the motion of unseen class, i.e., without access to corresponding data during training phase.
* Project Page: https://sites.google.com/view/vpeg-supp/home
Searching towards Class-Aware Generators for Conditional Generative Adversarial Networks
Jun 25, 2020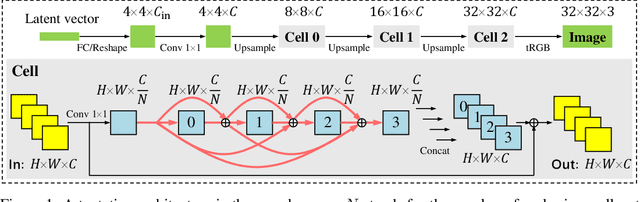
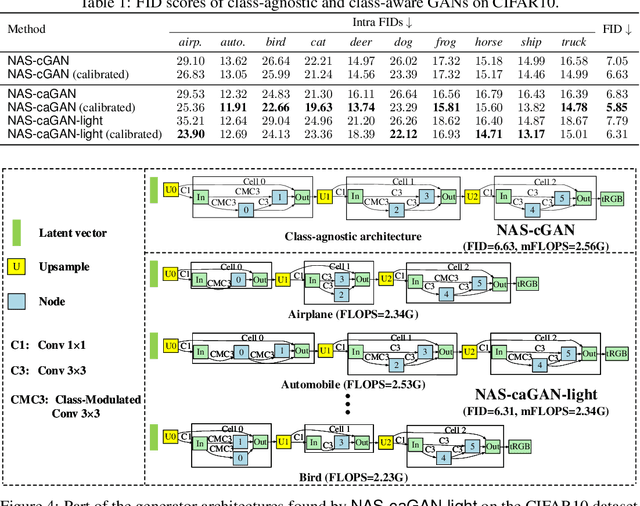
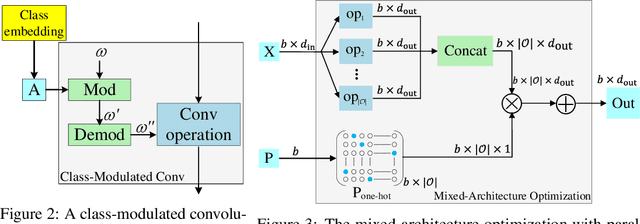
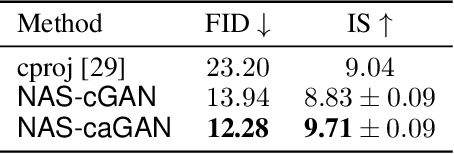
Abstract:Conditional Generative Adversarial Networks (cGAN) were designed to generate images based on the provided conditions, e.g., class-level distributions. However, existing methods have used the same generating architecture for all classes. This paper presents a novel idea that adopts NAS to find a distinct architecture for each class. The search space contains regular and class-modulated convolutions, where the latter is designed to introduce class-specific information while avoiding the reduction of training data for each class generator. The search algorithm follows a weight-sharing pipeline with mixed-architecture optimization so that the search cost does not grow with the number of classes. To learn the sampling policy, a Markov decision process is embedded into the search algorithm and a moving average is applied for better stability. We evaluate our approach on CIFAR10 and CIFAR100. Besides demonstrating superior performance, we deliver several insights that are helpful in designing efficient GAN models. Code is available \url{https://github.com/PeterouZh/NAS_cGAN}.
AlignShift: Bridging the Gap of Imaging Thickness in 3D Anisotropic Volumes
May 05, 2020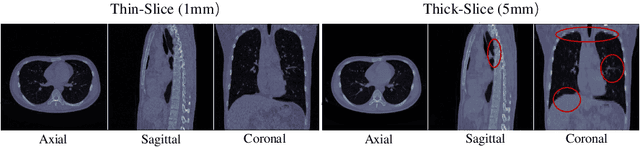
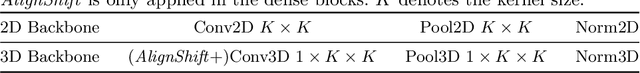
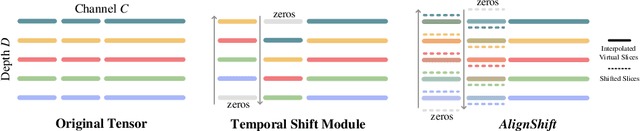
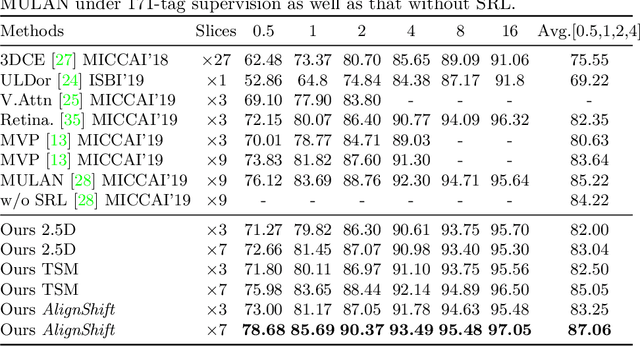
Abstract:This paper addresses a fundamental challenge in 3D medical image processing: how to deal with imaging thickness. For anisotropic medical volumes, there is a significant performance gap between thin-slice (mostly 1mm) and thick-slice (mostly 5mm) volumes. Prior arts tend to use 3D approaches for the thin-slice and 2D approaches for the thick-slice, respectively. We aim at a unified approach for both thin- and thick-slice medical volumes. Inspired by recent advances in video analysis, we propose AlignShift, a novel parameter-free operator to convert theoretically any 2D pretrained network into thickness-aware 3D network. Remarkably, the converted networks behave like 3D for the thin-slice, nevertheless degenerate to 2D for the thick-slice adaptively. The unified thickness-aware representation learning is achieved by shifting and fusing aligned "virtual slices" as per the input imaging thickness. Extensive experiments on public large-scale DeepLesion benchmark, consisting of 32K lesions for universal lesion detection, validate the effectiveness of our method, which outperforms previous state of the art by considerable margins, without whistles and bells. More importantly, to our knowledge, this is the first method that bridges the performance gap between thin- and thick-slice volumes by a unified framework. To improve research reproducibility, our code in PyTorch is open source at https://github.com/M3DV/AlignShift.
Relational Learning between Multiple Pulmonary Nodules via Deep Set Attention Transformers
Apr 12, 2020


Abstract:Diagnosis and treatment of multiple pulmonary nodules are clinically important but challenging. Prior studies on nodule characterization use solitary-nodule approaches on multiple nodular patients, which ignores the relations between nodules. In this study, we propose a multiple instance learning (MIL) approach and empirically prove the benefit to learn the relations between multiple nodules. By treating the multiple nodules from a same patient as a whole, critical relational information between solitary-nodule voxels is extracted. To our knowledge, it is the first study to learn the relations between multiple pulmonary nodules. Inspired by recent advances in natural language processing (NLP) domain, we introduce a self-attention transformer equipped with 3D CNN, named {NoduleSAT}, to replace typical pooling-based aggregation in multiple instance learning. Extensive experiments on lung nodule false positive reduction on LUNA16 database, and malignancy classification on LIDC-IDRI database, validate the effectiveness of the proposed method.
Cross-domain Detection via Graph-induced Prototype Alignment
Mar 28, 2020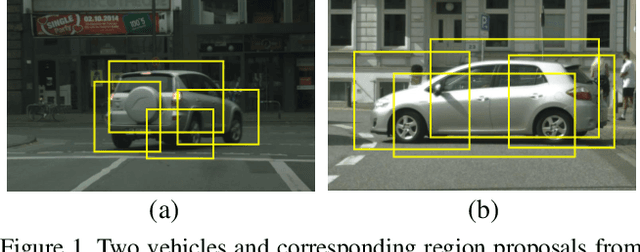
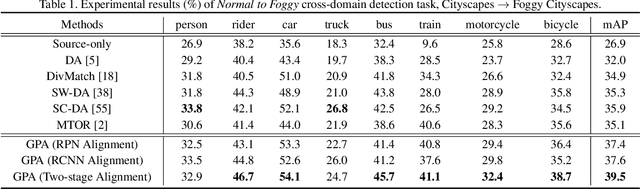
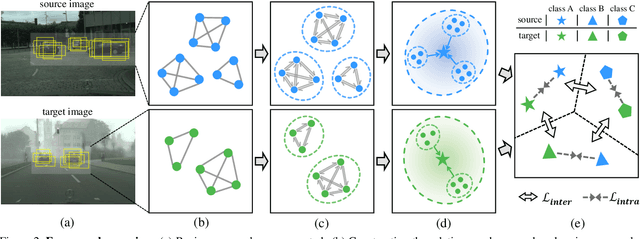
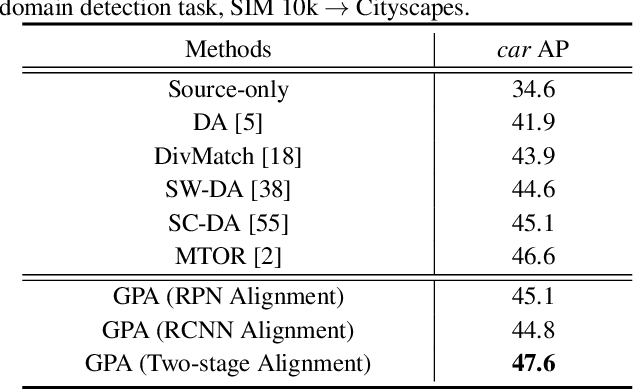
Abstract:Applying the knowledge of an object detector trained on a specific domain directly onto a new domain is risky, as the gap between two domains can severely degrade model's performance. Furthermore, since different instances commonly embody distinct modal information in object detection scenario, the feature alignment of source and target domain is hard to be realized. To mitigate these problems, we propose a Graph-induced Prototype Alignment (GPA) framework to seek for category-level domain alignment via elaborate prototype representations. In the nutshell, more precise instance-level features are obtained through graph-based information propagation among region proposals, and, on such basis, the prototype representation of each class is derived for category-level domain alignment. In addition, in order to alleviate the negative effect of class-imbalance on domain adaptation, we design a Class-reweighted Contrastive Loss to harmonize the adaptation training process. Combining with Faster R-CNN, the proposed framework conducts feature alignment in a two-stage manner. Comprehensive results on various cross-domain detection tasks demonstrate that our approach outperforms existing methods with a remarkable margin. Our code is available at https://github.com/ChrisAllenMing/GPA-detection.
Adversarial Domain Adaptation with Domain Mixup
Dec 04, 2019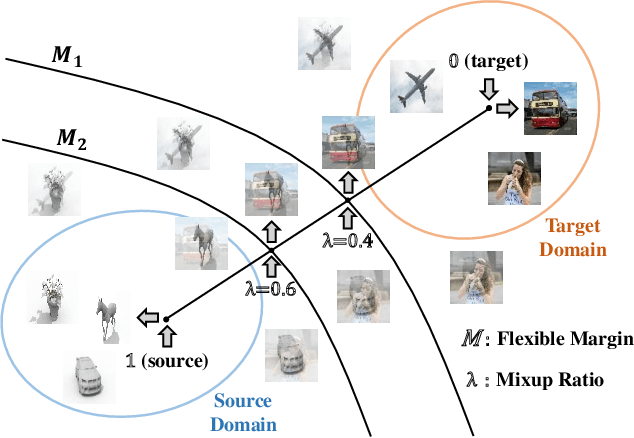
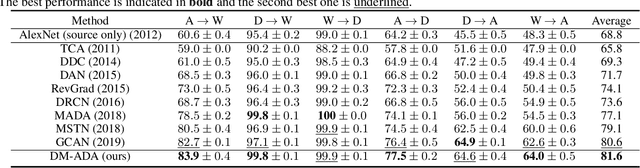
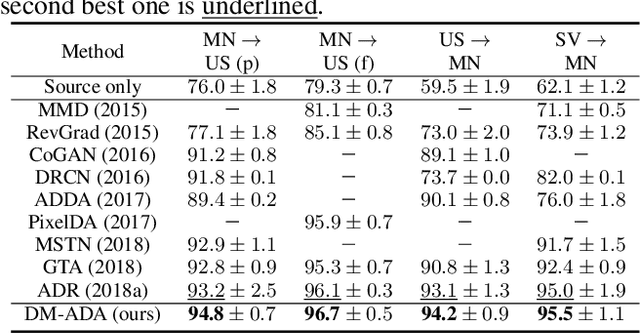
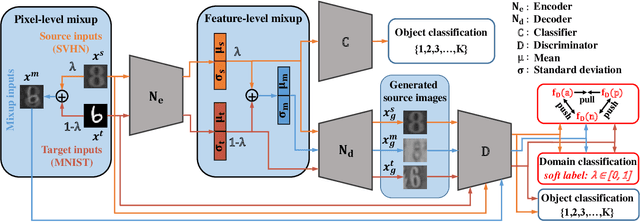

Abstract:Recent works on domain adaptation reveal the effectiveness of adversarial learning on filling the discrepancy between source and target domains. However, two common limitations exist in current adversarial-learning-based methods. First, samples from two domains alone are not sufficient to ensure domain-invariance at most part of latent space. Second, the domain discriminator involved in these methods can only judge real or fake with the guidance of hard label, while it is more reasonable to use soft scores to evaluate the generated images or features, i.e., to fully utilize the inter-domain information. In this paper, we present adversarial domain adaptation with domain mixup (DM-ADA), which guarantees domain-invariance in a more continuous latent space and guides the domain discriminator in judging samples' difference relative to source and target domains. Domain mixup is jointly conducted on pixel and feature level to improve the robustness of models. Extensive experiments prove that the proposed approach can achieve superior performance on tasks with various degrees of domain shift and data complexity.
Reinventing 2D Convolutions for 3D Medical Images
Nov 24, 2019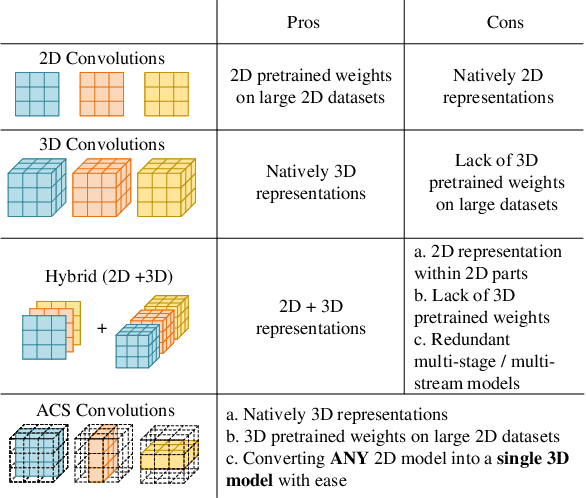
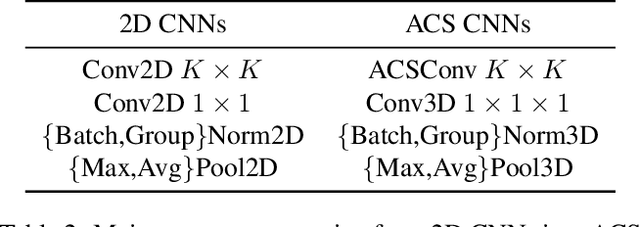
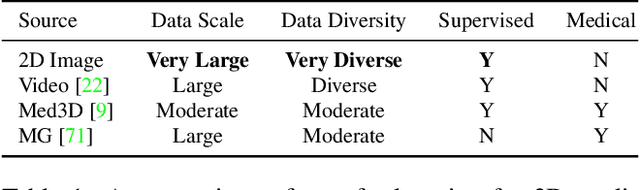
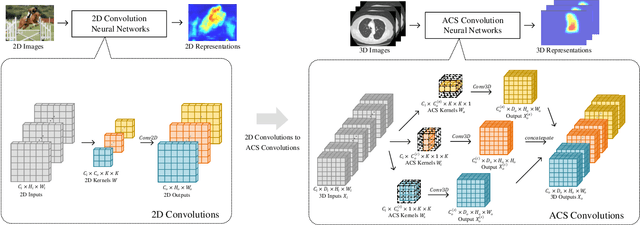

Abstract:There has been considerable debate over 2D and 3D representation learning on 3D medical images. 2D approaches could benefit from large-scale 2D pretraining, whereas they are generally weak in capturing large 3D contexts. 3D approaches are natively strong in 3D contexts, however few publicly available 3D medical dataset is large and diverse enough for universal 3D pretraining. Even for hybrid (2D + 3D) approaches, the intrinsic disadvantages within the 2D / 3D parts still exist. In this study, we bridge the gap between 2D and 3D convolutions by reinventing the 2D convolutions. We propose ACS (axial-coronal-sagittal) convolutions to perform natively 3D representation learning, while utilizing the pretrained weights from 2D counterparts. In ACS convolutions, 2D convolution kernels are split by channel into three parts, and convoluted separately on the three views (axial, coronal and sagittal) of 3D representations. Theoretically, ANY 2D CNN (ResNet, DenseNet, or DeepLab) is able to be converted into a 3D ACS CNN, with pretrained weights of same parameter sizes. Extensive experiments on proof-of-concept dataset and several medical benchmarks validate the consistent superiority of the pretrained ACS CNNs, over the 2D / 3D CNN counterparts with / without pretraining. Even without pretraining, the ACS convolution can be used as a plug-and-play replacement of standard 3D convolution, with smaller model size.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge