Ben Glocker
Biomedical Image Analysis Group, Department of Computing, Imperial College London
CrossMoDA 2021 challenge: Benchmark of Cross-Modality Domain Adaptation techniques for Vestibular Schwnannoma and Cochlea Segmentation
Jan 08, 2022
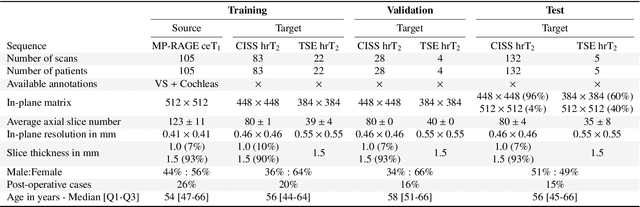
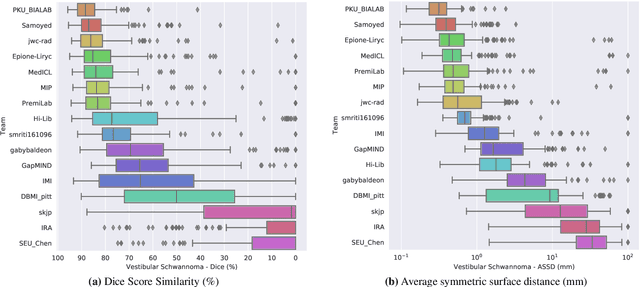
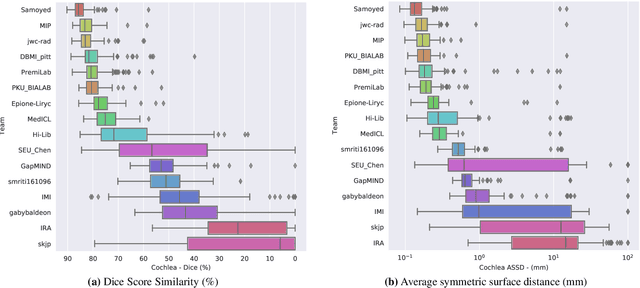
Abstract:Domain Adaptation (DA) has recently raised strong interests in the medical imaging community. While a large variety of DA techniques has been proposed for image segmentation, most of these techniques have been validated either on private datasets or on small publicly available datasets. Moreover, these datasets mostly addressed single-class problems. To tackle these limitations, the Cross-Modality Domain Adaptation (crossMoDA) challenge was organised in conjunction with the 24th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2021). CrossMoDA is the first large and multi-class benchmark for unsupervised cross-modality DA. The challenge's goal is to segment two key brain structures involved in the follow-up and treatment planning of vestibular schwannoma (VS): the VS and the cochleas. Currently, the diagnosis and surveillance in patients with VS are performed using contrast-enhanced T1 (ceT1) MRI. However, there is growing interest in using non-contrast sequences such as high-resolution T2 (hrT2) MRI. Therefore, we created an unsupervised cross-modality segmentation benchmark. The training set provides annotated ceT1 (N=105) and unpaired non-annotated hrT2 (N=105). The aim was to automatically perform unilateral VS and bilateral cochlea segmentation on hrT2 as provided in the testing set (N=137). A total of 16 teams submitted their algorithm for the evaluation phase. The level of performance reached by the top-performing teams is strikingly high (best median Dice - VS:88.4%; Cochleas:85.7%) and close to full supervision (median Dice - VS:92.5%; Cochleas:87.7%). All top-performing methods made use of an image-to-image translation approach to transform the source-domain images into pseudo-target-domain images. A segmentation network was then trained using these generated images and the manual annotations provided for the source image.
Algorithmic encoding of protected characteristics and its implications on disparities across subgroups
Oct 27, 2021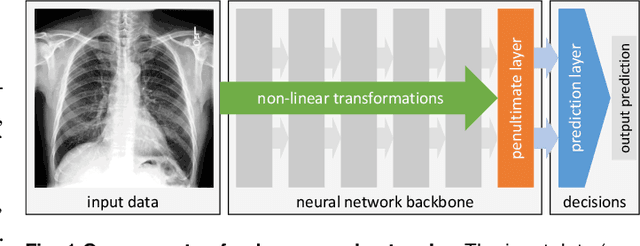
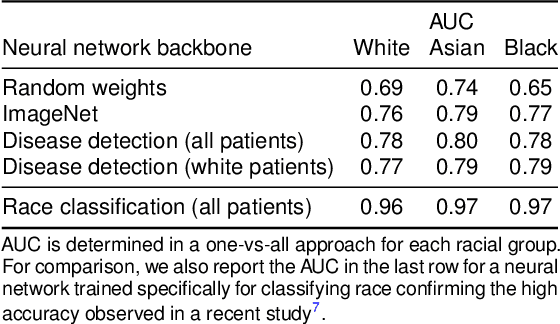
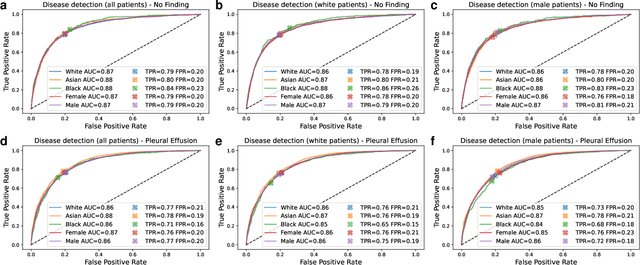
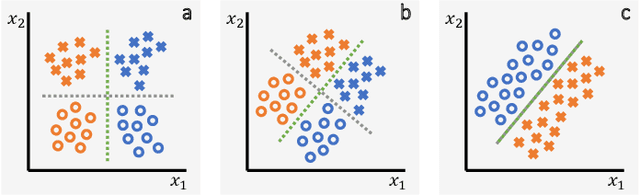
Abstract:It has been rightfully emphasized that the use of AI for clinical decision making could amplify health disparities. A machine learning model may pick up undesirable correlations, for example, between a patient's racial identity and clinical outcome. Such correlations are often present in (historical) data used for model development. There has been an increase in studies reporting biases in disease detection models across patient subgroups. Besides the scarcity of data from underserved populations, very little is known about how these biases are encoded and how one may reduce or even remove disparate performance. There is some speculation whether algorithms may recognize patient characteristics such as biological sex or racial identity, and then directly or indirectly use this information when making predictions. But it remains unclear how we can establish whether such information is actually used. This article aims to shed some light on these issues by exploring new methodology allowing intuitive inspections of the inner working of machine learning models for image-based detection of disease. We also evaluate an effective yet debatable technique for addressing disparities leveraging the automatic prediction of patient characteristics, resulting in models with comparable true and false positive rates across subgroups. Our findings may stimulate the discussion about safe and ethical use of AI.
Uncertainty quantification in non-rigid image registration via stochastic gradient Markov chain Monte Carlo
Oct 25, 2021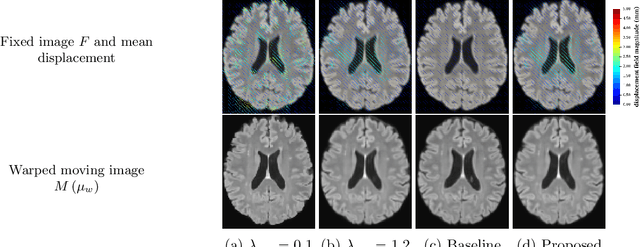
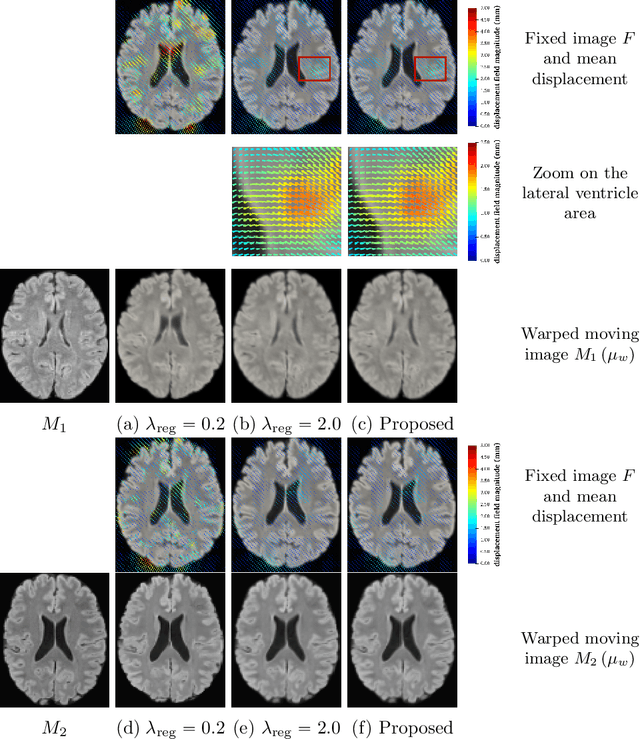

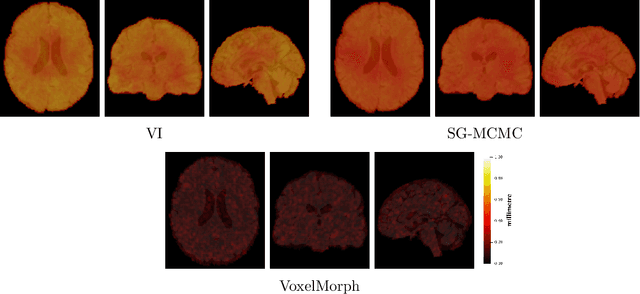
Abstract:We develop a new Bayesian model for non-rigid registration of three-dimensional medical images, with a focus on uncertainty quantification. Probabilistic registration of large images with calibrated uncertainty estimates is difficult for both computational and modelling reasons. To address the computational issues, we explore connections between the Markov chain Monte Carlo by backpropagation and the variational inference by backpropagation frameworks, in order to efficiently draw samples from the posterior distribution of transformation parameters. To address the modelling issues, we formulate a Bayesian model for image registration that overcomes the existing barriers when using a dense, high-dimensional, and diffeomorphic transformation parametrisation. This results in improved calibration of uncertainty estimates. We compare the model in terms of both image registration accuracy and uncertainty quantification to VoxelMorph, a state-of-the-art image registration model based on deep learning.
Is MC Dropout Bayesian?
Oct 08, 2021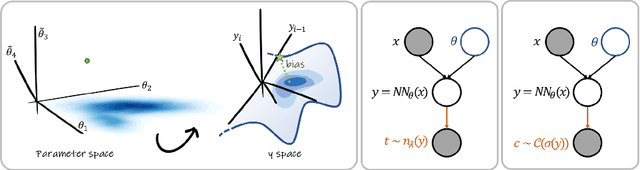
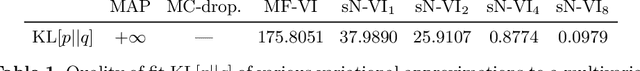
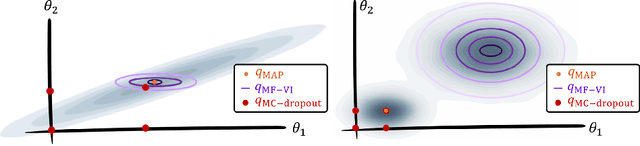

Abstract:MC Dropout is a mainstream "free lunch" method in medical imaging for approximate Bayesian computations (ABC). Its appeal is to solve out-of-the-box the daunting task of ABC and uncertainty quantification in Neural Networks (NNs); to fall within the variational inference (VI) framework; and to propose a highly multimodal, faithful predictive posterior. We question the properties of MC Dropout for approximate inference, as in fact MC Dropout changes the Bayesian model; its predictive posterior assigns $0$ probability to the true model on closed-form benchmarks; the multimodality of its predictive posterior is not a property of the true predictive posterior but a design artefact. To address the need for VI on arbitrary models, we share a generic VI engine within the pytorch framework. The code includes a carefully designed implementation of structured (diagonal plus low-rank) multivariate normal variational families, and mixtures thereof. It is intended as a go-to no-free-lunch approach, addressing shortcomings of mean-field VI with an adjustable trade-off between expressivity and computational complexity.
DeepMCAT: Large-Scale Deep Clustering for Medical Image Categorization
Sep 30, 2021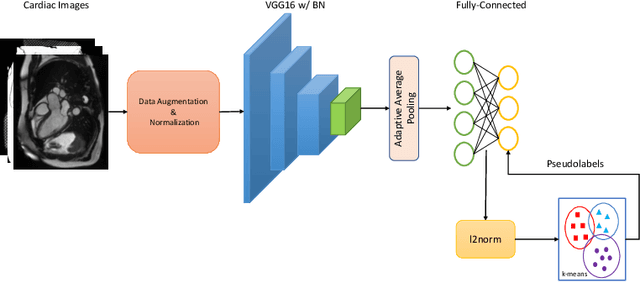
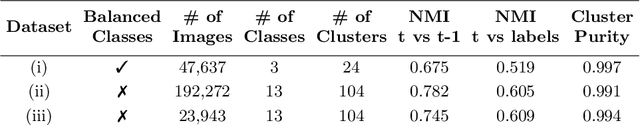
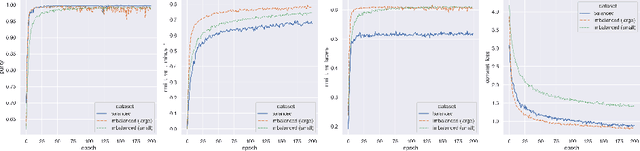
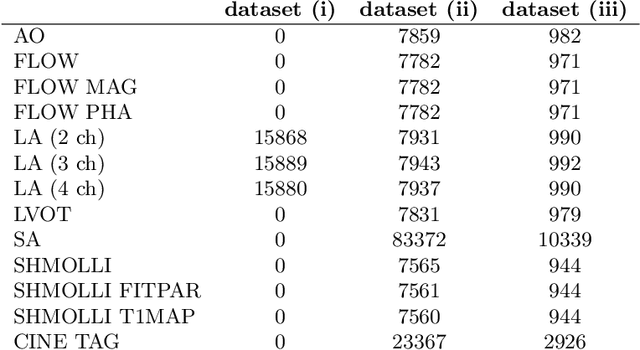
Abstract:In recent years, the research landscape of machine learning in medical imaging has changed drastically from supervised to semi-, weakly- or unsupervised methods. This is mainly due to the fact that ground-truth labels are time-consuming and expensive to obtain manually. Generating labels from patient metadata might be feasible but it suffers from user-originated errors which introduce biases. In this work, we propose an unsupervised approach for automatically clustering and categorizing large-scale medical image datasets, with a focus on cardiac MR images, and without using any labels. We investigated the end-to-end training using both class-balanced and imbalanced large-scale datasets. Our method was able to create clusters with high purity and achieved over 0.99 cluster purity on these datasets. The results demonstrate the potential of the proposed method for categorizing unstructured large medical databases, such as organizing clinical PACS systems in hospitals.
Class-Distribution-Aware Calibration for Long-Tailed Visual Recognition
Sep 11, 2021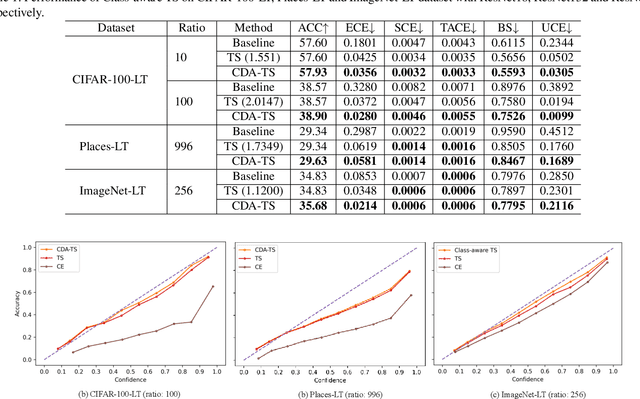
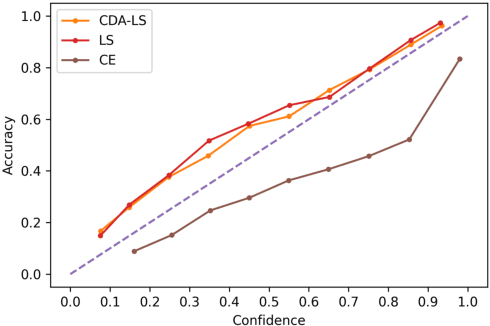
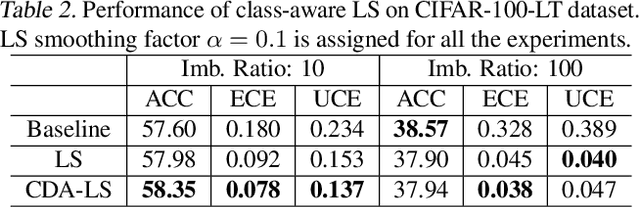
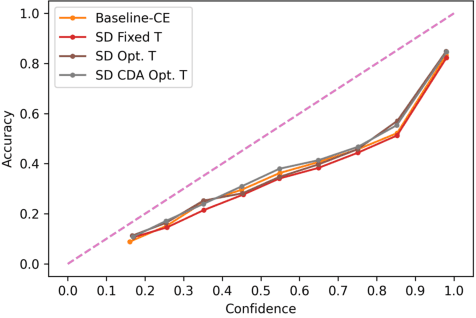
Abstract:Despite impressive accuracy, deep neural networks are often miscalibrated and tend to overly confident predictions. Recent techniques like temperature scaling (TS) and label smoothing (LS) show effectiveness in obtaining a well-calibrated model by smoothing logits and hard labels with scalar factors, respectively. However, the use of uniform TS or LS factor may not be optimal for calibrating models trained on a long-tailed dataset where the model produces overly confident probabilities for high-frequency classes. In this study, we propose class-distribution-aware TS (CDA-TS) and LS (CDA-LS) by incorporating class frequency information in model calibration in the context of long-tailed distribution. In CDA-TS, the scalar temperature value is replaced with the CDA temperature vector encoded with class frequency to compensate for the over-confidence. Similarly, CDA-LS uses a vector smoothing factor and flattens the hard labels according to their corresponding class distribution. We also integrate CDA optimal temperature vector with distillation loss, which reduces miscalibration in self-distillation (SD). We empirically show that class-distribution-aware TS and LS can accommodate the imbalanced data distribution yielding superior performance in both calibration error and predictive accuracy. We also observe that SD with an extremely imbalanced dataset is less effective in terms of calibration performance. Code is available in https://github.com/mobarakol/Class-Distribution-Aware-TS-LS.
Active label cleaning: Improving dataset quality under resource constraints
Sep 01, 2021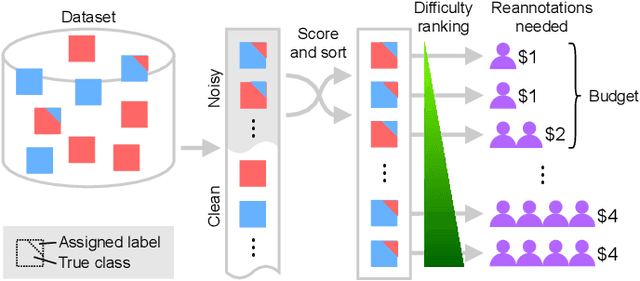
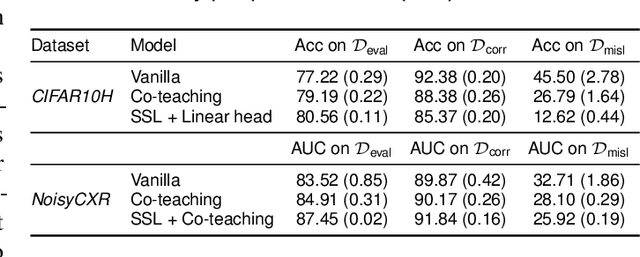
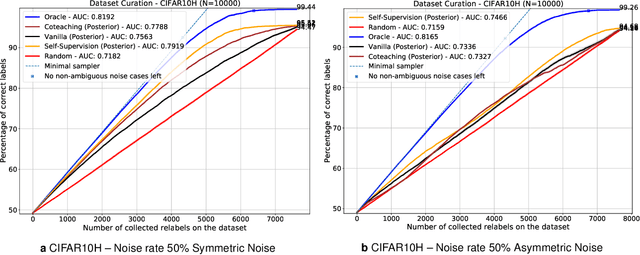
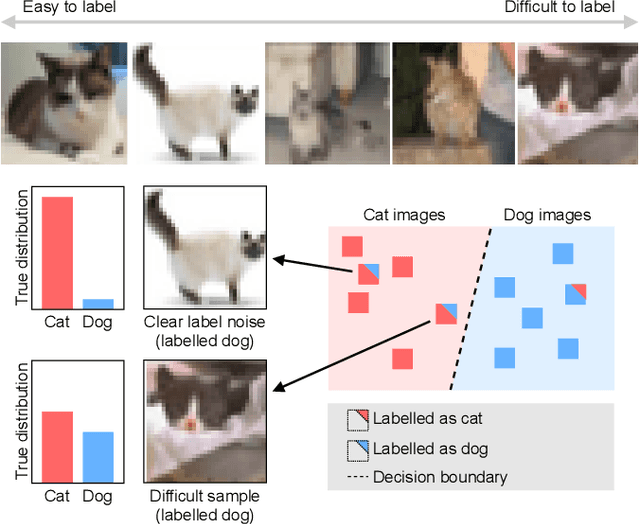
Abstract:Imperfections in data annotation, known as label noise, are detrimental to the training of machine learning models and have an often-overlooked confounding effect on the assessment of model performance. Nevertheless, employing experts to remove label noise by fully re-annotating large datasets is infeasible in resource-constrained settings, such as healthcare. This work advocates for a data-driven approach to prioritising samples for re-annotation - which we term "active label cleaning". We propose to rank instances according to estimated label correctness and labelling difficulty of each sample, and introduce a simulation framework to evaluate relabelling efficacy. Our experiments on natural images and on a new medical imaging benchmark show that cleaning noisy labels mitigates their negative impact on model training, evaluation, and selection. Crucially, the proposed active label cleaning enables correcting labels up to 4 times more effectively than typical random selection in realistic conditions, making better use of experts' valuable time for improving dataset quality.
The Pitfalls of Sample Selection: A Case Study on Lung Nodule Classification
Aug 11, 2021
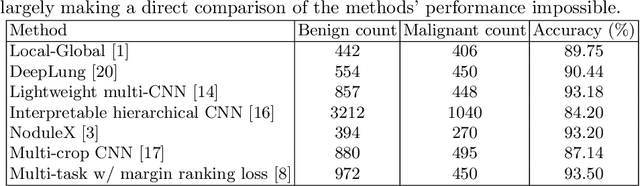
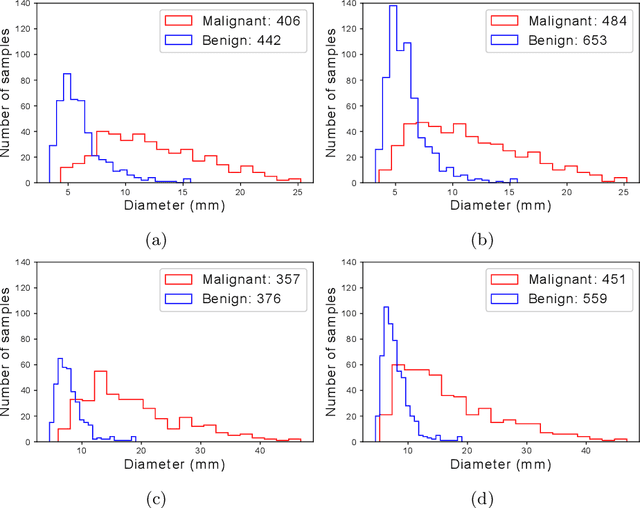
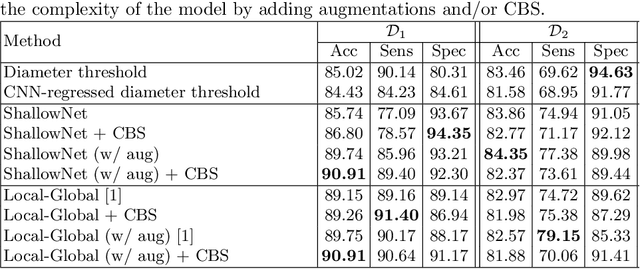
Abstract:Using publicly available data to determine the performance of methodological contributions is important as it facilitates reproducibility and allows scrutiny of the published results. In lung nodule classification, for example, many works report results on the publicly available LIDC dataset. In theory, this should allow a direct comparison of the performance of proposed methods and assess the impact of individual contributions. When analyzing seven recent works, however, we find that each employs a different data selection process, leading to largely varying total number of samples and ratios between benign and malignant cases. As each subset will have different characteristics with varying difficulty for classification, a direct comparison between the proposed methods is thus not always possible, nor fair. We study the particular effect of truthing when aggregating labels from multiple experts. We show that specific choices can have severe impact on the data distribution where it may be possible to achieve superior performance on one sample distribution but not on another. While we show that we can further improve on the state-of-the-art on one sample selection, we also find that on a more challenging sample selection, on the same database, the more advanced models underperform with respect to very simple baseline methods, highlighting that the selected data distribution may play an even more important role than the model architecture. This raises concerns about the validity of claimed methodological contributions. We believe the community should be aware of these pitfalls and make recommendations on how these can be avoided in future work.
The Effect of the Loss on Generalization: Empirical Study on Synthetic Lung Nodule Data
Aug 10, 2021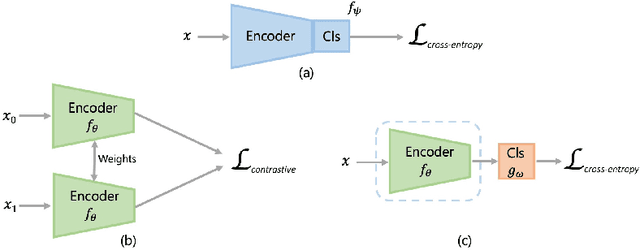
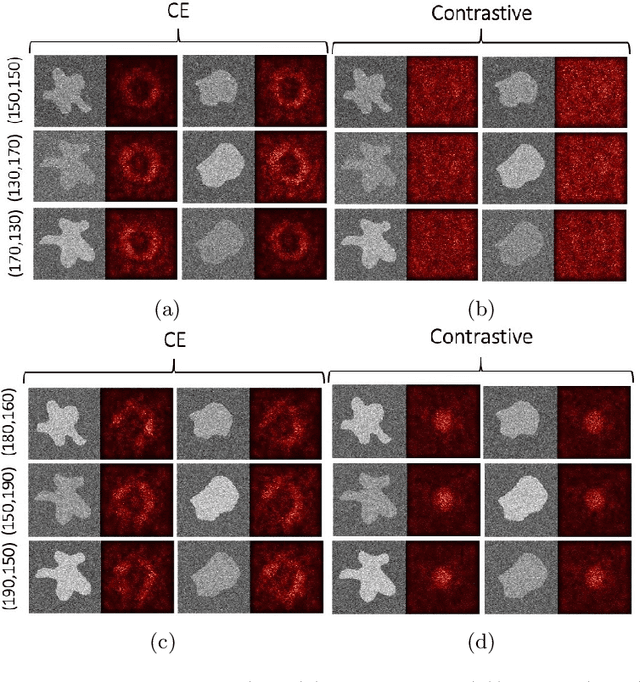
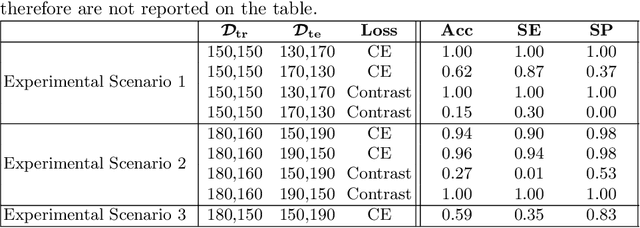
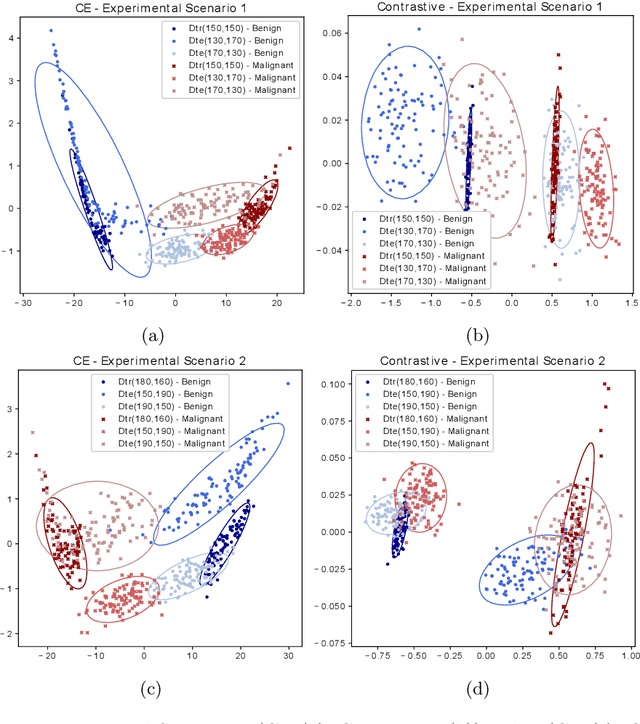

Abstract:Convolutional Neural Networks (CNNs) are widely used for image classification in a variety of fields, including medical imaging. While most studies deploy cross-entropy as the loss function in such tasks, a growing number of approaches have turned to a family of contrastive learning-based losses. Even though performance metrics such as accuracy, sensitivity and specificity are regularly used for the evaluation of CNN classifiers, the features that these classifiers actually learn are rarely identified and their effect on the classification performance on out-of-distribution test samples is insufficiently explored. In this paper, motivated by the real-world task of lung nodule classification, we investigate the features that a CNN learns when trained and tested on different distributions of a synthetic dataset with controlled modes of variation. We show that different loss functions lead to different features being learned and consequently affect the generalization ability of the classifier on unseen data. This study provides some important insights into the design of deep learning solutions for medical imaging tasks.
Transductive image segmentation: Self-training and effect of uncertainty estimation
Aug 02, 2021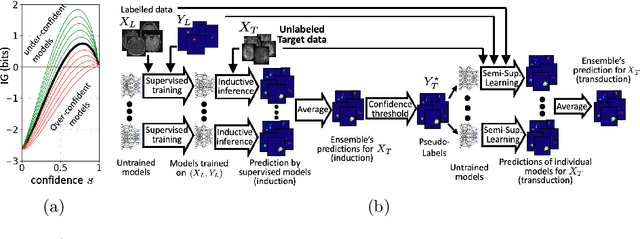

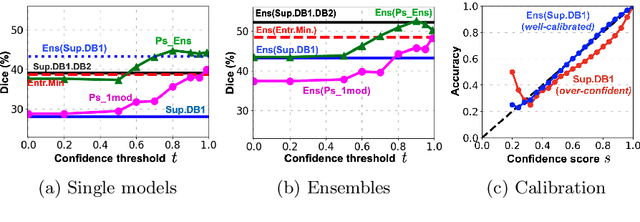
Abstract:Semi-supervised learning (SSL) uses unlabeled data during training to learn better models. Previous studies on SSL for medical image segmentation focused mostly on improving model generalization to unseen data. In some applications, however, our primary interest is not generalization but to obtain optimal predictions on a specific unlabeled database that is fully available during model development. Examples include population studies for extracting imaging phenotypes. This work investigates an often overlooked aspect of SSL, transduction. It focuses on the quality of predictions made on the unlabeled data of interest when they are included for optimization during training, rather than improving generalization. We focus on the self-training framework and explore its potential for transduction. We analyze it through the lens of Information Gain and reveal that learning benefits from the use of calibrated or under-confident models. Our extensive experiments on a large MRI database for multi-class segmentation of traumatic brain lesions shows promising results when comparing transductive with inductive predictions. We believe this study will inspire further research on transductive learning, a well-suited paradigm for medical image analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge