"Information": models, code, and papers
Speaker-Text Retrieval via Contrastive Learning
Dec 11, 2023In this study, we introduce a novel cross-modal retrieval task involving speaker descriptions and their corresponding audio samples. Utilizing pre-trained speaker and text encoders, we present a simple learning framework based on contrastive learning. Additionally, we explore the impact of incorporating speaker labels into the training process. Our findings establish the effectiveness of linking speaker and text information for the task for both English and Japanese languages, across diverse data configurations. Additional visual analysis unveils potential nuanced associations between speaker clustering and retrieval performance.
Age of Information Analysis for CR-NOMA Aided Uplink Systems with Randomly Arrived Packets
Nov 05, 2023This paper studies the application of cognitive radio inspired non-orthogonal multiple access (CR-NOMA) to reduce age of information (AoI) for uplink transmission. In particular, a time division multiple access (TDMA) based legacy network is considered, where each user is allocated with a dedicated time slot to transmit its status update information. The CR-NOMA is implemented as an add-on to the TDMA legacy network, which enables each user to have more opportunities to transmit by sharing other user's time slots. A rigorous analytical framework is developed to obtain the expressions for AoIs achieved by CR-NOMA with and without re-transmission, by taking the randomness of the status update generating process into consideration. Numerical results are presented to verify the accuracy of the developed analysis. It is shown that the AoI can be significantly reduced by applying CR-NOMA compared to TDMA. Moreover, the use of re-transmission is helpful to reduce AoI, especially when the status arrival rate is low.
Multi-view Inversion for 3D-aware Generative Adversarial Networks
Dec 08, 2023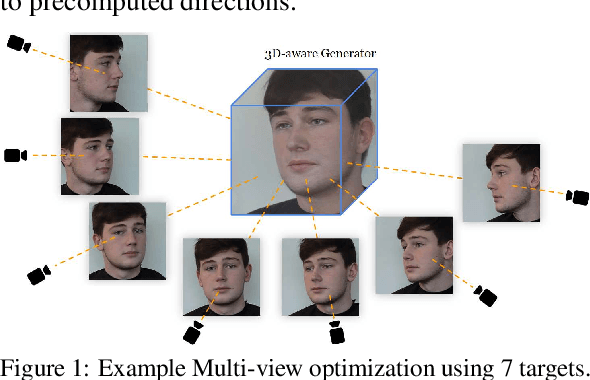
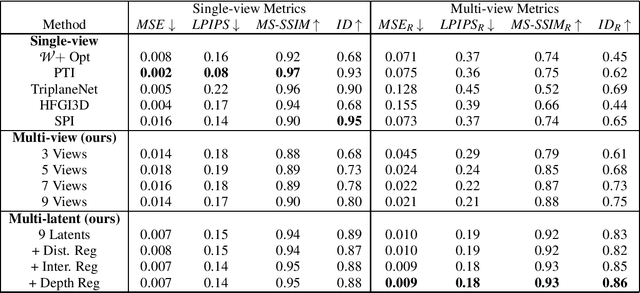

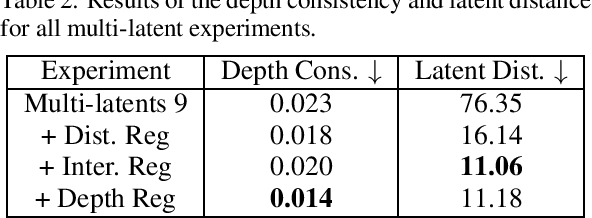
Current 3D GAN inversion methods for human heads typically use only one single frontal image to reconstruct the whole 3D head model. This leaves out meaningful information when multi-view data or dynamic videos are available. Our method builds on existing state-of-the-art 3D GAN inversion techniques to allow for consistent and simultaneous inversion of multiple views of the same subject. We employ a multi-latent extension to handle inconsistencies present in dynamic face videos to re-synthesize consistent 3D representations from the sequence. As our method uses additional information about the target subject, we observe significant enhancements in both geometric accuracy and image quality, particularly when rendering from wide viewing angles. Moreover, we demonstrate the editability of our inverted 3D renderings, which distinguishes them from NeRF-based scene reconstructions.
Graph-based Trajectory Prediction with Cooperative Information
Oct 24, 2023For automated driving, predicting the future trajectories of other road users in complex traffic situations is a hard problem. Modern neural networks use the past trajectories of traffic participants as well as map data to gather hints about the possible driver intention and likely maneuvers. With increasing connectivity between cars and other traffic actors, cooperative information is another source of data that can be used as inputs for trajectory prediction algorithms. Connected actors might transmit their intended path or even complete planned trajectories to other actors, which simplifies the prediction problem due to the imposed constraints. In this work, we outline the benefits of using this source of data for trajectory prediction and propose a graph-based neural network architecture that can leverage this additional data. We show that the network performance increases substantially if cooperative data is present. Also, our proposed training scheme improves the network's performance even for cases where no cooperative information is available. We also show that the network can deal with inaccurate cooperative data, which allows it to be used in real automated driving environments.
CoCoGen: Physically-Consistent and Conditioned Score-based Generative Models for Forward and Inverse Problems
Dec 16, 2023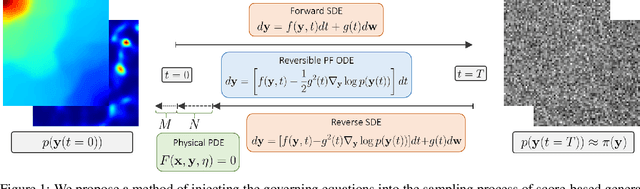
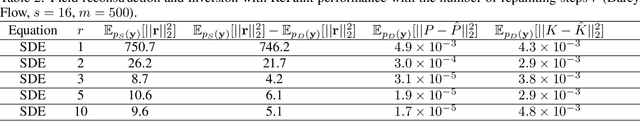
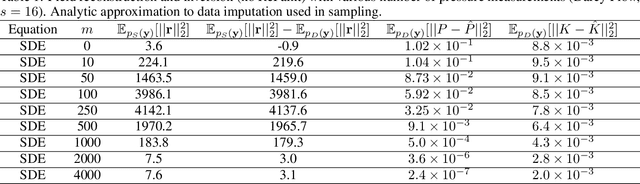
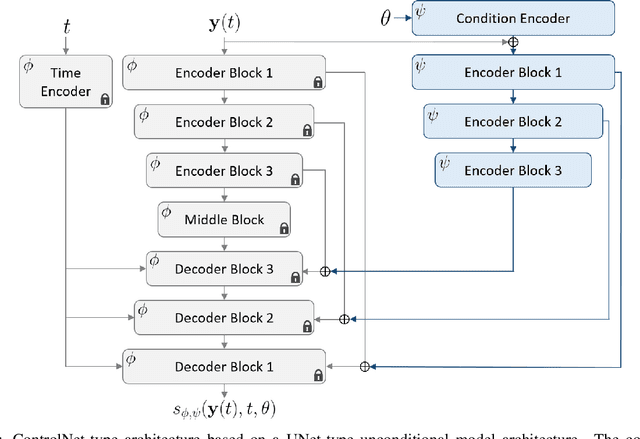

Recent advances in generative artificial intelligence have had a significant impact on diverse domains spanning computer vision, natural language processing, and drug discovery. This work extends the reach of generative models into physical problem domains, particularly addressing the efficient enforcement of physical laws and conditioning for forward and inverse problems involving partial differential equations (PDEs). Our work introduces two key contributions: firstly, we present an efficient approach to promote consistency with the underlying PDE. By incorporating discretized information into score-based generative models, our method generates samples closely aligned with the true data distribution, showcasing residuals comparable to data generated through conventional PDE solvers, significantly enhancing fidelity. Secondly, we showcase the potential and versatility of score-based generative models in various physics tasks, specifically highlighting surrogate modeling as well as probabilistic field reconstruction and inversion from sparse measurements. A robust foundation is laid by designing unconditional score-based generative models that utilize reversible probability flow ordinary differential equations. Leveraging conditional models that require minimal training, we illustrate their flexibility when combined with a frozen unconditional model. These conditional models generate PDE solutions by incorporating parameters, macroscopic quantities, or partial field measurements as guidance. The results illustrate the inherent flexibility of score-based generative models and explore the synergy between unconditional score-based generative models and the present physically-consistent sampling approach, emphasizing the power and flexibility in solving for and inverting physical fields governed by differential equations, and in other scientific machine learning tasks.
RIGHT: Retrieval-augmented Generation for Mainstream Hashtag Recommendation
Dec 16, 2023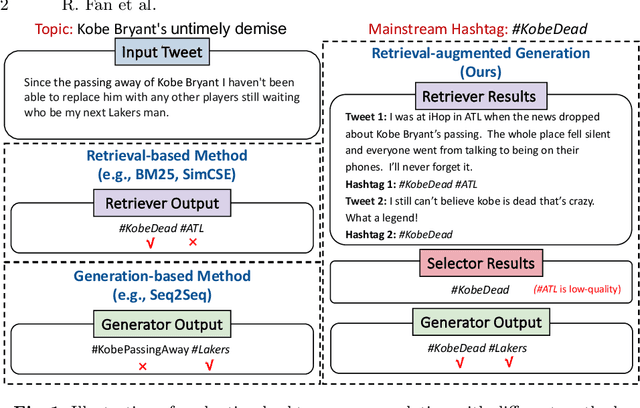
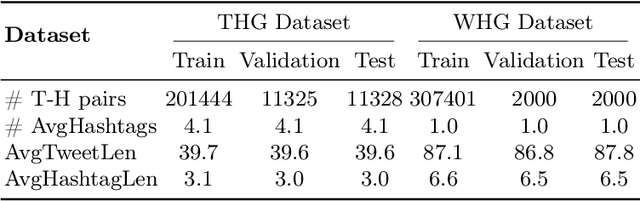
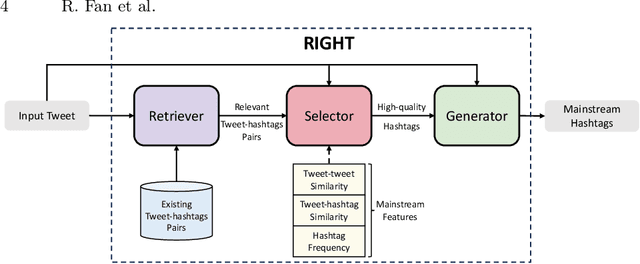

Automatic mainstream hashtag recommendation aims to accurately provide users with concise and popular topical hashtags before publication. Generally, mainstream hashtag recommendation faces challenges in the comprehensive difficulty of newly posted tweets in response to new topics, and the accurate identification of mainstream hashtags beyond semantic correctness. However, previous retrieval-based methods based on a fixed predefined mainstream hashtag list excel in producing mainstream hashtags, but fail to understand the constant flow of up-to-date information. Conversely, generation-based methods demonstrate a superior ability to comprehend newly posted tweets, but their capacity is constrained to identifying mainstream hashtags without additional features. Inspired by the recent success of the retrieval-augmented technique, in this work, we attempt to adopt this framework to combine the advantages of both approaches. Meantime, with the help of the generator component, we could rethink how to further improve the quality of the retriever component at a low cost. Therefore, we propose RetrIeval-augmented Generative Mainstream HashTag Recommender (RIGHT), which consists of three components: 1) a retriever seeks relevant hashtags from the entire tweet-hashtags set; 2) a selector enhances mainstream identification by introducing global signals; and 3) a generator incorporates input tweets and selected hashtags to directly generate the desired hashtags. The experimental results show that our method achieves significant improvements over state-of-the-art baselines. Moreover, RIGHT can be easily integrated into large language models, improving the performance of ChatGPT by more than 10%.
scBiGNN: Bilevel Graph Representation Learning for Cell Type Classification from Single-cell RNA Sequencing Data
Dec 16, 2023Single-cell RNA sequencing (scRNA-seq) technology provides high-throughput gene expression data to study the cellular heterogeneity and dynamics of complex organisms. Graph neural networks (GNNs) have been widely used for automatic cell type classification, which is a fundamental problem to solve in scRNA-seq analysis. However, existing methods do not sufficiently exploit both gene-gene and cell-cell relationships, and thus the true potential of GNNs is not realized. In this work, we propose a bilevel graph representation learning method, named scBiGNN, to simultaneously mine the relationships at both gene and cell levels for more accurate single-cell classification. Specifically, scBiGNN comprises two GNN modules to identify cell types. A gene-level GNN is established to adaptively learn gene-gene interactions and cell representations via the self-attention mechanism, and a cell-level GNN builds on the cell-cell graph that is constructed from the cell representations generated by the gene-level GNN. To tackle the scalability issue for processing a large number of cells, scBiGNN adopts an Expectation Maximization (EM) framework in which the two modules are alternately trained via the E-step and M-step to learn from each other. Through this interaction, the gene- and cell-level structural information is integrated to gradually enhance the classification performance of both GNN modules. Experiments on benchmark datasets demonstrate that our scBiGNN outperforms a variety of existing methods for cell type classification from scRNA-seq data.
CLIPSyntel: CLIP and LLM Synergy for Multimodal Question Summarization in Healthcare
Dec 16, 2023In the era of modern healthcare, swiftly generating medical question summaries is crucial for informed and timely patient care. Despite the increasing complexity and volume of medical data, existing studies have focused solely on text-based summarization, neglecting the integration of visual information. Recognizing the untapped potential of combining textual queries with visual representations of medical conditions, we introduce the Multimodal Medical Question Summarization (MMQS) Dataset. This dataset, a major contribution to our work, pairs medical queries with visual aids, facilitating a richer and more nuanced understanding of patient needs. We also propose a framework, utilizing the power of Contrastive Language Image Pretraining(CLIP) and Large Language Models(LLMs), consisting of four modules that identify medical disorders, generate relevant context, filter medical concepts, and craft visually aware summaries. Our comprehensive framework harnesses the power of CLIP, a multimodal foundation model, and various general-purpose LLMs, comprising four main modules: the medical disorder identification module, the relevant context generation module, the context filtration module for distilling relevant medical concepts and knowledge, and finally, a general-purpose LLM to generate visually aware medical question summaries. Leveraging our MMQS dataset, we showcase how visual cues from images enhance the generation of medically nuanced summaries. This multimodal approach not only enhances the decision-making process in healthcare but also fosters a more nuanced understanding of patient queries, laying the groundwork for future research in personalized and responsive medical care
Sentiment Analysis and Text Analysis of the Public Discourse on Twitter about COVID-19 and MPox
Dec 17, 2023
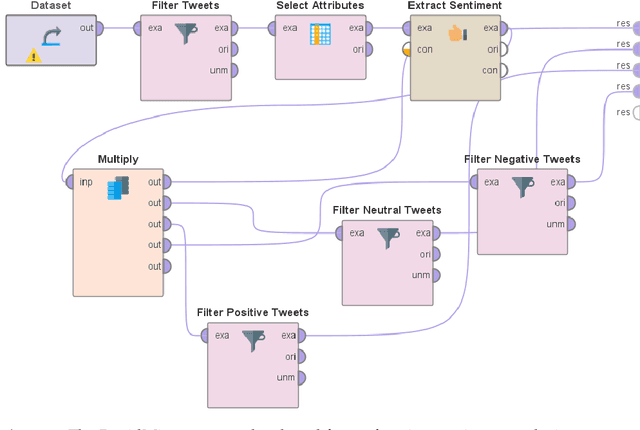

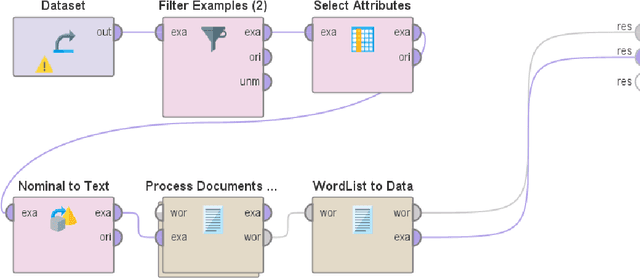
Mining and analysis of the big data of Twitter conversations have been of significant interest to the scientific community in the fields of healthcare, epidemiology, big data, data science, computer science, and their related areas, as can be seen from several works in the last few years that focused on sentiment analysis and other forms of text analysis of tweets related to Ebola, E-Coli, Dengue, Human Papillomavirus, Middle East Respiratory Syndrome, Measles, Zika virus, H1N1, influenza like illness, swine flu, flu, Cholera, Listeriosis, cancer, Liver Disease, Inflammatory Bowel Disease, kidney disease, lupus, Parkinsons, Diphtheria, and West Nile virus. The recent outbreaks of COVID-19 and MPox have served as catalysts for Twitter usage related to seeking and sharing information, views, opinions, and sentiments involving both of these viruses. None of the prior works in this field analyzed tweets focusing on both COVID-19 and MPox simultaneously. To address this research gap, a total of 61,862 tweets that focused on MPox and COVID-19 simultaneously, posted between 7 May 2022 and 3 March 2023, were studied. The findings and contributions of this study are manifold. First, the results of sentiment analysis using the VADER approach show that nearly half the tweets had a negative sentiment. It was followed by tweets that had a positive sentiment and tweets that had a neutral sentiment, respectively. Second, this paper presents the top 50 hashtags used in these tweets. Third, it presents the top 100 most frequently used words in these tweets after performing tokenization, removal of stopwords, and word frequency analysis. Finally, a comprehensive comparative study that compares the contributions of this paper with 49 prior works in this field is presented to further uphold the relevance and novelty of this work.
Self-MI: Efficient Multimodal Fusion via Self-Supervised Multi-Task Learning with Auxiliary Mutual Information Maximization
Nov 07, 2023Multimodal representation learning poses significant challenges in capturing informative and distinct features from multiple modalities. Existing methods often struggle to exploit the unique characteristics of each modality due to unified multimodal annotations. In this study, we propose Self-MI in the self-supervised learning fashion, which also leverage Contrastive Predictive Coding (CPC) as an auxiliary technique to maximize the Mutual Information (MI) between unimodal input pairs and the multimodal fusion result with unimodal inputs. Moreover, we design a label generation module, $ULG_{MI}$ for short, that enables us to create meaningful and informative labels for each modality in a self-supervised manner. By maximizing the Mutual Information, we encourage better alignment between the multimodal fusion and the individual modalities, facilitating improved multimodal fusion. Extensive experiments on three benchmark datasets including CMU-MOSI, CMU-MOSEI, and SIMS, demonstrate the effectiveness of Self-MI in enhancing the multimodal fusion task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge