"Image": models, code, and papers
Multimodal Graph Learning for Generative Tasks
Oct 11, 2023
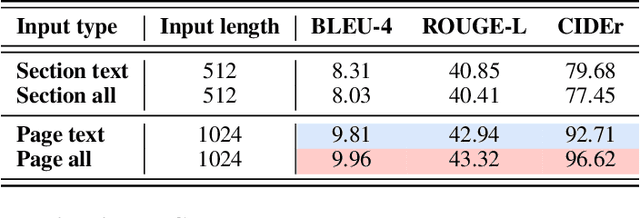
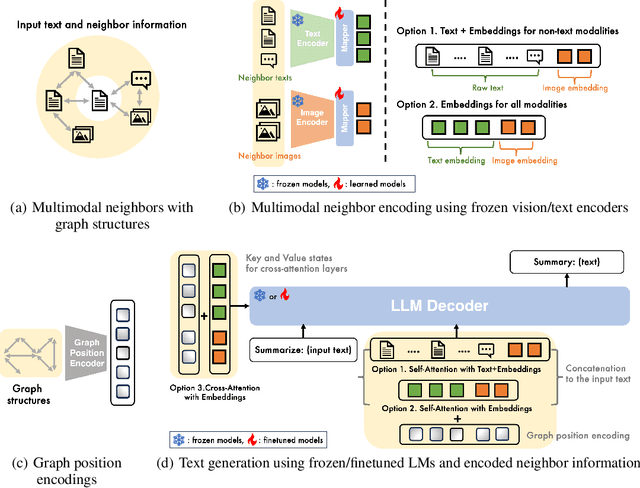
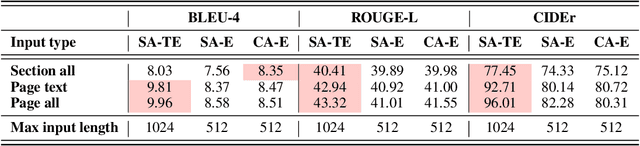
Multimodal learning combines multiple data modalities, broadening the types and complexity of data our models can utilize: for example, from plain text to image-caption pairs. Most multimodal learning algorithms focus on modeling simple one-to-one pairs of data from two modalities, such as image-caption pairs, or audio-text pairs. However, in most real-world settings, entities of different modalities interact with each other in more complex and multifaceted ways, going beyond one-to-one mappings. We propose to represent these complex relationships as graphs, allowing us to capture data with any number of modalities, and with complex relationships between modalities that can flexibly vary from one sample to another. Toward this goal, we propose Multimodal Graph Learning (MMGL), a general and systematic framework for capturing information from multiple multimodal neighbors with relational structures among them. In particular, we focus on MMGL for generative tasks, building upon pretrained Language Models (LMs), aiming to augment their text generation with multimodal neighbor contexts. We study three research questions raised by MMGL: (1) how can we infuse multiple neighbor information into the pretrained LMs, while avoiding scalability issues? (2) how can we infuse the graph structure information among multimodal neighbors into the LMs? and (3) how can we finetune the pretrained LMs to learn from the neighbor context in a parameter-efficient manner? We conduct extensive experiments to answer these three questions on MMGL and analyze the empirical results to pave the way for future MMGL research.
Attention-Map Augmentation for Hypercomplex Breast Cancer Classification
Oct 11, 2023Breast cancer is the most widespread neoplasm among women and early detection of this disease is critical. Deep learning techniques have become of great interest to improve diagnostic performance. Nonetheless, discriminating between malignant and benign masses from whole mammograms remains challenging due to them being almost identical to an untrained eye and the region of interest (ROI) occupying a minuscule portion of the entire image. In this paper, we propose a framework, parameterized hypercomplex attention maps (PHAM), to overcome these problems. Specifically, we deploy an augmentation step based on computing attention maps. Then, the attention maps are used to condition the classification step by constructing a multi-dimensional input comprised of the original breast cancer image and the corresponding attention map. In this step, a parameterized hypercomplex neural network (PHNN) is employed to perform breast cancer classification. The framework offers two main advantages. First, attention maps provide critical information regarding the ROI and allow the neural model to concentrate on it. Second, the hypercomplex architecture has the ability to model local relations between input dimensions thanks to hypercomplex algebra rules, thus properly exploiting the information provided by the attention map. We demonstrate the efficacy of the proposed framework on both mammography images as well as histopathological ones, surpassing attention-based state-of-the-art networks and the real-valued counterpart of our method. The code of our work is available at https://github.com/elelo22/AttentionBCS.
Minimalist and High-Performance Semantic Segmentation with Plain Vision Transformers
Oct 19, 2023In the wake of Masked Image Modeling (MIM), a diverse range of plain, non-hierarchical Vision Transformer (ViT) models have been pre-trained with extensive datasets, offering new paradigms and significant potential for semantic segmentation. Current state-of-the-art systems incorporate numerous inductive biases and employ cumbersome decoders. Building upon the original motivations of plain ViTs, which are simplicity and generality, we explore high-performance `minimalist' systems to this end. Our primary purpose is to provide simple and efficient baselines for practical semantic segmentation with plain ViTs. Specifically, we first explore the feasibility and methodology for achieving high-performance semantic segmentation using the last feature map. As a result, we introduce the PlainSeg, a model comprising only three 3$\times$3 convolutions in addition to the transformer layers (either encoder or decoder). In this process, we offer insights into two underlying principles: (i) high-resolution features are crucial to high performance in spite of employing simple up-sampling techniques and (ii) the slim transformer decoder requires a much larger learning rate than the wide transformer decoder. On this basis, we further present the PlainSeg-Hier, which allows for the utilization of hierarchical features. Extensive experiments on four popular benchmarks demonstrate the high performance and efficiency of our methods. They can also serve as powerful tools for assessing the transfer ability of base models in semantic segmentation. Code is available at \url{https://github.com/ydhongHIT/PlainSeg}.
MIML: Multiplex Image Machine Learning for High Precision Cell Classification via Mechanical Traits within Microfluidic Systems
Sep 15, 2023Label-free cell classification is advantageous for supplying pristine cells for further use or examination, yet existing techniques frequently fall short in terms of specificity and speed. In this study, we address these limitations through the development of a novel machine learning framework, Multiplex Image Machine Learning (MIML). This architecture uniquely combines label-free cell images with biomechanical property data, harnessing the vast, often underutilized morphological information intrinsic to each cell. By integrating both types of data, our model offers a more holistic understanding of the cellular properties, utilizing morphological information typically discarded in traditional machine learning models. This approach has led to a remarkable 98.3\% accuracy in cell classification, a substantial improvement over models that only consider a single data type. MIML has been proven effective in classifying white blood cells and tumor cells, with potential for broader application due to its inherent flexibility and transfer learning capability. It's particularly effective for cells with similar morphology but distinct biomechanical properties. This innovative approach has significant implications across various fields, from advancing disease diagnostics to understanding cellular behavior.
Foundation Model-oriented Robustness: Robust Image Model Evaluation with Pretrained Models
Aug 23, 2023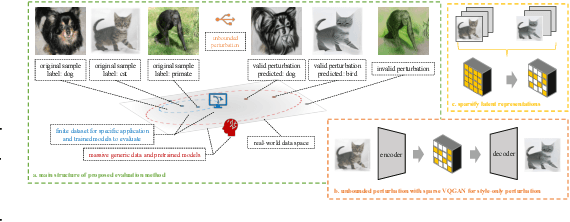
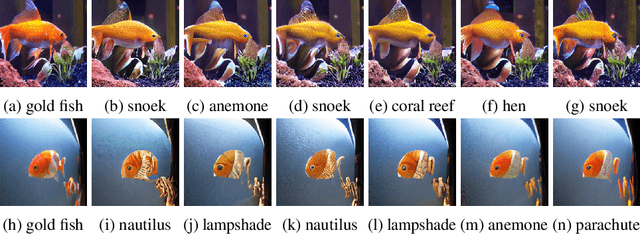
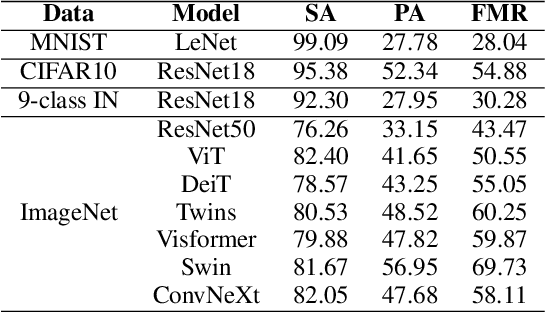
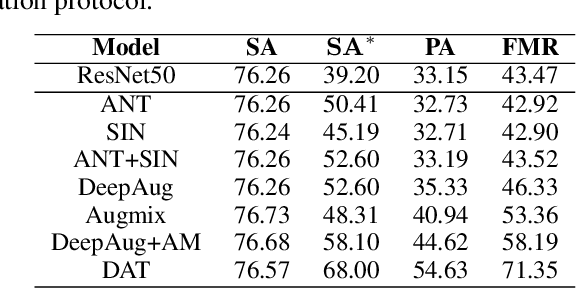

Machine learning has demonstrated remarkable performance over finite datasets, yet whether the scores over the fixed benchmarks can sufficiently indicate the model's performance in the real world is still in discussion. In reality, an ideal robust model will probably behave similarly to the oracle (e.g., the human users), thus a good evaluation protocol is probably to evaluate the models' behaviors in comparison to the oracle. In this paper, we introduce a new robustness measurement that directly measures the image classification model's performance compared with a surrogate oracle (i.e., a foundation model). Besides, we design a simple method that can accomplish the evaluation beyond the scope of the benchmarks. Our method extends the image datasets with new samples that are sufficiently perturbed to be distinct from the ones in the original sets, but are still bounded within the same image-label structure the original test image represents, constrained by a foundation model pretrained with a large amount of samples. As a result, our new method will offer us a new way to evaluate the models' robustness performance, free of limitations of fixed benchmarks or constrained perturbations, although scoped by the power of the oracle. In addition to the evaluation results, we also leverage our generated data to understand the behaviors of the model and our new evaluation strategies.
MetaWeather: Few-Shot Weather-Degraded Image Restoration via Degradation Pattern Matching
Aug 28, 2023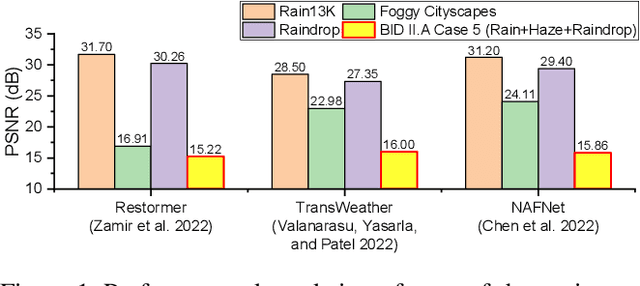
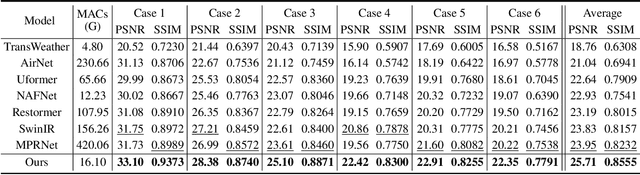
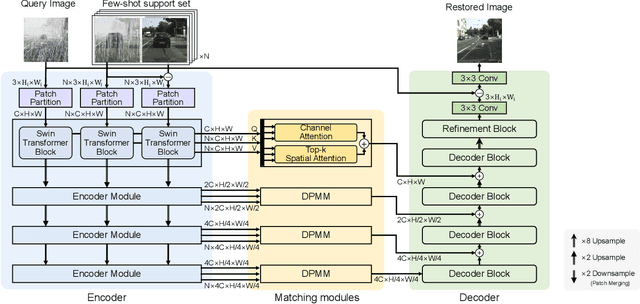
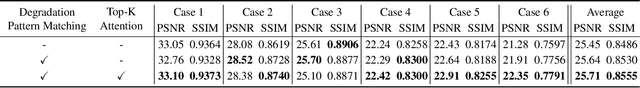
Real-world vision tasks frequently suffer from the appearance of adverse weather conditions including rain, fog, snow, and raindrops in captured images. Recently, several generic methods for restoring weather-degraded images have been proposed, aiming to remove multiple types of adverse weather effects present in the images. However, these methods have considered weather as discrete and mutually exclusive variables, leading to failure in generalizing to unforeseen weather conditions beyond the scope of the training data, such as the co-occurrence of rain, fog, and raindrops. To this end, weather-degraded image restoration models should have flexible adaptability to the current unknown weather condition to ensure reliable and optimal performance. The adaptation method should also be able to cope with data scarcity for real-world adaptation. This paper proposes MetaWeather, a few-shot weather-degraded image restoration method for arbitrary weather conditions. For this, we devise the core piece of MetaWeather, coined Degradation Pattern Matching Module (DPMM), which leverages representations from a few-shot support set by matching features between input and sample images under new weather conditions. In addition, we build meta-knowledge with episodic meta-learning on top of our MetaWeather architecture to provide flexible adaptability. In the meta-testing phase, we adopt a parameter-efficient fine-tuning method to preserve the prebuilt knowledge and avoid the overfitting problem. Experiments on the BID Task II.A dataset show our method achieves the best performance on PSNR and SSIM compared to state-of-the-art image restoration methods. Code is available at (TBA).
On the Localization of Ultrasound Image Slices within Point Distribution Models
Sep 01, 2023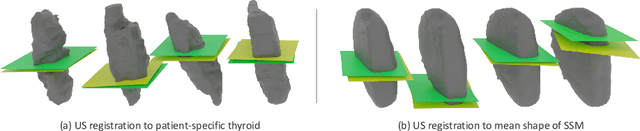
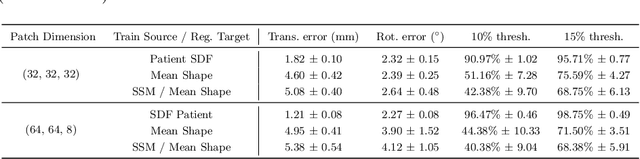
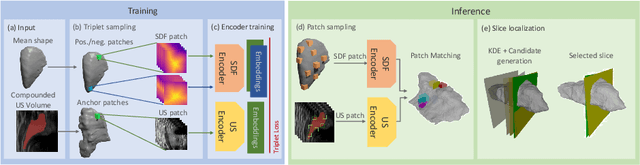
Thyroid disorders are most commonly diagnosed using high-resolution Ultrasound (US). Longitudinal nodule tracking is a pivotal diagnostic protocol for monitoring changes in pathological thyroid morphology. This task, however, imposes a substantial cognitive load on clinicians due to the inherent challenge of maintaining a mental 3D reconstruction of the organ. We thus present a framework for automated US image slice localization within a 3D shape representation to ease how such sonographic diagnoses are carried out. Our proposed method learns a common latent embedding space between US image patches and the 3D surface of an individual's thyroid shape, or a statistical aggregation in the form of a statistical shape model (SSM), via contrastive metric learning. Using cross-modality registration and Procrustes analysis, we leverage features from our model to register US slices to a 3D mesh representation of the thyroid shape. We demonstrate that our multi-modal registration framework can localize images on the 3D surface topology of a patient-specific organ and the mean shape of an SSM. Experimental results indicate slice positions can be predicted within an average of 1.2 mm of the ground-truth slice location on the patient-specific 3D anatomy and 4.6 mm on the SSM, exemplifying its usefulness for slice localization during sonographic acquisitions. Code is publically available: \href{https://github.com/vuenc/slice-to-shape}{https://github.com/vuenc/slice-to-shape}
MarineGPT: Unlocking Secrets of Ocean to the Public
Oct 20, 2023Large language models (LLMs), such as ChatGPT/GPT-4, have proven to be powerful tools in promoting the user experience as an AI assistant. The continuous works are proposing multi-modal large language models (MLLM), empowering LLMs with the ability to sense multiple modality inputs through constructing a joint semantic space (e.g. visual-text space). Though significant success was achieved in LLMs and MLLMs, exploring LLMs and MLLMs in domain-specific applications that required domain-specific knowledge and expertise has been less conducted, especially for \textbf{marine domain}. Different from general-purpose MLLMs, the marine-specific MLLM is required to yield much more \textbf{sensitive}, \textbf{informative}, and \textbf{scientific} responses. In this work, we demonstrate that the existing MLLMs optimized on huge amounts of readily available general-purpose training data show a minimal ability to understand domain-specific intents and then generate informative and satisfactory responses. To address these issues, we propose \textbf{MarineGPT}, the first vision-language model specially designed for the marine domain, unlocking the secrets of the ocean to the public. We present our \textbf{Marine-5M} dataset with more than 5 million marine image-text pairs to inject domain-specific marine knowledge into our model and achieve better marine vision and language alignment. Our MarineGPT not only pushes the boundaries of marine understanding to the general public but also offers a standard protocol for adapting a general-purpose assistant to downstream domain-specific experts. We pave the way for a wide range of marine applications while setting valuable data and pre-trained models for future research in both academic and industrial communities.
Multimodal Object Query Initialization for 3D Object Detection
Oct 16, 20233D object detection models that exploit both LiDAR and camera sensor features are top performers in large-scale autonomous driving benchmarks. A transformer is a popular network architecture used for this task, in which so-called object queries act as candidate objects. Initializing these object queries based on current sensor inputs is a common practice. For this, existing methods strongly rely on LiDAR data however, and do not fully exploit image features. Besides, they introduce significant latency. To overcome these limitations we propose EfficientQ3M, an efficient, modular, and multimodal solution for object query initialization for transformer-based 3D object detection models. The proposed initialization method is combined with a "modality-balanced" transformer decoder where the queries can access all sensor modalities throughout the decoder. In experiments, we outperform the state of the art in transformer-based LiDAR object detection on the competitive nuScenes benchmark and showcase the benefits of input-dependent multimodal query initialization, while being more efficient than the available alternatives for LiDAR-camera initialization. The proposed method can be applied with any combination of sensor modalities as input, demonstrating its modularity.
Style transfer between Microscopy and Magnetic Resonance Imaging via Generative Adversarial Network in small sample size settings
Oct 16, 2023Cross-modal augmentation of Magnetic Resonance Imaging (MRI) and microscopic imaging based on the same tissue samples is promising because it can allow histopathological analysis in the absence of an underlying invasive biopsy procedure. Here, we tested a method for generating microscopic histological images from MRI scans of the corpus callosum using conditional generative adversarial network (cGAN) architecture. To our knowledge, this is the first multimodal translation of the brain MRI to histological volumetric representation of the same sample. The technique was assessed by training paired image translation models taking sets of images from MRI scans and microscopy. The use of cGAN for this purpose is challenging because microscopy images are large in size and typically have low sample availability. The current work demonstrates that the framework reliably synthesizes histology images from MRI scans of corpus callosum, emphasizing the network's ability to train on high resolution histologies paired with relatively lower-resolution MRI scans. With the ultimate goal of avoiding biopsies, the proposed tool can be used for educational purposes.
* 2023 IEEE International Conference on Image Processing (ICIP)
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge