"Image": models, code, and papers
Hard-Attention for Scalable Image Classification
Feb 20, 2021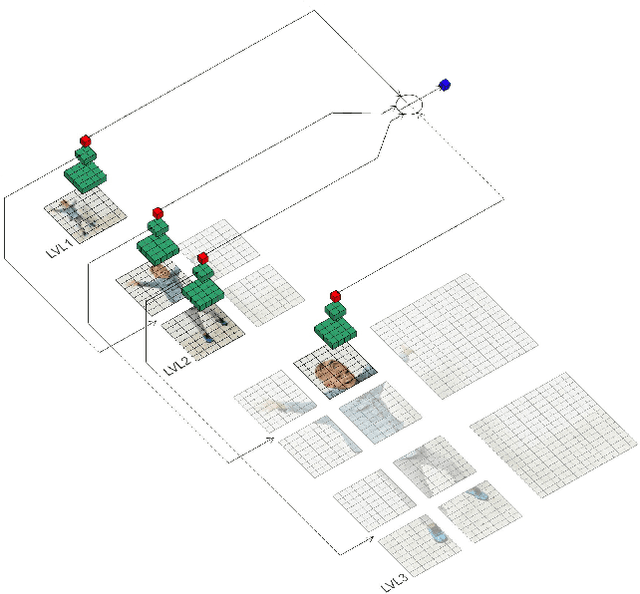
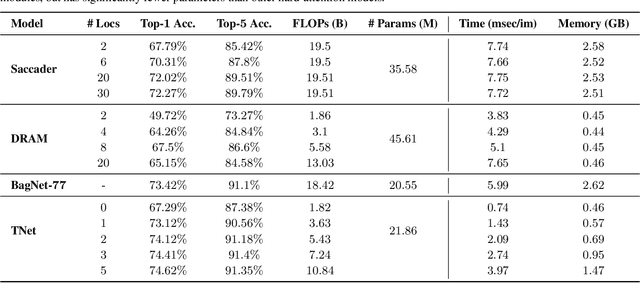
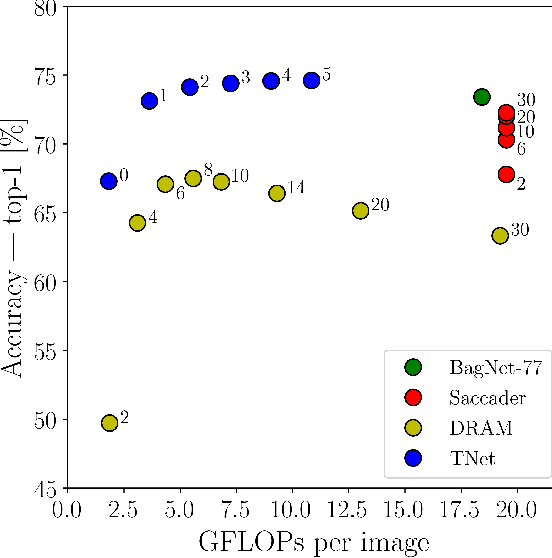
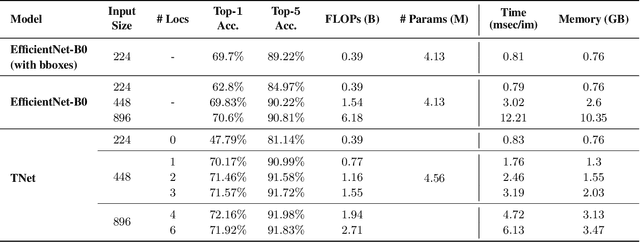
Deep neural networks (DNNs) are typically optimized for a specific input resolution (e.g. $224 \times 224$ px) and their adoption to inputs of higher resolution (e.g., satellite or medical images) remains challenging, as it leads to excessive computation and memory overhead, and may require substantial engineering effort (e.g., streaming). We show that multi-scale hard-attention can be an effective solution to this problem. We propose a novel architecture, TNet, which traverses an image pyramid in a top-down fashion, visiting only the most informative regions along the way. We compare our model against strong hard-attention baselines, achieving a better trade-off between resources and accuracy on ImageNet. We further verify the efficacy of our model on satellite images (fMoW dataset) of size up to $896 \times 896$ px. In addition, our hard-attention mechanism guarantees predictions with a degree of interpretability, without extra cost beyond inference. We also show that we can reduce data acquisition and annotation cost, since our model attends only to a fraction of the highest resolution content, while using only image-level labels without bounding boxes.
Unsupervised Deep Learning Methods for Biological Image Reconstruction
May 17, 2021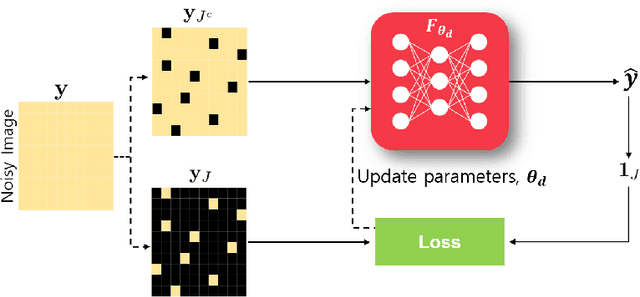
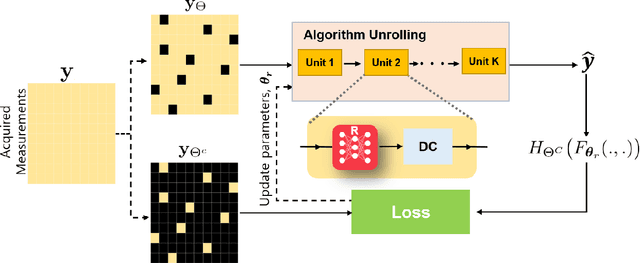
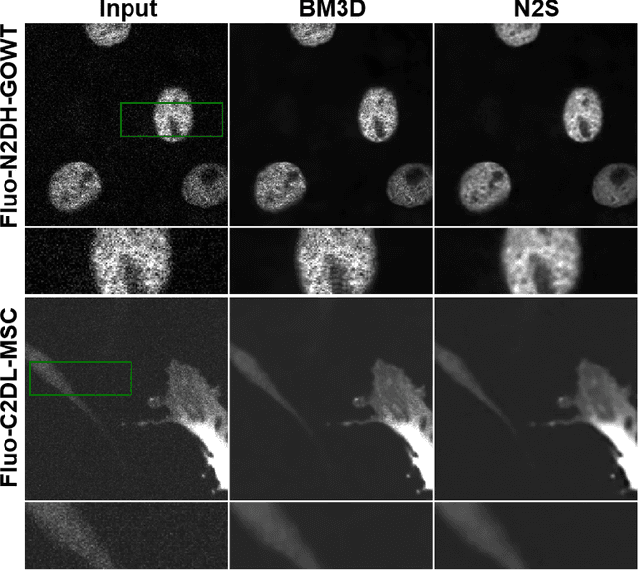
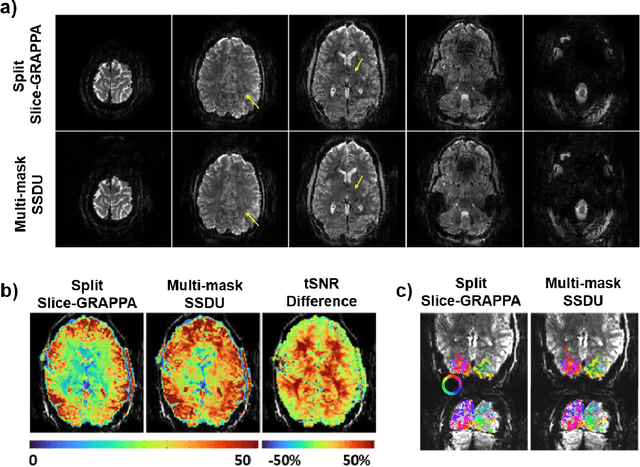
Recently, deep learning approaches have become the main research frontier for biological image reconstruction problems thanks to their high performance, along with their ultra-fast reconstruction times. However, due to the difficulty of obtaining matched reference data for supervised learning, there has been increasing interest in unsupervised learning approaches that do not need paired reference data. In particular, self-supervised learning and generative models have been successfully used for various biological imaging applications. In this paper, we overview these approaches from a coherent perspective in the context of classical inverse problems, and discuss their applications to biological imaging.
A Dempster-Shafer approach to trustworthy AI with application to fetal brain MRI segmentation
Apr 05, 2022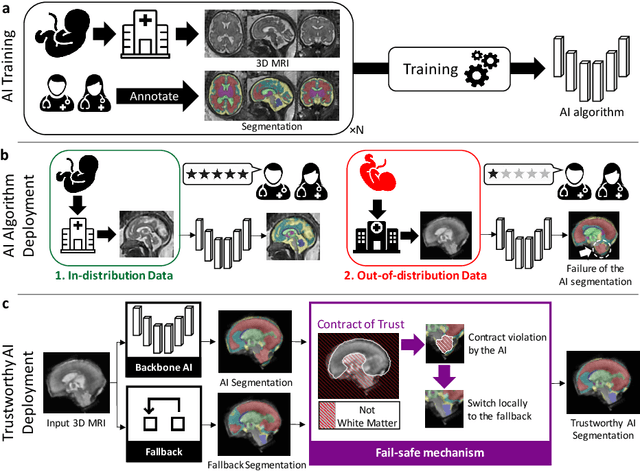
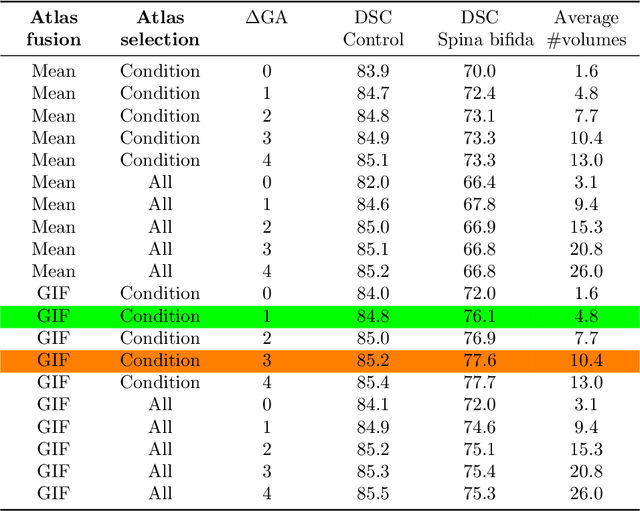
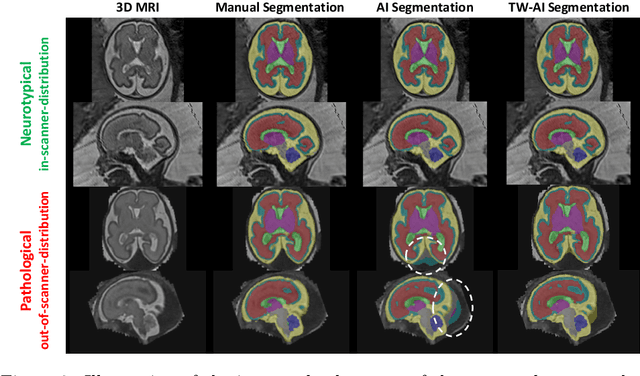
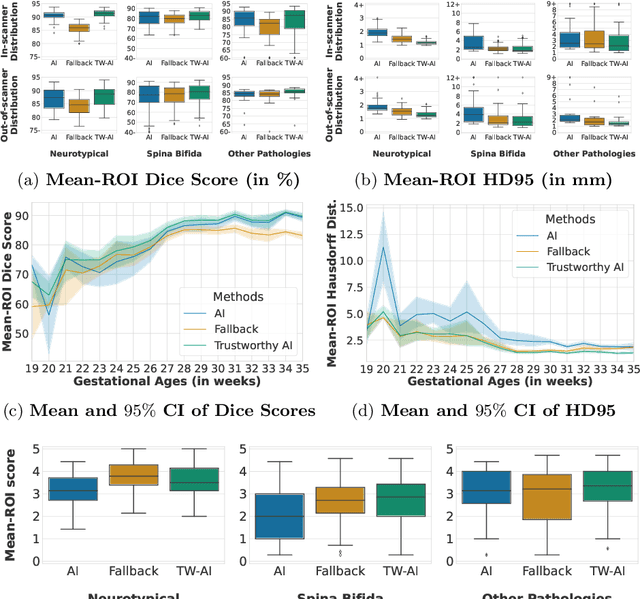
Deep learning models for medical image segmentation can fail unexpectedly and spectacularly for pathological cases and for images acquired at different centers than those used for training, with labeling errors that violate expert knowledge about the anatomy and the intensity distribution of the regions to be segmented. Such errors undermine the trustworthiness of deep learning models developed for medical image segmentation. Mechanisms with a fallback method for detecting and correcting such failures are essential for safely translating this technology into clinics and are likely to be a requirement of future regulations on artificial intelligence (AI). Here, we propose a principled trustworthy AI theoretical framework and a practical system that can augment any backbone AI system using a fallback method and a fail-safe mechanism based on Dempster-Shafer theory. Our approach relies on an actionable definition of trustworthy AI. Our method automatically discards the voxel-level labeling predicted by the backbone AI that are likely to violate expert knowledge and relies on a fallback atlas-based segmentation method for those voxels. We demonstrate the effectiveness of the proposed trustworthy AI approach on the largest reported annotated dataset of fetal T2w MRI consisting of 540 manually annotated fetal brain 3D MRIs with neurotypical or abnormal brain development and acquired from 13 sources of data across 6 countries. We show that our trustworthy AI method improves the robustness of a state-of-the-art backbone AI for fetal brain MRI segmentation on MRIs acquired across various centers and for fetuses with various brain abnormalities.
Graph Convolutional Networks in Feature Space for Image Deblurring and Super-resolution
May 21, 2021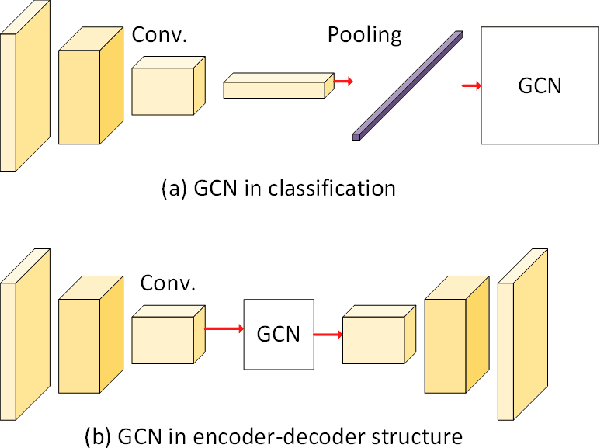
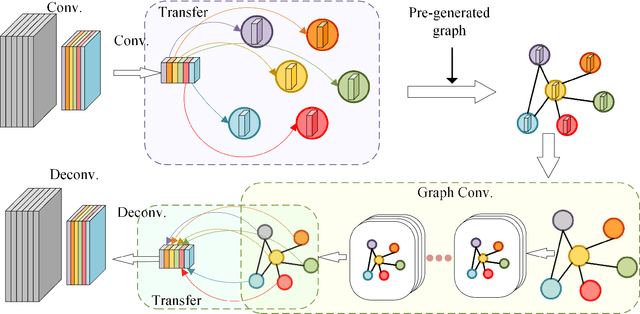
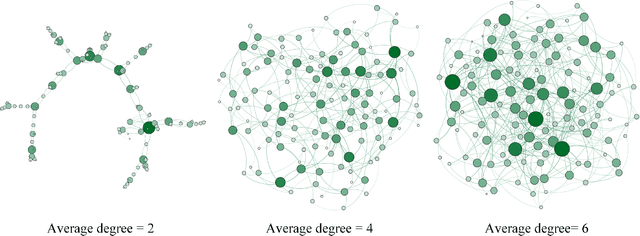

Graph convolutional networks (GCNs) have achieved great success in dealing with data of non-Euclidean structures. Their success directly attributes to fitting graph structures effectively to data such as in social media and knowledge databases. For image processing applications, the use of graph structures and GCNs have not been fully explored. In this paper, we propose a novel encoder-decoder network with added graph convolutions by converting feature maps to vertexes of a pre-generated graph to synthetically construct graph-structured data. By doing this, we inexplicitly apply graph Laplacian regularization to the feature maps, making them more structured. The experiments show that it significantly boosts performance for image restoration tasks, including deblurring and super-resolution. We believe it opens up opportunities for GCN-based approaches in more applications.
* Accepted by IJCNN 2021 (Oral)
Leveraging Deepfakes to Close the Domain Gap between Real and Synthetic Images in Facial Capture Pipelines
Apr 27, 2022



We propose an end-to-end pipeline for both building and tracking 3D facial models from personalized in-the-wild (cellphone, webcam, youtube clips, etc.) video data. First, we present a method for automatic data curation and retrieval based on a hierarchical clustering framework typical of collision detection algorithms in traditional computer graphics pipelines. Subsequently, we utilize synthetic turntables and leverage deepfake technology in order to build a synthetic multi-view stereo pipeline for appearance capture that is robust to imperfect synthetic geometry and image misalignment. The resulting model is fit with an animation rig, which is then used to track facial performances. Notably, our novel use of deepfake technology enables us to perform robust tracking of in-the-wild data using differentiable renderers despite a significant synthetic-to-real domain gap. Finally, we outline how we train a motion capture regressor, leveraging the aforementioned techniques to avoid the need for real-world ground truth data and/or a high-end calibrated camera capture setup.
The Tree Loss: Improving Generalization with Many Classes
Apr 16, 2022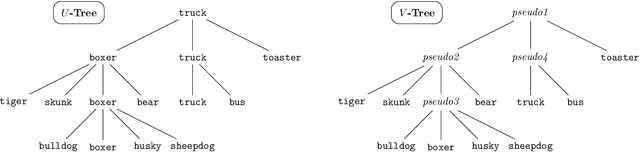
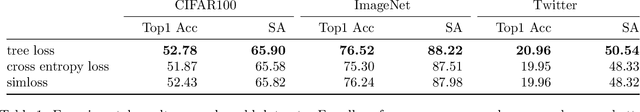
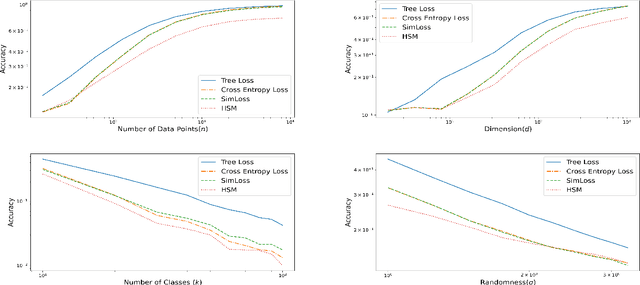
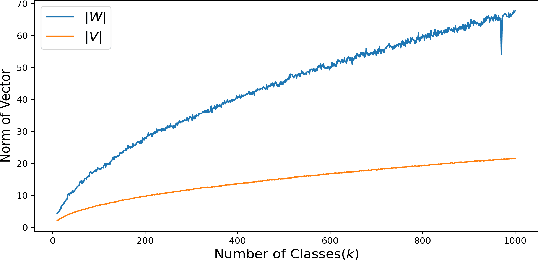
Multi-class classification problems often have many semantically similar classes. For example, 90 of ImageNet's 1000 classes are for different breeds of dog. We should expect that these semantically similar classes will have similar parameter vectors, but the standard cross entropy loss does not enforce this constraint. We introduce the tree loss as a drop-in replacement for the cross entropy loss. The tree loss re-parameterizes the parameter matrix in order to guarantee that semantically similar classes will have similar parameter vectors. Using simple properties of stochastic gradient descent, we show that the tree loss's generalization error is asymptotically better than the cross entropy loss's. We then validate these theoretical results on synthetic data, image data (CIFAR100, ImageNet), and text data (Twitter).
Is Geometry Enough for Matching in Visual Localization?
Mar 24, 2022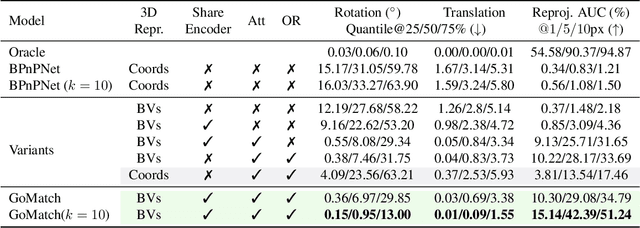
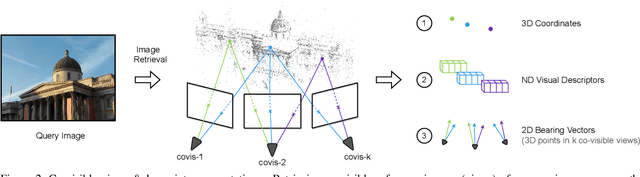
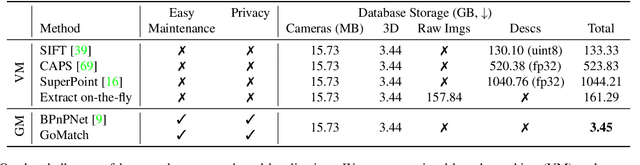
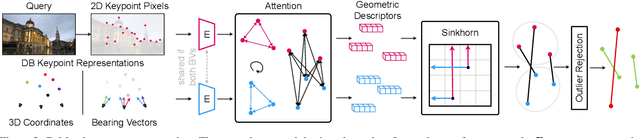
In this paper, we propose to go beyond the well-established approach to vision-based localization that relies on visual descriptor matching between a query image and a 3D point cloud. While matching keypoints via visual descriptors makes localization highly accurate, it has significant storage demands, raises privacy concerns and increases map maintenance complexity. To elegantly address those practical challenges for large-scale localization, we present GoMatch, an alternative to visual-based matching that solely relies on geometric information for matching image keypoints to maps, represented as sets of bearing vectors. Our novel bearing vectors representation of 3D points, significantly relieves the cross-domain challenge in geometric-based matching that prevented prior work to tackle localization in a realistic environment. With additional careful architecture design, GoMatch improves over prior geometric-based matching work with a reduction of ($10.67m, 95.7^{\circ}$) and ($1.43m$, $34.7^{\circ}$) in average median pose errors on Cambridge Landmarks and 7-Scenes, while requiring as little as $1.5/1.7\%$ of storage capacity in comparison to the best visual-based matching methods. This confirms its potential and feasibility for real-world localization and opens the door to future efforts in advancing city-scale visual localization methods that do not require storing visual descriptors.
Data--driven Image Restoration with Option--driven Learning for Big and Small Astronomical Image Datasets
Nov 07, 2020
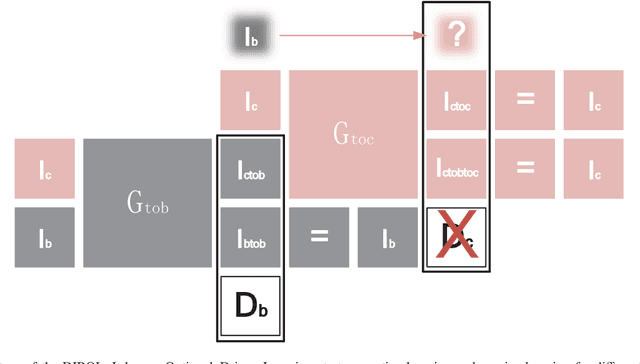
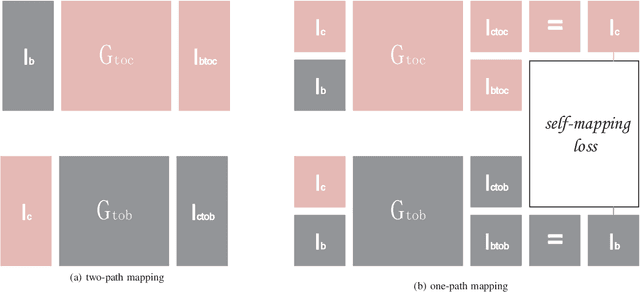

Image restoration methods are commonly used to improve the quality of astronomical images. In recent years, developments of deep neural networks and increments of the number of astronomical images have evoked a lot of data--driven image restoration methods. However, most of these methods belong to supervised learning algorithms, which require paired images either from real observations or simulated data as training set. For some applications, it is hard to get enough paired images from real observations and simulated images are quite different from real observed ones. In this paper, we propose a new data--driven image restoration method based on generative adversarial networks with option--driven learning. Our method uses several high resolution images as references and applies different learning strategies when the number of reference images is different. For sky surveys with variable observation conditions, our method can obtain very stable image restoration results, regardless of the number of reference images.
DivergentNets: Medical Image Segmentation by Network Ensemble
Jul 01, 2021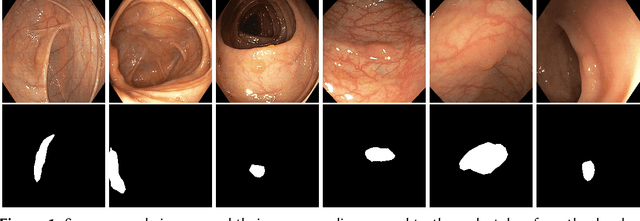
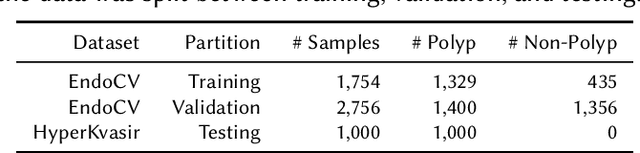
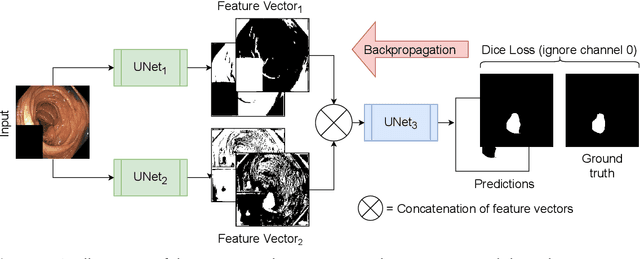
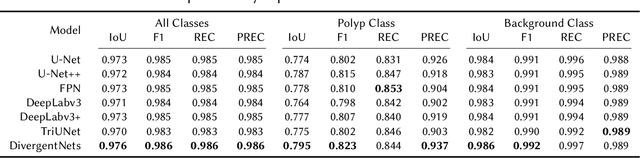
Detection of colon polyps has become a trending topic in the intersecting fields of machine learning and gastrointestinal endoscopy. The focus has mainly been on per-frame classification. More recently, polyp segmentation has gained attention in the medical community. Segmentation has the advantage of being more accurate than per-frame classification or object detection as it can show the affected area in greater detail. For our contribution to the EndoCV 2021 segmentation challenge, we propose two separate approaches. First, a segmentation model named TriUNet composed of three separate UNet models. Second, we combine TriUNet with an ensemble of well-known segmentation models, namely UNet++, FPN, DeepLabv3, and DeepLabv3+, into a model called DivergentNets to produce more generalizable medical image segmentation masks. In addition, we propose a modified Dice loss that calculates loss only for a single class when performing multiclass segmentation, forcing the model to focus on what is most important. Overall, the proposed methods achieved the best average scores for each respective round in the challenge, with TriUNet being the winning model in Round I and DivergentNets being the winning model in Round II of the segmentation generalization challenge at EndoCV 2021. The implementation of our approach is made publicly available on GitHub.
* the winning model of the segmentation generalization challenge at EndoCV 2021
CrossPoint: Self-Supervised Cross-Modal Contrastive Learning for 3D Point Cloud Understanding
Mar 24, 2022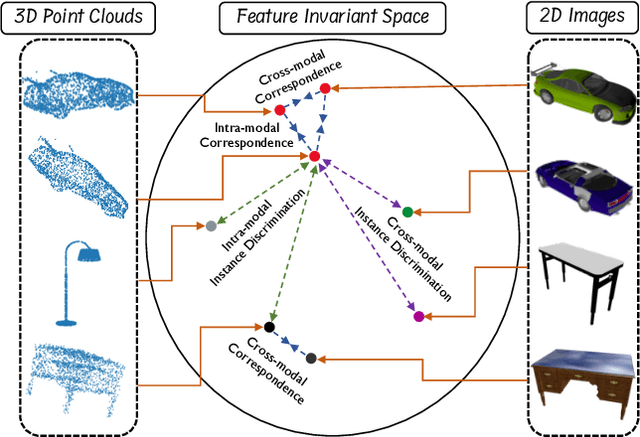
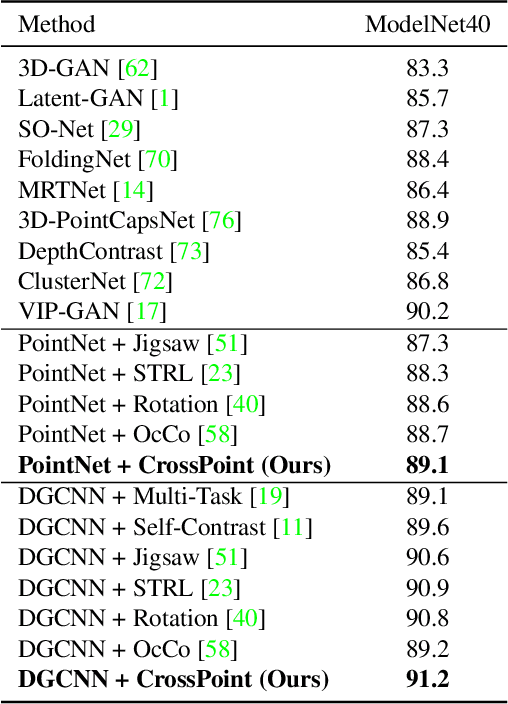
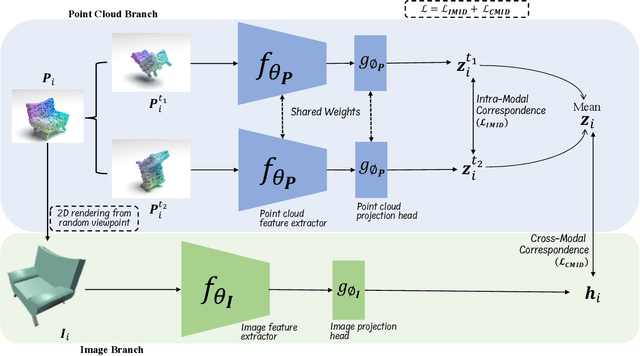
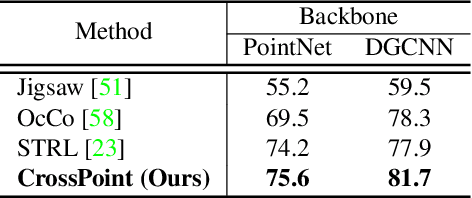
Manual annotation of large-scale point cloud dataset for varying tasks such as 3D object classification, segmentation and detection is often laborious owing to the irregular structure of point clouds. Self-supervised learning, which operates without any human labeling, is a promising approach to address this issue. We observe in the real world that humans are capable of mapping the visual concepts learnt from 2D images to understand the 3D world. Encouraged by this insight, we propose CrossPoint, a simple cross-modal contrastive learning approach to learn transferable 3D point cloud representations. It enables a 3D-2D correspondence of objects by maximizing agreement between point clouds and the corresponding rendered 2D image in the invariant space, while encouraging invariance to transformations in the point cloud modality. Our joint training objective combines the feature correspondences within and across modalities, thus ensembles a rich learning signal from both 3D point cloud and 2D image modalities in a self-supervised fashion. Experimental results show that our approach outperforms the previous unsupervised learning methods on a diverse range of downstream tasks including 3D object classification and segmentation. Further, the ablation studies validate the potency of our approach for a better point cloud understanding. Code and pretrained models are available at http://github.com/MohamedAfham/CrossPoint.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge