Yunhe Gao
VerSe: Integrating Multiple Queries as Prompts for Versatile Cardiac MRI Segmentation
Dec 20, 2024Abstract:Despite the advances in learning-based image segmentation approach, the accurate segmentation of cardiac structures from magnetic resonance imaging (MRI) remains a critical challenge. While existing automatic segmentation methods have shown promise, they still require extensive manual corrections of the segmentation results by human experts, particularly in complex regions such as the basal and apical parts of the heart. Recent efforts have been made on developing interactive image segmentation methods that enable human-in-the-loop learning. However, they are semi-automatic and inefficient, due to their reliance on click-based prompts, especially for 3D cardiac MRI volumes. To address these limitations, we propose VerSe, a Versatile Segmentation framework to unify automatic and interactive segmentation through mutiple queries. Our key innovation lies in the joint learning of object and click queries as prompts for a shared segmentation backbone. VerSe supports both fully automatic segmentation, through object queries, and interactive mask refinement, by providing click queries when needed. With the proposed integrated prompting scheme, VerSe demonstrates significant improvement in performance and efficiency over existing methods, on both cardiac MRI and out-of-distribution medical imaging datasets. The code is available at https://github.com/bangwayne/Verse.
Aligning Human Knowledge with Visual Concepts Towards Explainable Medical Image Classification
Jun 08, 2024


Abstract:Although explainability is essential in the clinical diagnosis, most deep learning models still function as black boxes without elucidating their decision-making process. In this study, we investigate the explainable model development that can mimic the decision-making process of human experts by fusing the domain knowledge of explicit diagnostic criteria. We introduce a simple yet effective framework, Explicd, towards Explainable language-informed criteria-based diagnosis. Explicd initiates its process by querying domain knowledge from either large language models (LLMs) or human experts to establish diagnostic criteria across various concept axes (e.g., color, shape, texture, or specific patterns of diseases). By leveraging a pretrained vision-language model, Explicd injects these criteria into the embedding space as knowledge anchors, thereby facilitating the learning of corresponding visual concepts within medical images. The final diagnostic outcome is determined based on the similarity scores between the encoded visual concepts and the textual criteria embeddings. Through extensive evaluation of five medical image classification benchmarks, Explicd has demonstrated its inherent explainability and extends to improve classification performance compared to traditional black-box models.
Implicit In-context Learning
May 23, 2024



Abstract:In-context Learning (ICL) empowers large language models (LLMs) to adapt to unseen tasks during inference by prefixing a few demonstration examples prior to test queries. Despite its versatility, ICL incurs substantial computational and memory overheads compared to zero-shot learning and is susceptible to the selection and order of demonstration examples. In this work, we introduce Implicit In-context Learning (I2CL), an innovative paradigm that addresses the challenges associated with traditional ICL by absorbing demonstration examples within the activation space. I2CL first generates a condensed vector representation, namely a context vector, from the demonstration examples. It then integrates the context vector during inference by injecting a linear combination of the context vector and query activations into the model's residual streams. Empirical evaluation on nine real-world tasks across three model architectures demonstrates that I2CL achieves few-shot performance with zero-shot cost and exhibits robustness against the variation of demonstration examples. Furthermore, I2CL facilitates a novel representation of "task-ids", enhancing task similarity detection and enabling effective transfer learning. We provide a comprehensive analysis of I2CL, offering deeper insights into its mechanisms and broader implications for ICL. The source code is available at: https://github.com/LzVv123456/I2CL.
Deep Deformable Models: Learning 3D Shape Abstractions with Part Consistency
Sep 02, 2023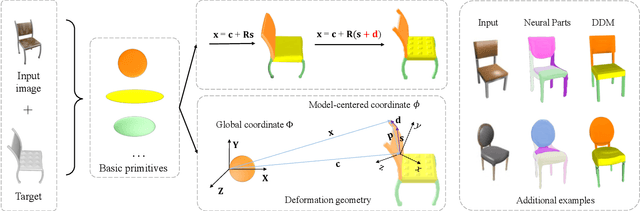
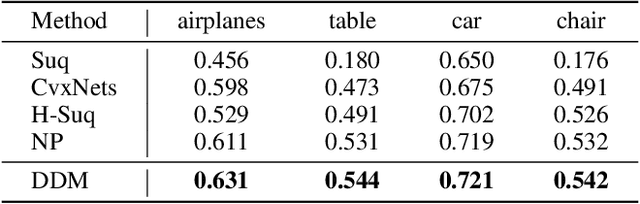
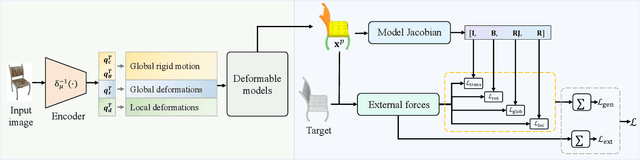
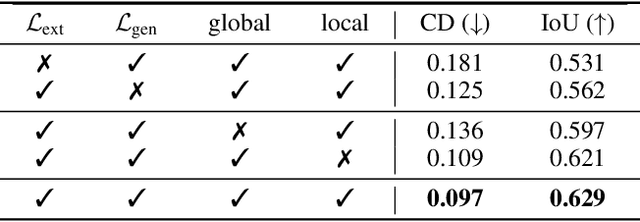
Abstract:The task of shape abstraction with semantic part consistency is challenging due to the complex geometries of natural objects. Recent methods learn to represent an object shape using a set of simple primitives to fit the target. \textcolor{black}{However, in these methods, the primitives used do not always correspond to real parts or lack geometric flexibility for semantic interpretation.} In this paper, we investigate salient and efficient primitive descriptors for accurate shape abstractions, and propose \textit{Deep Deformable Models (DDMs)}. DDM employs global deformations and diffeomorphic local deformations. These properties enable DDM to abstract complex object shapes with significantly fewer primitives that offer broader geometry coverage and finer details. DDM is also capable of learning part-level semantic correspondences due to the differentiable and invertible properties of our primitive deformation. Moreover, DDM learning formulation is based on dynamic and kinematic modeling, which enables joint regularization of each sub-transformation during primitive fitting. Extensive experiments on \textit{ShapeNet} demonstrate that DDM outperforms the state-of-the-art in terms of reconstruction and part consistency by a notable margin.
Training Like a Medical Resident: Universal Medical Image Segmentation via Context Prior Learning
Jun 06, 2023



Abstract:A major enduring focus of clinical workflows is disease analytics and diagnosis, leading to medical imaging datasets where the modalities and annotations are strongly tied to specific clinical objectives. To date, building task-specific segmentation models is intuitive yet a restrictive approach, lacking insights gained from widespread imaging cohorts. Inspired by the training of medical residents, we explore universal medical image segmentation, whose goal is to learn from diverse medical imaging sources covering a range of clinical targets, body regions, and image modalities. Following this paradigm, we propose Hermes, a context prior learning approach that addresses the challenges related to the heterogeneity on data, modality, and annotations in the proposed universal paradigm. In a collection of seven diverse datasets, we demonstrate the appealing merits of the universal paradigm over the traditional task-specific training paradigm. By leveraging the synergy among various tasks, Hermes shows superior performance and model scalability. Our in-depth investigation on two additional datasets reveals Hermes' strong capabilities for transfer learning, incremental learning, and generalization to different downstream tasks. The code is available: https://github.com/yhygao/universal-medical-image-segmentation.
Visual Prompt Tuning for Test-time Domain Adaptation
Oct 10, 2022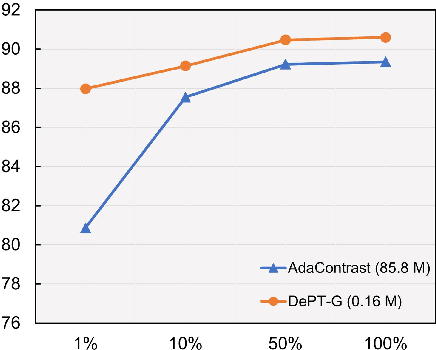
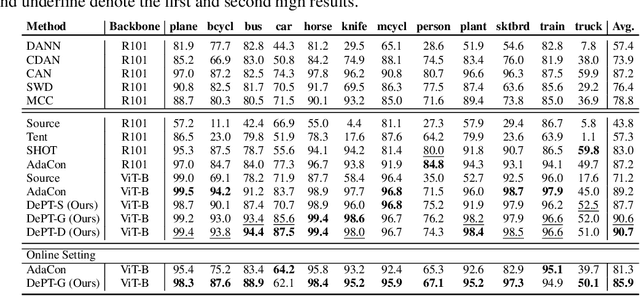
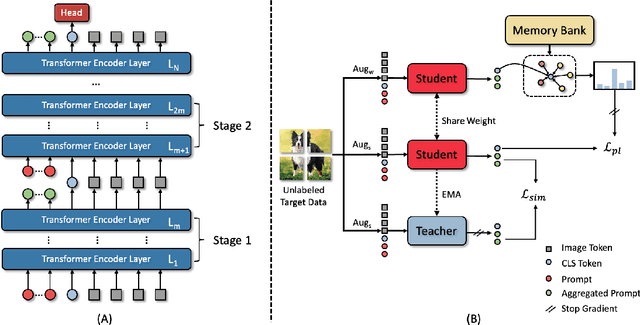
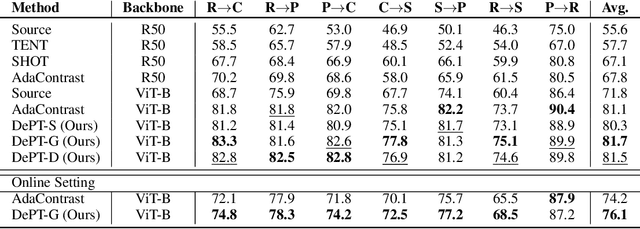
Abstract:Models should have the ability to adapt to unseen data during test-time to avoid performance drop caused by inevitable distribution shifts in real-world deployment scenarios. In this work, we tackle the practical yet challenging test-time adaptation (TTA) problem, where a model adapts to the target domain without accessing the source data. We propose a simple recipe called data-efficient prompt tuning (DePT) with two key ingredients. First, DePT plugs visual prompts into the vision Transformer and only tunes these source-initialized prompts during adaptation. We find such parameter-efficient finetuning can efficiently adapt the model representation to the target domain without overfitting to the noise in the learning objective. Second, DePT bootstraps the source representation to the target domain by memory bank-based online pseudo labeling. A hierarchical self-supervised regularization specially designed for prompts is jointly optimized to alleviate error accumulation during self-training. With much fewer tunable parameters, DePT demonstrates not only state-of-the-art performance on major adaptation benchmarks, but also superior data efficiency, i.e., adaptation with only 1\% or 10\% data without much performance degradation compared to 100\% data. In addition, DePT is also versatile to be extended to online or multi-source TTA settings.
TransFusion: Multi-view Divergent Fusion for Medical Image Segmentation with Transformers
Mar 21, 2022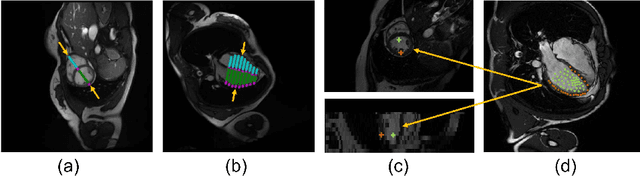
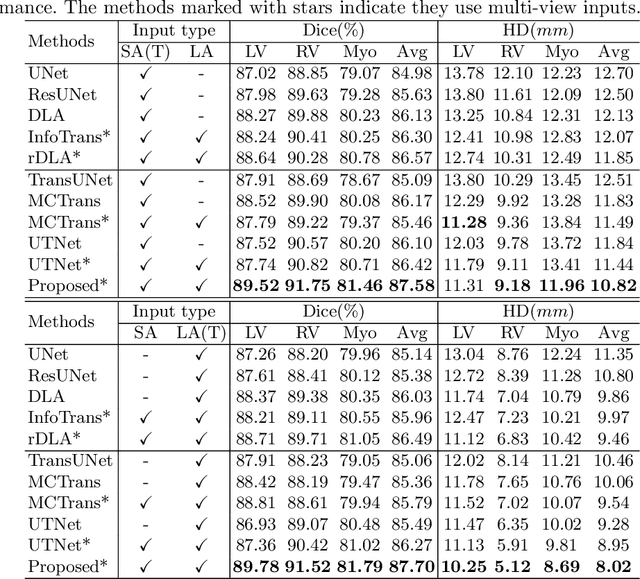
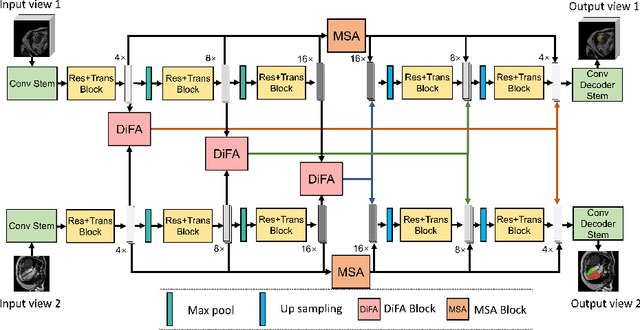
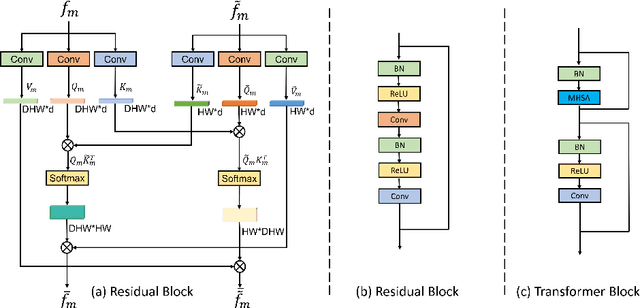
Abstract:Combining information from multi-view images is crucial to improve the performance and robustness of automated methods for disease diagnosis. However, due to the non-alignment characteristics of multi-view images, building correlation and data fusion across views largely remain an open problem. In this study, we present TransFusion, a Transformer-based architecture to merge divergent multi-view imaging information using convolutional layers and powerful attention mechanisms. In particular, the Divergent Fusion Attention (DiFA) module is proposed for rich cross-view context modeling and semantic dependency mining, addressing the critical issue of capturing long-range correlations between unaligned data from different image views. We further propose the Multi-Scale Attention (MSA) to collect global correspondence of multi-scale feature representations. We evaluate TransFusion on the Multi-Disease, Multi-View \& Multi-Center Right Ventricular Segmentation in Cardiac MRI (M\&Ms-2) challenge cohort. TransFusion demonstrates leading performance against the state-of-the-art methods and opens up new perspectives for multi-view imaging integration towards robust medical image segmentation.
A Multi-scale Transformer for Medical Image Segmentation: Architectures, Model Efficiency, and Benchmarks
Mar 03, 2022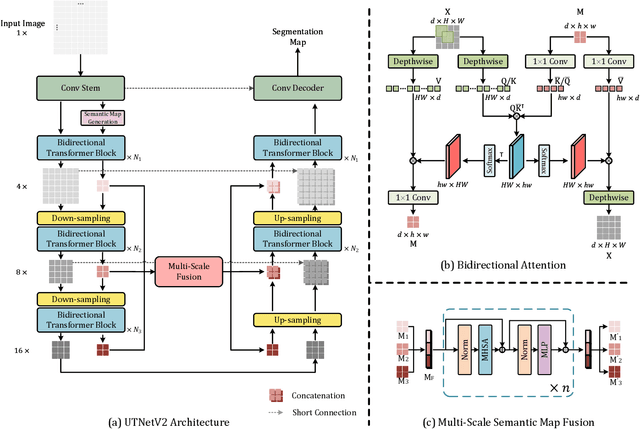
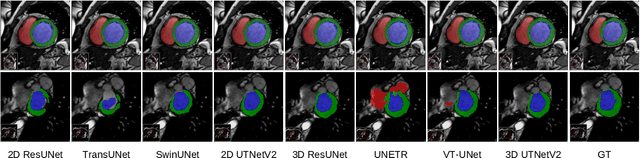
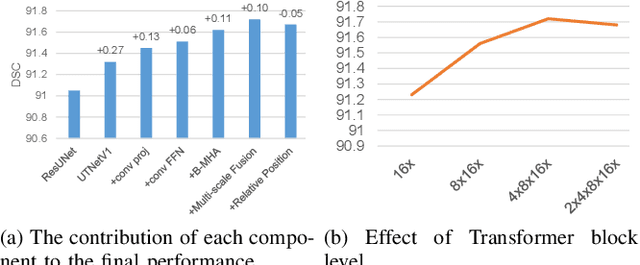
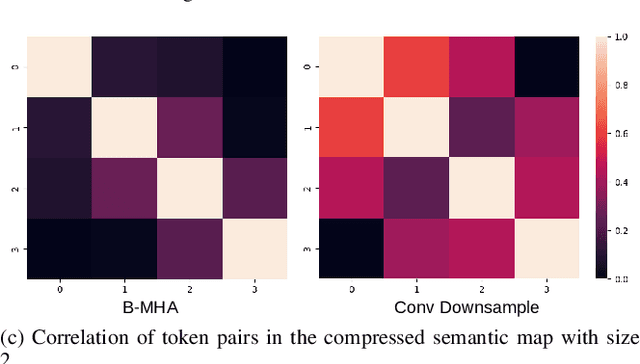
Abstract:Transformers have emerged to be successful in a number of natural language processing and vision tasks, but their potential applications to medical imaging remain largely unexplored due to the unique difficulties of this field. In this study, we present UTNetV2, a simple yet powerful backbone model that combines the strengths of the convolutional neural network and Transformer for enhancing performance and efficiency in medical image segmentation. The critical design of UTNetV2 includes three innovations: (1) We used a hybrid hierarchical architecture by introducing depthwise separable convolution to projection and feed-forward network in the Transformer block, which brings local relationship modeling and desirable properties of CNNs (translation invariance) to Transformer, thus eliminate the requirement of large-scale pre-training. (2) We proposed efficient bidirectional attention (B-MHA) that reduces the quadratic computation complexity of self-attention to linear by introducing an adaptively updated semantic map. The efficient attention makes it possible to capture long-range relationship and correct the fine-grained errors in high-resolution token maps. (3) The semantic maps in the B-MHA allow us to perform semantically and spatially global multi-scale feature fusion without introducing much computational overhead. Furthermore, we provide a fair comparison codebase of CNN-based and Transformer-based on various medical image segmentation tasks to evaluate the merits and defects of both architectures. UTNetV2 demonstrated state-of-the-art performance across various settings, including large-scale datasets, small-scale datasets, 2D and 3D settings.
Modality Bank: Learn multi-modality images across data centers without sharing medical data
Jan 22, 2022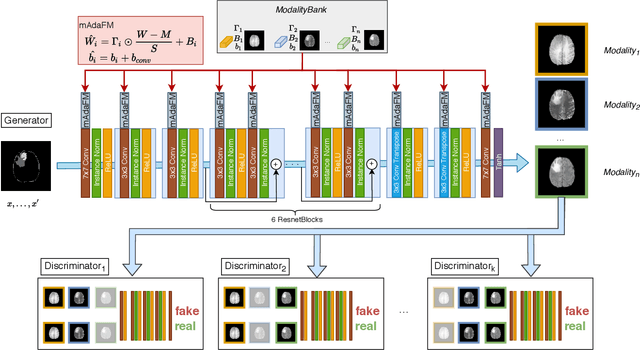
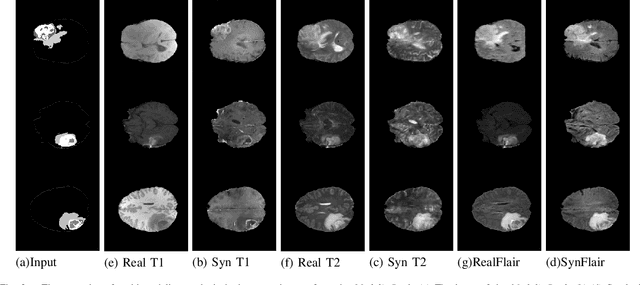
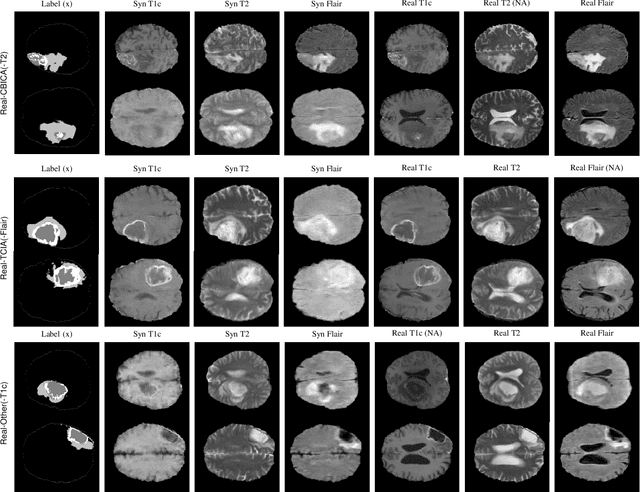
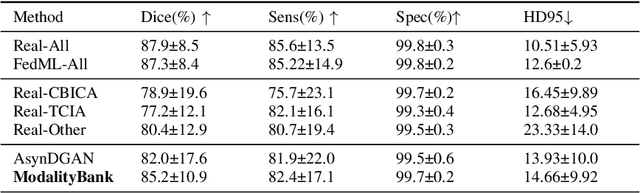
Abstract:Multi-modality images have been widely used and provide comprehensive information for medical image analysis. However, acquiring all modalities among all institutes is costly and often impossible in clinical settings. To leverage more comprehensive multi-modality information, we propose a privacy secured decentralized multi-modality adaptive learning architecture named ModalityBank. Our method could learn a set of effective domain-specific modulation parameters plugged into a common domain-agnostic network. We demonstrate by switching different sets of configurations, the generator could output high-quality images for a specific modality. Our method could also complete the missing modalities across all data centers, thus could be used for modality completion purposes. The downstream task trained from the synthesized multi-modality samples could achieve higher performance than learning from one real data center and achieve close-to-real performance compare with all real images.
UTNet: A Hybrid Transformer Architecture for Medical Image Segmentation
Jul 02, 2021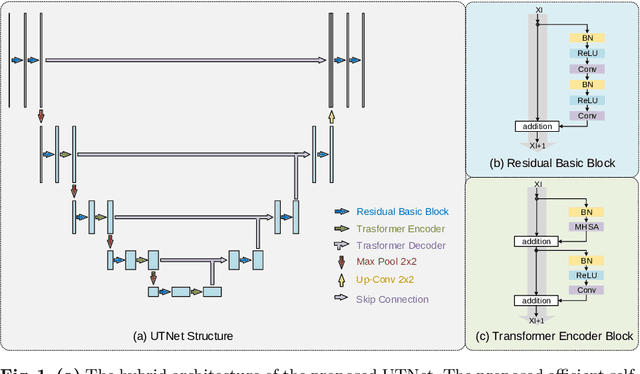
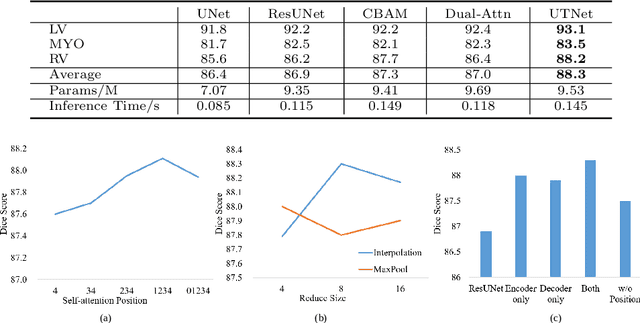
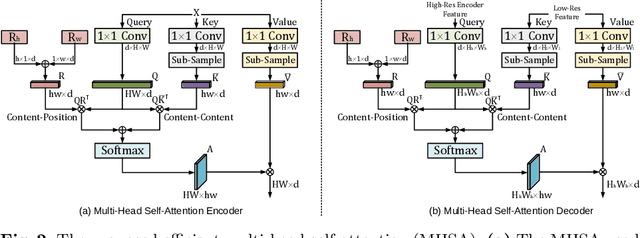
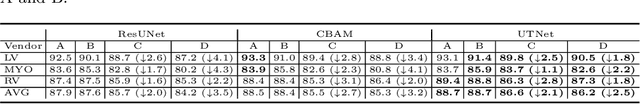
Abstract:Transformer architecture has emerged to be successful in a number of natural language processing tasks. However, its applications to medical vision remain largely unexplored. In this study, we present UTNet, a simple yet powerful hybrid Transformer architecture that integrates self-attention into a convolutional neural network for enhancing medical image segmentation. UTNet applies self-attention modules in both encoder and decoder for capturing long-range dependency at different scales with minimal overhead. To this end, we propose an efficient self-attention mechanism along with relative position encoding that reduces the complexity of self-attention operation significantly from $O(n^2)$ to approximate $O(n)$. A new self-attention decoder is also proposed to recover fine-grained details from the skipped connections in the encoder. Our approach addresses the dilemma that Transformer requires huge amounts of data to learn vision inductive bias. Our hybrid layer design allows the initialization of Transformer into convolutional networks without a need of pre-training. We have evaluated UTNet on the multi-label, multi-vendor cardiac magnetic resonance imaging cohort. UTNet demonstrates superior segmentation performance and robustness against the state-of-the-art approaches, holding the promise to generalize well on other medical image segmentations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge