Xipeng Pan
WSSS4LUAD: Grand Challenge on Weakly-supervised Tissue Semantic Segmentation for Lung Adenocarcinoma
Apr 14, 2022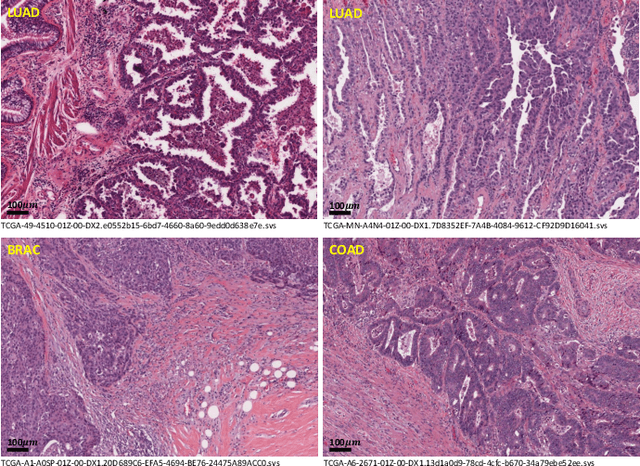
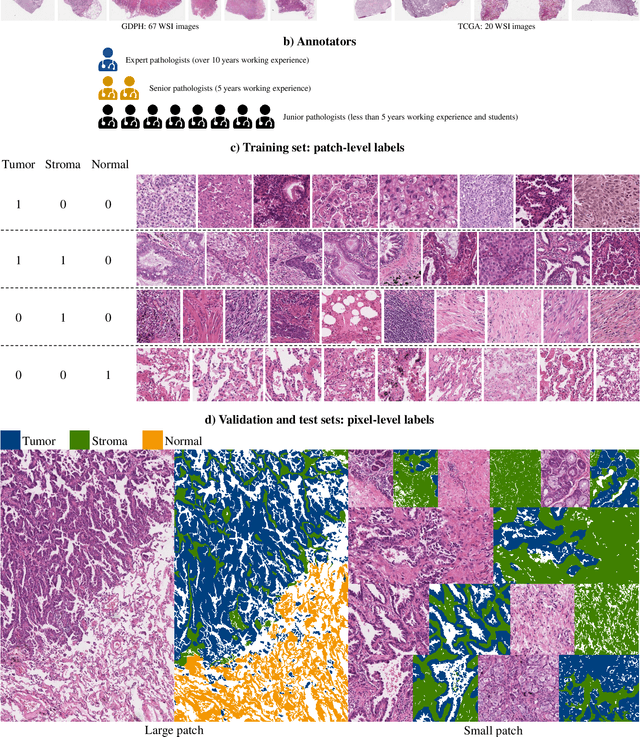
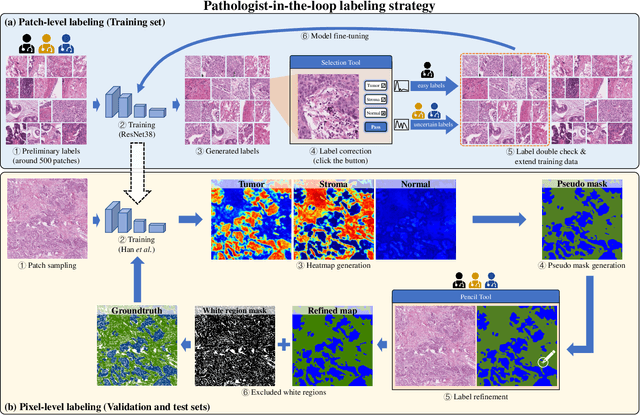
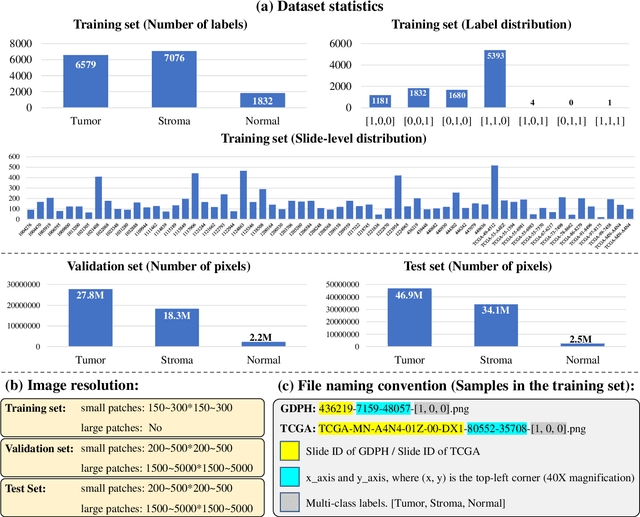
Abstract:Lung cancer is the leading cause of cancer death worldwide, and adenocarcinoma (LUAD) is the most common subtype. Exploiting the potential value of the histopathology images can promote precision medicine in oncology. Tissue segmentation is the basic upstream task of histopathology image analysis. Existing deep learning models have achieved superior segmentation performance but require sufficient pixel-level annotations, which is time-consuming and expensive. To enrich the label resources of LUAD and to alleviate the annotation efforts, we organize this challenge WSSS4LUAD to call for the outstanding weakly-supervised semantic segmentation (WSSS) techniques for histopathology images of LUAD. Participants have to design the algorithm to segment tumor epithelial, tumor-associated stroma and normal tissue with only patch-level labels. This challenge includes 10,091 patch-level annotations (the training set) and over 130 million labeled pixels (the validation and test sets), from 87 WSIs (67 from GDPH, 20 from TCGA). All the labels were generated by a pathologist-in-the-loop pipeline with the help of AI models and checked by the label review board. Among 532 registrations, 28 teams submitted the results in the test phase with over 1,000 submissions. Finally, the first place team achieved mIoU of 0.8413 (tumor: 0.8389, stroma: 0.7931, normal: 0.8919). According to the technical reports of the top-tier teams, CAM is still the most popular approach in WSSS. Cutmix data augmentation has been widely adopted to generate more reliable samples. With the success of this challenge, we believe that WSSS approaches with patch-level annotations can be a complement to the traditional pixel annotations while reducing the annotation efforts. The entire dataset has been released to encourage more researches on computational pathology in LUAD and more novel WSSS techniques.
A Standardized Pipeline for Colon Nuclei Identification and Counting Challenge
Mar 20, 2022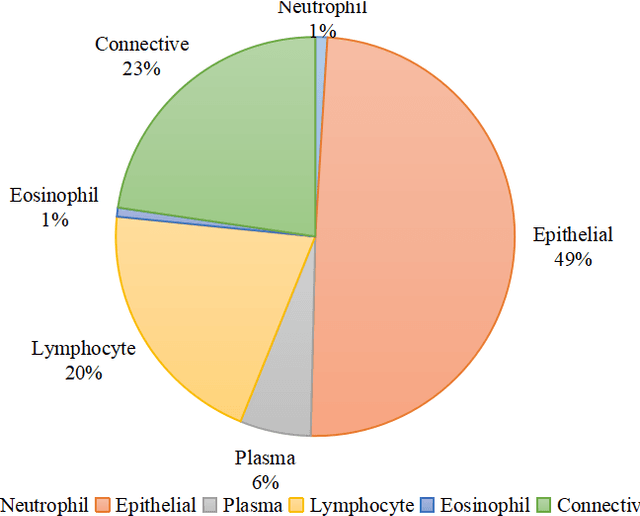
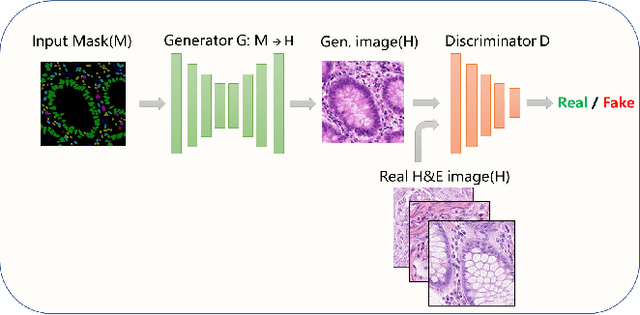
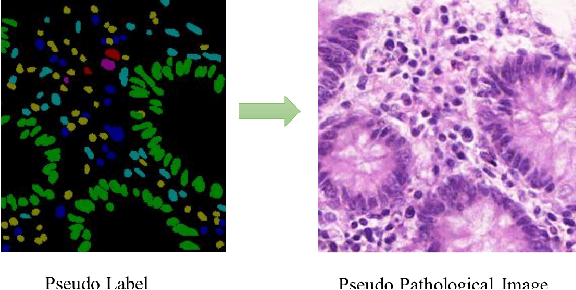
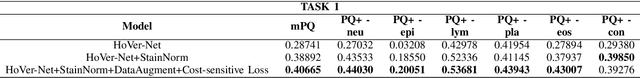
Abstract:Nuclear segmentation and classification is an essential step for computational pathology. TIA lab from Warwick University organized a nuclear segmentation and classification challenge (CoNIC) for H&E stained histopathology images in colorectal cancer with two highly correlated tasks, nuclei segmentation and classification task and cellular composition task. There are a few obstacles we have to address in this challenge, 1) limited training samples, 2) color variation, 3) imbalanced annotations, 4) similar morphological appearance among classes. To deal with these challenges, we proposed a standardized pipeline for nuclear segmentation and classification by integrating several pluggable components. First, we built a GAN-based model to automatically generate pseudo images for data augmentation. Then we trained a self-supervised stain normalization model to solve the color variation problem. Next we constructed a baseline model HoVer-Net with cost-sensitive loss to encourage the model pay more attention on the minority classes. According to the results of the leaderboard, our proposed pipeline achieves 0.40665 mPQ+ (Rank 49th) and 0.62199 r2 (Rank 10th) in the preliminary test phase.
PHTrans: Parallelly Aggregating Global and Local Representations for Medical Image Segmentation
Mar 13, 2022
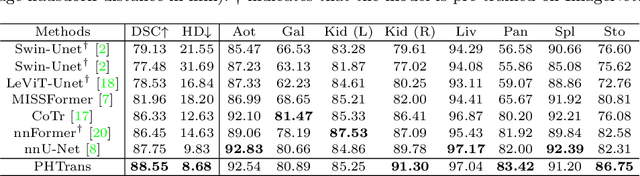

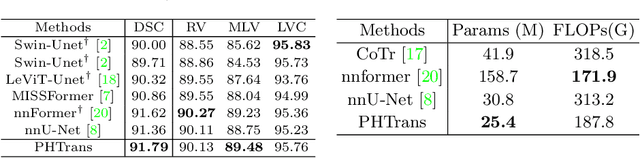
Abstract:The success of Transformer in computer vision has attracted increasing attention in the medical imaging community. Especially for medical image segmentation, many excellent hybrid architectures based on convolutional neural networks (CNNs) and Transformer have been presented and achieve impressive performance. However, most of these methods, which embed modular Transformer into CNNs, struggle to reach their full potential. In this paper, we propose a novel hybrid architecture for medical image segmentation called PHTrans, which parallelly hybridizes Transformer and CNN in main building blocks to produce hierarchical representations from global and local features and adaptively aggregate them, aiming to fully exploit their strengths to obtain better segmentation performance. Specifically, PHTrans follows the U-shaped encoder-decoder design and introduces the parallel hybird module in deep stages, where convolution blocks and the modified 3D Swin Transformer learn local features and global dependencies separately, then a sequence-to-volume operation unifies the dimensions of the outputs to achieve feature aggregation. Extensive experimental results on both Multi-Atlas Labeling Beyond the Cranial Vault and Automated Cardiac Diagnosis Challeng datasets corroborate its effectiveness, consistently outperforming state-of-the-art methods.
PDBL: Improving Histopathological Tissue Classification with Plug-and-Play Pyramidal Deep-Broad Learning
Nov 04, 2021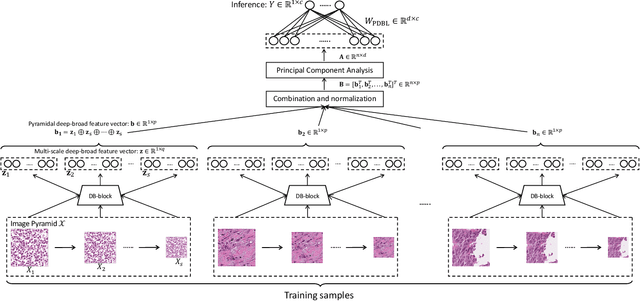
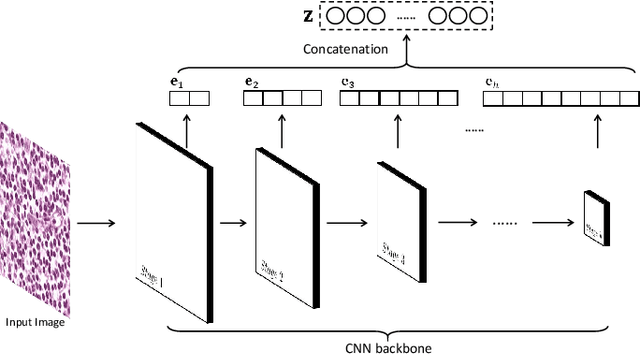

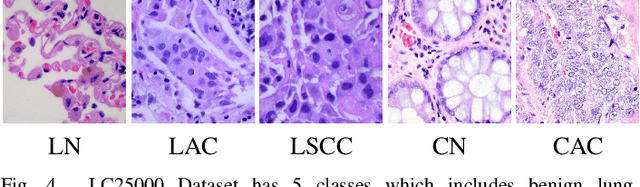
Abstract:Histopathological tissue classification is a fundamental task in pathomics cancer research. Precisely differentiating different tissue types is a benefit for the downstream researches, like cancer diagnosis, prognosis and etc. Existing works mostly leverage the popular classification backbones in computer vision to achieve histopathological tissue classification. In this paper, we proposed a super lightweight plug-and-play module, named Pyramidal Deep-Broad Learning (PDBL), for any well-trained classification backbone to further improve the classification performance without a re-training burden. We mimic how pathologists observe pathology slides in different magnifications and construct an image pyramid for the input image in order to obtain the pyramidal contextual information. For each level in the pyramid, we extract the multi-scale deep-broad features by our proposed Deep-Broad block (DB-block). We equipped PDBL in three popular classification backbones, ShuffLeNetV2, EfficientNetb0, and ResNet50 to evaluate the effectiveness and efficiency of our proposed module on two datasets (Kather Multiclass Dataset and the LC25000 Dataset). Experimental results demonstrate the proposed PDBL can steadily improve the tissue-level classification performance for any CNN backbones, especially for the lightweight models when given a small among of training samples (less than 10%), which greatly saves the computational time and annotation efforts.
Multi-Layer Pseudo-Supervision for Histopathology Tissue Semantic Segmentation using Patch-level Classification Labels
Oct 14, 2021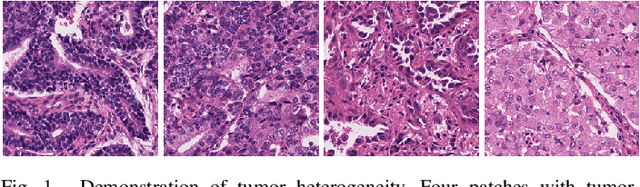

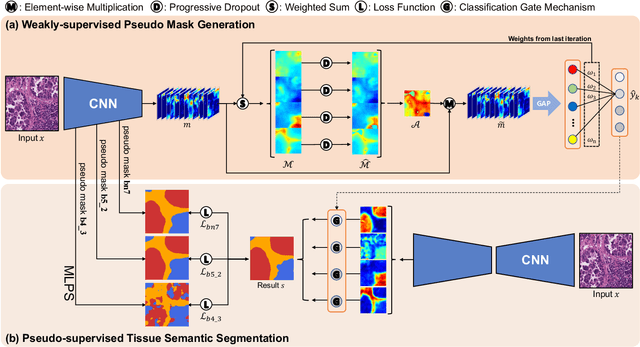
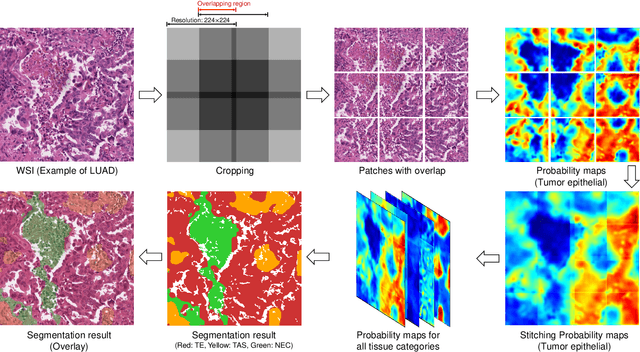
Abstract:Tissue-level semantic segmentation is a vital step in computational pathology. Fully-supervised models have already achieved outstanding performance with dense pixel-level annotations. However, drawing such labels on the giga-pixel whole slide images is extremely expensive and time-consuming. In this paper, we use only patch-level classification labels to achieve tissue semantic segmentation on histopathology images, finally reducing the annotation efforts. We proposed a two-step model including a classification and a segmentation phases. In the classification phase, we proposed a CAM-based model to generate pseudo masks by patch-level labels. In the segmentation phase, we achieved tissue semantic segmentation by our proposed Multi-Layer Pseudo-Supervision. Several technical novelties have been proposed to reduce the information gap between pixel-level and patch-level annotations. As a part of this paper, we introduced a new weakly-supervised semantic segmentation (WSSS) dataset for lung adenocarcinoma (LUAD-HistoSeg). We conducted several experiments to evaluate our proposed model on two datasets. Our proposed model outperforms two state-of-the-art WSSS approaches. Note that we can achieve comparable quantitative and qualitative results with the fully-supervised model, with only around a 2\% gap for MIoU and FwIoU. By comparing with manual labeling, our model can greatly save the annotation time from hours to minutes. The source code is available at: \url{https://github.com/ChuHan89/WSSS-Tissue}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge